- Skip to secondary menu
- Skip to main content
- Skip to primary sidebar
Statistics By Jim
Making statistics intuitive

Null Hypothesis: Definition, Rejecting & Examples
By Jim Frost 6 Comments
What is a Null Hypothesis?
The null hypothesis in statistics states that there is no difference between groups or no relationship between variables. It is one of two mutually exclusive hypotheses about a population in a hypothesis test.

- Null Hypothesis H 0 : No effect exists in the population.
- Alternative Hypothesis H A : The effect exists in the population.
In every study or experiment, researchers assess an effect or relationship. This effect can be the effectiveness of a new drug, building material, or other intervention that has benefits. There is a benefit or connection that the researchers hope to identify. Unfortunately, no effect may exist. In statistics, we call this lack of an effect the null hypothesis. Researchers assume that this notion of no effect is correct until they have enough evidence to suggest otherwise, similar to how a trial presumes innocence.
In this context, the analysts don’t necessarily believe the null hypothesis is correct. In fact, they typically want to reject it because that leads to more exciting finds about an effect or relationship. The new vaccine works!
You can think of it as the default theory that requires sufficiently strong evidence to reject. Like a prosecutor, researchers must collect sufficient evidence to overturn the presumption of no effect. Investigators must work hard to set up a study and a data collection system to obtain evidence that can reject the null hypothesis.
Related post : What is an Effect in Statistics?
Null Hypothesis Examples
Null hypotheses start as research questions that the investigator rephrases as a statement indicating there is no effect or relationship.
| Does the vaccine prevent infections? | The vaccine does not affect the infection rate. |
| Does the new additive increase product strength? | The additive does not affect mean product strength. |
| Does the exercise intervention increase bone mineral density? | The intervention does not affect bone mineral density. |
| As screen time increases, does test performance decrease? | There is no relationship between screen time and test performance. |
After reading these examples, you might think they’re a bit boring and pointless. However, the key is to remember that the null hypothesis defines the condition that the researchers need to discredit before suggesting an effect exists.
Let’s see how you reject the null hypothesis and get to those more exciting findings!
When to Reject the Null Hypothesis
So, you want to reject the null hypothesis, but how and when can you do that? To start, you’ll need to perform a statistical test on your data. The following is an overview of performing a study that uses a hypothesis test.
The first step is to devise a research question and the appropriate null hypothesis. After that, the investigators need to formulate an experimental design and data collection procedures that will allow them to gather data that can answer the research question. Then they collect the data. For more information about designing a scientific study that uses statistics, read my post 5 Steps for Conducting Studies with Statistics .
After data collection is complete, statistics and hypothesis testing enter the picture. Hypothesis testing takes your sample data and evaluates how consistent they are with the null hypothesis. The p-value is a crucial part of the statistical results because it quantifies how strongly the sample data contradict the null hypothesis.
When the sample data provide sufficient evidence, you can reject the null hypothesis. In a hypothesis test, this process involves comparing the p-value to your significance level .
Rejecting the Null Hypothesis
Reject the null hypothesis when the p-value is less than or equal to your significance level. Your sample data favor the alternative hypothesis, which suggests that the effect exists in the population. For a mnemonic device, remember—when the p-value is low, the null must go!
When you can reject the null hypothesis, your results are statistically significant. Learn more about Statistical Significance: Definition & Meaning .
Failing to Reject the Null Hypothesis
Conversely, when the p-value is greater than your significance level, you fail to reject the null hypothesis. The sample data provides insufficient data to conclude that the effect exists in the population. When the p-value is high, the null must fly!
Note that failing to reject the null is not the same as proving it. For more information about the difference, read my post about Failing to Reject the Null .
That’s a very general look at the process. But I hope you can see how the path to more exciting findings depends on being able to rule out the less exciting null hypothesis that states there’s nothing to see here!
Let’s move on to learning how to write the null hypothesis for different types of effects, relationships, and tests.
Related posts : How Hypothesis Tests Work and Interpreting P-values
How to Write a Null Hypothesis
The null hypothesis varies by the type of statistic and hypothesis test. Remember that inferential statistics use samples to draw conclusions about populations. Consequently, when you write a null hypothesis, it must make a claim about the relevant population parameter . Further, that claim usually indicates that the effect does not exist in the population. Below are typical examples of writing a null hypothesis for various parameters and hypothesis tests.
Related posts : Descriptive vs. Inferential Statistics and Populations, Parameters, and Samples in Inferential Statistics
Group Means
T-tests and ANOVA assess the differences between group means. For these tests, the null hypothesis states that there is no difference between group means in the population. In other words, the experimental conditions that define the groups do not affect the mean outcome. Mu (µ) is the population parameter for the mean, and you’ll need to include it in the statement for this type of study.
For example, an experiment compares the mean bone density changes for a new osteoporosis medication. The control group does not receive the medicine, while the treatment group does. The null states that the mean bone density changes for the control and treatment groups are equal.
- Null Hypothesis H 0 : Group means are equal in the population: µ 1 = µ 2 , or µ 1 – µ 2 = 0
- Alternative Hypothesis H A : Group means are not equal in the population: µ 1 ≠ µ 2 , or µ 1 – µ 2 ≠ 0.
Group Proportions
Proportions tests assess the differences between group proportions. For these tests, the null hypothesis states that there is no difference between group proportions. Again, the experimental conditions did not affect the proportion of events in the groups. P is the population proportion parameter that you’ll need to include.
For example, a vaccine experiment compares the infection rate in the treatment group to the control group. The treatment group receives the vaccine, while the control group does not. The null states that the infection rates for the control and treatment groups are equal.
- Null Hypothesis H 0 : Group proportions are equal in the population: p 1 = p 2 .
- Alternative Hypothesis H A : Group proportions are not equal in the population: p 1 ≠ p 2 .
Correlation and Regression Coefficients
Some studies assess the relationship between two continuous variables rather than differences between groups.
In these studies, analysts often use either correlation or regression analysis . For these tests, the null states that there is no relationship between the variables. Specifically, it says that the correlation or regression coefficient is zero. As one variable increases, there is no tendency for the other variable to increase or decrease. Rho (ρ) is the population correlation parameter and beta (β) is the regression coefficient parameter.
For example, a study assesses the relationship between screen time and test performance. The null states that there is no correlation between this pair of variables. As screen time increases, test performance does not tend to increase or decrease.
- Null Hypothesis H 0 : The correlation in the population is zero: ρ = 0.
- Alternative Hypothesis H A : The correlation in the population is not zero: ρ ≠ 0.
For all these cases, the analysts define the hypotheses before the study. After collecting the data, they perform a hypothesis test to determine whether they can reject the null hypothesis.
The preceding examples are all for two-tailed hypothesis tests. To learn about one-tailed tests and how to write a null hypothesis for them, read my post One-Tailed vs. Two-Tailed Tests .
Related post : Understanding Correlation
Neyman, J; Pearson, E. S. (January 1, 1933). On the Problem of the most Efficient Tests of Statistical Hypotheses . Philosophical Transactions of the Royal Society A . 231 (694–706): 289–337.
Share this:

Reader Interactions
January 11, 2024 at 2:57 pm
Thanks for the reply.
January 10, 2024 at 1:23 pm
Hi Jim, In your comment you state that equivalence test null and alternate hypotheses are reversed. For hypothesis tests of data fits to a probability distribution, the null hypothesis is that the probability distribution fits the data. Is this correct?
January 10, 2024 at 2:15 pm
Those two separate things, equivalence testing and normality tests. But, yes, you’re correct for both.
Hypotheses are switched for equivalence testing. You need to “work” (i.e., collect a large sample of good quality data) to be able to reject the null that the groups are different to be able to conclude they’re the same.
With typical hypothesis tests, if you have low quality data and a low sample size, you’ll fail to reject the null that they’re the same, concluding they’re equivalent. But that’s more a statement about the low quality and small sample size than anything to do with the groups being equal.
So, equivalence testing make you work to obtain a finding that the groups are the same (at least within some amount you define as a trivial difference).
For normality testing, and other distribution tests, the null states that the data follow the distribution (normal or whatever). If you reject the null, you have sufficient evidence to conclude that your sample data don’t follow the probability distribution. That’s a rare case where you hope to fail to reject the null. And it suffers from the problem I describe above where you might fail to reject the null simply because you have a small sample size. In that case, you’d conclude the data follow the probability distribution but it’s more that you don’t have enough data for the test to register the deviation. In this scenario, if you had a larger sample size, you’d reject the null and conclude it doesn’t follow that distribution.
I don’t know of any equivalence testing type approach for distribution fit tests where you’d need to work to show the data follow a distribution, although I haven’t looked for one either!
February 20, 2022 at 9:26 pm
Is a null hypothesis regularly (always) stated in the negative? “there is no” or “does not”
February 23, 2022 at 9:21 pm
Typically, the null hypothesis includes an equal sign. The null hypothesis states that the population parameter equals a particular value. That value is usually one that represents no effect. In the case of a one-sided hypothesis test, the null still contains an equal sign but it’s “greater than or equal to” or “less than or equal to.” If you wanted to translate the null hypothesis from its native mathematical expression, you could use the expression “there is no effect.” But the mathematical form more specifically states what it’s testing.
It’s the alternative hypothesis that typically contains does not equal.
There are some exceptions. For example, in an equivalence test where the researchers want to show that two things are equal, the null hypothesis states that they’re not equal.
In short, the null hypothesis states the condition that the researchers hope to reject. They need to work hard to set up an experiment and data collection that’ll gather enough evidence to be able to reject the null condition.
February 15, 2022 at 9:32 am
Dear sir I always read your notes on Research methods.. Kindly tell is there any available Book on all these..wonderfull Urgent
Comments and Questions Cancel reply

- Science Notes Posts
- Contact Science Notes
- Todd Helmenstine Biography
- Anne Helmenstine Biography
- Free Printable Periodic Tables (PDF and PNG)
- Periodic Table Wallpapers
- Interactive Periodic Table
- Periodic Table Posters
- Science Experiments for Kids
- How to Grow Crystals
- Chemistry Projects
- Fire and Flames Projects
- Holiday Science
- Chemistry Problems With Answers
- Physics Problems
- Unit Conversion Example Problems
- Chemistry Worksheets
- Biology Worksheets
- Periodic Table Worksheets
- Physical Science Worksheets
- Science Lab Worksheets
- My Amazon Books
Null Hypothesis Examples

The null hypothesis (H 0 ) is the hypothesis that states there is no statistical difference between two sample sets. In other words, it assumes the independent variable does not have an effect on the dependent variable in a scientific experiment .
The null hypothesis is the most powerful type of hypothesis in the scientific method because it’s the easiest one to test with a high confidence level using statistics. If the null hypothesis is accepted, then it’s evidence any observed differences between two experiment groups are due to random chance. If the null hypothesis is rejected, then it’s strong evidence there is a true difference between test sets or that the independent variable affects the dependent variable.
- The null hypothesis is a nullifiable hypothesis. A researcher seeks to reject it because this result strongly indicates observed differences are real and not just due to chance.
- The null hypothesis may be accepted or rejected, but not proven. There is always a level of confidence in the outcome.
What Is the Null Hypothesis?
The null hypothesis is written as H 0 , which is read as H-zero, H-nought, or H-null. It is associated with another hypothesis, called the alternate or alternative hypothesis H A or H 1 . When the null hypothesis and alternate hypothesis are written mathematically, they cover all possible outcomes of an experiment.
An experimenter tests the null hypothesis with a statistical analysis called a significance test. The significance test determines the likelihood that the results of the test are not due to chance. Usually, a researcher uses a confidence level of 95% or 99% (p-value of 0.05 or 0.01). But, even if the confidence in the test is high, there is always a small chance the outcome is incorrect. This means you can’t prove a null hypothesis. It’s also a good reason why it’s important to repeat experiments.
Exact and Inexact Null Hypothesis
The most common type of null hypothesis assumes no difference between two samples or groups or no measurable effect of a treatment. This is the exact hypothesis . If you’re asked to state a null hypothesis for a science class, this is the one to write. It is the easiest type of hypothesis to test and is the only one accepted for certain types of analysis. Examples include:
There is no difference between two groups H 0 : μ 1 = μ 2 (where H 0 = the null hypothesis, μ 1 = the mean of population 1, and μ 2 = the mean of population 2)
Both groups have value of 100 (or any number or quality) H 0 : μ = 100
However, sometimes a researcher may test an inexact hypothesis . This type of hypothesis specifies ranges or intervals. Examples include:
Recovery time from a treatment is the same or worse than a placebo: H 0 : μ ≥ placebo time
There is a 5% or less difference between two groups: H 0 : 95 ≤ μ ≤ 105
An inexact hypothesis offers “directionality” about a phenomenon. For example, an exact hypothesis can indicate whether or not a treatment has an effect, while an inexact hypothesis can tell whether an effect is positive of negative. However, an inexact hypothesis may be harder to test and some scientists and statisticians disagree about whether it’s a true null hypothesis .
How to State the Null Hypothesis
To state the null hypothesis, first state what you expect the experiment to show. Then, rephrase the statement in a form that assumes there is no relationship between the variables or that a treatment has no effect.
Example: A researcher tests whether a new drug speeds recovery time from a certain disease. The average recovery time without treatment is 3 weeks.
- State the goal of the experiment: “I hope the average recovery time with the new drug will be less than 3 weeks.”
- Rephrase the hypothesis to assume the treatment has no effect: “If the drug doesn’t shorten recovery time, then the average time will be 3 weeks or longer.” Mathematically: H 0 : μ ≥ 3
This null hypothesis (inexact hypothesis) covers both the scenario in which the drug has no effect and the one in which the drugs makes the recovery time longer. The alternate hypothesis is that average recovery time will be less than three weeks:
H A : μ < 3
Of course, the researcher could test the no-effect hypothesis (exact null hypothesis): H 0 : μ = 3
The danger of testing this hypothesis is that rejecting it only implies the drug affected recovery time (not whether it made it better or worse). This is because the alternate hypothesis is:
H A : μ ≠ 3 (which includes μ <3 and μ >3)
Even though the no-effect null hypothesis yields less information, it’s used because it’s easier to test using statistics. Basically, testing whether something is unchanged/changed is easier than trying to quantify the nature of the change.
Remember, a researcher hopes to reject the null hypothesis because this supports the alternate hypothesis. Also, be sure the null and alternate hypothesis cover all outcomes. Finally, remember a simple true/false, equal/unequal, yes/no exact hypothesis is easier to test than a more complex inexact hypothesis.
| Does chewing willow bark relieve pain? | Pain relief is the same compared with a . (exact) Pain relief after chewing willow bark is the same or worse versus taking a placebo. (inexact) | Pain relief is different compared with a placebo. (exact) Pain relief is better compared to a placebo. (inexact) |
| Do cats care about the shape of their food? | Cats show no food preference based on shape. (exact) | Cat show a food preference based on shape. (exact) |
| Do teens use mobile devices more than adults? | Teens and adults use mobile devices the same amount. (exact) Teens use mobile devices less than or equal to adults. (inexact) | Teens and adults used mobile devices different amounts. (exact) Teens use mobile devices more than adults. (inexact) |
| Does the color of light influence plant growth? | The color of light has no effect on plant growth. (exact) | The color of light affects plant growth. (exact) |
- Adèr, H. J.; Mellenbergh, G. J. & Hand, D. J. (2007). Advising on Research Methods: A Consultant’s Companion . Huizen, The Netherlands: Johannes van Kessel Publishing. ISBN 978-90-79418-01-5 .
- Cox, D. R. (2006). Principles of Statistical Inference . Cambridge University Press. ISBN 978-0-521-68567-2 .
- Everitt, Brian (1998). The Cambridge Dictionary of Statistics . Cambridge, UK New York: Cambridge University Press. ISBN 978-0521593465.
- Weiss, Neil A. (1999). Introductory Statistics (5th ed.). ISBN 9780201598773.
Related Posts
Have a thesis expert improve your writing
Check your thesis for plagiarism in 10 minutes, generate your apa citations for free.
- Knowledge Base
- Null and Alternative Hypotheses | Definitions & Examples
Null and Alternative Hypotheses | Definitions & Examples
Published on 5 October 2022 by Shaun Turney . Revised on 6 December 2022.
The null and alternative hypotheses are two competing claims that researchers weigh evidence for and against using a statistical test :
- Null hypothesis (H 0 ): There’s no effect in the population .
- Alternative hypothesis (H A ): There’s an effect in the population.
The effect is usually the effect of the independent variable on the dependent variable .
Table of contents
Answering your research question with hypotheses, what is a null hypothesis, what is an alternative hypothesis, differences between null and alternative hypotheses, how to write null and alternative hypotheses, frequently asked questions about null and alternative hypotheses.
The null and alternative hypotheses offer competing answers to your research question . When the research question asks “Does the independent variable affect the dependent variable?”, the null hypothesis (H 0 ) answers “No, there’s no effect in the population.” On the other hand, the alternative hypothesis (H A ) answers “Yes, there is an effect in the population.”
The null and alternative are always claims about the population. That’s because the goal of hypothesis testing is to make inferences about a population based on a sample . Often, we infer whether there’s an effect in the population by looking at differences between groups or relationships between variables in the sample.
You can use a statistical test to decide whether the evidence favors the null or alternative hypothesis. Each type of statistical test comes with a specific way of phrasing the null and alternative hypothesis. However, the hypotheses can also be phrased in a general way that applies to any test.
The null hypothesis is the claim that there’s no effect in the population.
If the sample provides enough evidence against the claim that there’s no effect in the population ( p ≤ α), then we can reject the null hypothesis . Otherwise, we fail to reject the null hypothesis.
Although “fail to reject” may sound awkward, it’s the only wording that statisticians accept. Be careful not to say you “prove” or “accept” the null hypothesis.
Null hypotheses often include phrases such as “no effect”, “no difference”, or “no relationship”. When written in mathematical terms, they always include an equality (usually =, but sometimes ≥ or ≤).
Examples of null hypotheses
The table below gives examples of research questions and null hypotheses. There’s always more than one way to answer a research question, but these null hypotheses can help you get started.
| ( ) | ||
| Does tooth flossing affect the number of cavities? | Tooth flossing has on the number of cavities. | test: The mean number of cavities per person does not differ between the flossing group (µ ) and the non-flossing group (µ ) in the population; µ = µ . |
| Does the amount of text highlighted in the textbook affect exam scores? | The amount of text highlighted in the textbook has on exam scores. | : There is no relationship between the amount of text highlighted and exam scores in the population; β = 0. |
| Does daily meditation decrease the incidence of depression? | Daily meditation the incidence of depression.* | test: The proportion of people with depression in the daily-meditation group ( ) is greater than or equal to the no-meditation group ( ) in the population; ≥ . |
*Note that some researchers prefer to always write the null hypothesis in terms of “no effect” and “=”. It would be fine to say that daily meditation has no effect on the incidence of depression and p 1 = p 2 .
The alternative hypothesis (H A ) is the other answer to your research question . It claims that there’s an effect in the population.
Often, your alternative hypothesis is the same as your research hypothesis. In other words, it’s the claim that you expect or hope will be true.
The alternative hypothesis is the complement to the null hypothesis. Null and alternative hypotheses are exhaustive, meaning that together they cover every possible outcome. They are also mutually exclusive, meaning that only one can be true at a time.
Alternative hypotheses often include phrases such as “an effect”, “a difference”, or “a relationship”. When alternative hypotheses are written in mathematical terms, they always include an inequality (usually ≠, but sometimes > or <). As with null hypotheses, there are many acceptable ways to phrase an alternative hypothesis.
Examples of alternative hypotheses
The table below gives examples of research questions and alternative hypotheses to help you get started with formulating your own.
| Does tooth flossing affect the number of cavities? | Tooth flossing has an on the number of cavities. | test: The mean number of cavities per person differs between the flossing group (µ ) and the non-flossing group (µ ) in the population; µ ≠ µ . |
| Does the amount of text highlighted in a textbook affect exam scores? | The amount of text highlighted in the textbook has an on exam scores. | : There is a relationship between the amount of text highlighted and exam scores in the population; β ≠ 0. |
| Does daily meditation decrease the incidence of depression? | Daily meditation the incidence of depression. | test: The proportion of people with depression in the daily-meditation group ( ) is less than the no-meditation group ( ) in the population; < . |
Null and alternative hypotheses are similar in some ways:
- They’re both answers to the research question
- They both make claims about the population
- They’re both evaluated by statistical tests.
However, there are important differences between the two types of hypotheses, summarized in the following table.
| A claim that there is in the population. | A claim that there is in the population. | |
|
| ||
| Equality symbol (=, ≥, or ≤) | Inequality symbol (≠, <, or >) | |
| Rejected | Supported | |
| Failed to reject | Not supported |
To help you write your hypotheses, you can use the template sentences below. If you know which statistical test you’re going to use, you can use the test-specific template sentences. Otherwise, you can use the general template sentences.
The only thing you need to know to use these general template sentences are your dependent and independent variables. To write your research question, null hypothesis, and alternative hypothesis, fill in the following sentences with your variables:
Does independent variable affect dependent variable ?
- Null hypothesis (H 0 ): Independent variable does not affect dependent variable .
- Alternative hypothesis (H A ): Independent variable affects dependent variable .
Test-specific
Once you know the statistical test you’ll be using, you can write your hypotheses in a more precise and mathematical way specific to the test you chose. The table below provides template sentences for common statistical tests.
| ( ) | ||
| test
with two groups | The mean dependent variable does not differ between group 1 (µ ) and group 2 (µ ) in the population; µ = µ . | The mean dependent variable differs between group 1 (µ ) and group 2 (µ ) in the population; µ ≠ µ . |
| with three groups | The mean dependent variable does not differ between group 1 (µ ), group 2 (µ ), and group 3 (µ ) in the population; µ = µ = µ . | The mean dependent variable of group 1 (µ ), group 2 (µ ), and group 3 (µ ) are not all equal in the population. |
| There is no correlation between independent variable and dependent variable in the population; ρ = 0. | There is a correlation between independent variable and dependent variable in the population; ρ ≠ 0. | |
| There is no relationship between independent variable and dependent variable in the population; β = 0. | There is a relationship between independent variable and dependent variable in the population; β ≠ 0. | |
| Two-proportions test | The dependent variable expressed as a proportion does not differ between group 1 ( ) and group 2 ( ) in the population; = . | The dependent variable expressed as a proportion differs between group 1 ( ) and group 2 ( ) in the population; ≠ . |
Note: The template sentences above assume that you’re performing one-tailed tests . One-tailed tests are appropriate for most studies.
The null hypothesis is often abbreviated as H 0 . When the null hypothesis is written using mathematical symbols, it always includes an equality symbol (usually =, but sometimes ≥ or ≤).
The alternative hypothesis is often abbreviated as H a or H 1 . When the alternative hypothesis is written using mathematical symbols, it always includes an inequality symbol (usually ≠, but sometimes < or >).
A research hypothesis is your proposed answer to your research question. The research hypothesis usually includes an explanation (‘ x affects y because …’).
A statistical hypothesis, on the other hand, is a mathematical statement about a population parameter. Statistical hypotheses always come in pairs: the null and alternative hypotheses. In a well-designed study , the statistical hypotheses correspond logically to the research hypothesis.
Cite this Scribbr article
If you want to cite this source, you can copy and paste the citation or click the ‘Cite this Scribbr article’ button to automatically add the citation to our free Reference Generator.
Turney, S. (2022, December 06). Null and Alternative Hypotheses | Definitions & Examples. Scribbr. Retrieved 9 September 2024, from https://www.scribbr.co.uk/stats/null-and-alternative-hypothesis/
Is this article helpful?

Shaun Turney
Other students also liked, levels of measurement: nominal, ordinal, interval, ratio, the standard normal distribution | calculator, examples & uses, types of variables in research | definitions & examples.
Null Hypothesis Definition and Examples, How to State
What is the null hypothesis, how to state the null hypothesis, null hypothesis overview.

Why is it Called the “Null”?
The word “null” in this context means that it’s a commonly accepted fact that researchers work to nullify . It doesn’t mean that the statement is null (i.e. amounts to nothing) itself! (Perhaps the term should be called the “nullifiable hypothesis” as that might cause less confusion).
Why Do I need to Test it? Why not just prove an alternate one?
The short answer is, as a scientist, you are required to ; It’s part of the scientific process. Science uses a battery of processes to prove or disprove theories, making sure than any new hypothesis has no flaws. Including both a null and an alternate hypothesis is one safeguard to ensure your research isn’t flawed. Not including the null hypothesis in your research is considered very bad practice by the scientific community. If you set out to prove an alternate hypothesis without considering it, you are likely setting yourself up for failure. At a minimum, your experiment will likely not be taken seriously.

- Null hypothesis : H 0 : The world is flat.
- Alternate hypothesis: The world is round.
Several scientists, including Copernicus , set out to disprove the null hypothesis. This eventually led to the rejection of the null and the acceptance of the alternate. Most people accepted it — the ones that didn’t created the Flat Earth Society !. What would have happened if Copernicus had not disproved the it and merely proved the alternate? No one would have listened to him. In order to change people’s thinking, he first had to prove that their thinking was wrong .
How to State the Null Hypothesis from a Word Problem
You’ll be asked to convert a word problem into a hypothesis statement in statistics that will include a null hypothesis and an alternate hypothesis . Breaking your problem into a few small steps makes these problems much easier to handle.
Step 2: Convert the hypothesis to math . Remember that the average is sometimes written as μ.
H 1 : μ > 8.2
Broken down into (somewhat) English, that’s H 1 (The hypothesis): μ (the average) > (is greater than) 8.2
Step 3: State what will happen if the hypothesis doesn’t come true. If the recovery time isn’t greater than 8.2 weeks, there are only two possibilities, that the recovery time is equal to 8.2 weeks or less than 8.2 weeks.
H 0 : μ ≤ 8.2
Broken down again into English, that’s H 0 (The null hypothesis): μ (the average) ≤ (is less than or equal to) 8.2
How to State the Null Hypothesis: Part Two
But what if the researcher doesn’t have any idea what will happen.
Example Problem: A researcher is studying the effects of radical exercise program on knee surgery patients. There is a good chance the therapy will improve recovery time, but there’s also the possibility it will make it worse. Average recovery times for knee surgery patients is 8.2 weeks.
Step 1: State what will happen if the experiment doesn’t make any difference. That’s the null hypothesis–that nothing will happen. In this experiment, if nothing happens, then the recovery time will stay at 8.2 weeks.
H 0 : μ = 8.2
Broken down into English, that’s H 0 (The null hypothesis): μ (the average) = (is equal to) 8.2
Step 2: Figure out the alternate hypothesis . The alternate hypothesis is the opposite of the null hypothesis. In other words, what happens if our experiment makes a difference?
H 1 : μ ≠ 8.2
In English again, that’s H 1 (The alternate hypothesis): μ (the average) ≠ (is not equal to) 8.2
That’s How to State the Null Hypothesis!
Check out our Youtube channel for more stats tips!
Gonick, L. (1993). The Cartoon Guide to Statistics . HarperPerennial. Kotz, S.; et al., eds. (2006), Encyclopedia of Statistical Sciences , Wiley.
What is The Null Hypothesis & When Do You Reject The Null Hypothesis
Julia Simkus
Editor at Simply Psychology
BA (Hons) Psychology, Princeton University
Julia Simkus is a graduate of Princeton University with a Bachelor of Arts in Psychology. She is currently studying for a Master's Degree in Counseling for Mental Health and Wellness in September 2023. Julia's research has been published in peer reviewed journals.
Learn about our Editorial Process
Saul McLeod, PhD
Editor-in-Chief for Simply Psychology
BSc (Hons) Psychology, MRes, PhD, University of Manchester
Saul McLeod, PhD., is a qualified psychology teacher with over 18 years of experience in further and higher education. He has been published in peer-reviewed journals, including the Journal of Clinical Psychology.
Olivia Guy-Evans, MSc
Associate Editor for Simply Psychology
BSc (Hons) Psychology, MSc Psychology of Education
Olivia Guy-Evans is a writer and associate editor for Simply Psychology. She has previously worked in healthcare and educational sectors.
On This Page:
A null hypothesis is a statistical concept suggesting no significant difference or relationship between measured variables. It’s the default assumption unless empirical evidence proves otherwise.
The null hypothesis states no relationship exists between the two variables being studied (i.e., one variable does not affect the other).
The null hypothesis is the statement that a researcher or an investigator wants to disprove.
Testing the null hypothesis can tell you whether your results are due to the effects of manipulating the dependent variable or due to random chance.
How to Write a Null Hypothesis
Null hypotheses (H0) start as research questions that the investigator rephrases as statements indicating no effect or relationship between the independent and dependent variables.
It is a default position that your research aims to challenge or confirm.
For example, if studying the impact of exercise on weight loss, your null hypothesis might be:
There is no significant difference in weight loss between individuals who exercise daily and those who do not.
Examples of Null Hypotheses
| Research Question | Null Hypothesis |
|---|---|
| Do teenagers use cell phones more than adults? | Teenagers and adults use cell phones the same amount. |
| Do tomato plants exhibit a higher rate of growth when planted in compost rather than in soil? | Tomato plants show no difference in growth rates when planted in compost rather than soil. |
| Does daily meditation decrease the incidence of depression? | Daily meditation does not decrease the incidence of depression. |
| Does daily exercise increase test performance? | There is no relationship between daily exercise time and test performance. |
| Does the new vaccine prevent infections? | The vaccine does not affect the infection rate. |
| Does flossing your teeth affect the number of cavities? | Flossing your teeth has no effect on the number of cavities. |
When Do We Reject The Null Hypothesis?
We reject the null hypothesis when the data provide strong enough evidence to conclude that it is likely incorrect. This often occurs when the p-value (probability of observing the data given the null hypothesis is true) is below a predetermined significance level.
If the collected data does not meet the expectation of the null hypothesis, a researcher can conclude that the data lacks sufficient evidence to back up the null hypothesis, and thus the null hypothesis is rejected.
Rejecting the null hypothesis means that a relationship does exist between a set of variables and the effect is statistically significant ( p > 0.05).
If the data collected from the random sample is not statistically significance , then the null hypothesis will be accepted, and the researchers can conclude that there is no relationship between the variables.
You need to perform a statistical test on your data in order to evaluate how consistent it is with the null hypothesis. A p-value is one statistical measurement used to validate a hypothesis against observed data.
Calculating the p-value is a critical part of null-hypothesis significance testing because it quantifies how strongly the sample data contradicts the null hypothesis.
The level of statistical significance is often expressed as a p -value between 0 and 1. The smaller the p-value, the stronger the evidence that you should reject the null hypothesis.
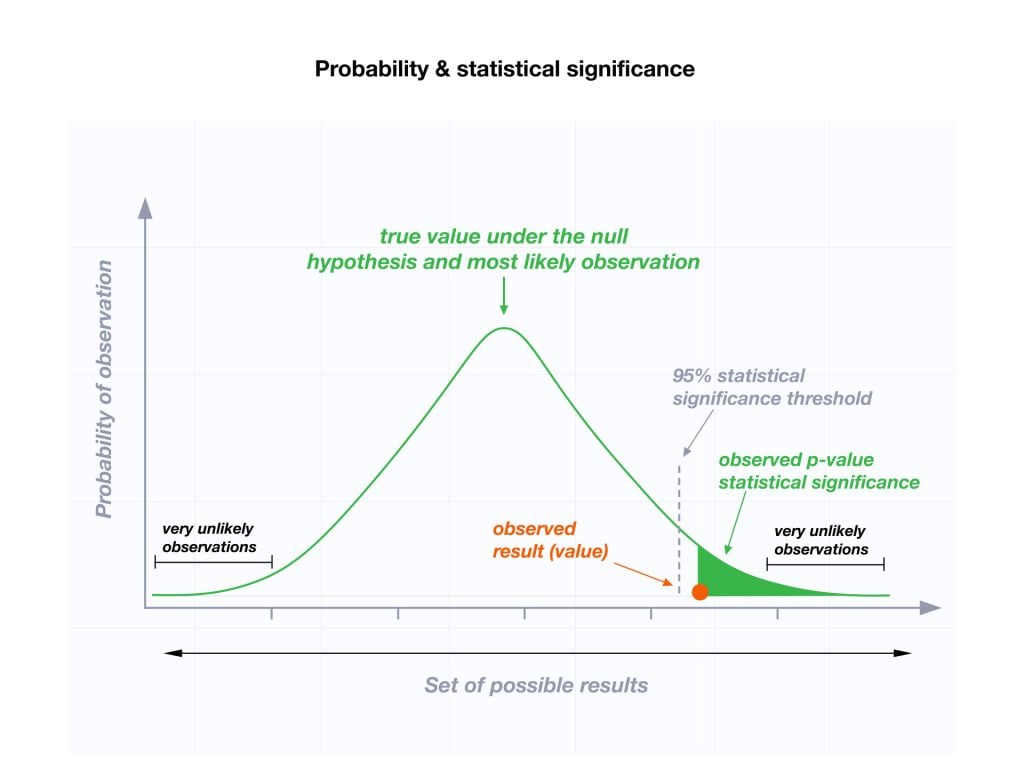
Usually, a researcher uses a confidence level of 95% or 99% (p-value of 0.05 or 0.01) as general guidelines to decide if you should reject or keep the null.
When your p-value is less than or equal to your significance level, you reject the null hypothesis.
In other words, smaller p-values are taken as stronger evidence against the null hypothesis. Conversely, when the p-value is greater than your significance level, you fail to reject the null hypothesis.
In this case, the sample data provides insufficient data to conclude that the effect exists in the population.
Because you can never know with complete certainty whether there is an effect in the population, your inferences about a population will sometimes be incorrect.
When you incorrectly reject the null hypothesis, it’s called a type I error. When you incorrectly fail to reject it, it’s called a type II error.
Why Do We Never Accept The Null Hypothesis?
The reason we do not say “accept the null” is because we are always assuming the null hypothesis is true and then conducting a study to see if there is evidence against it. And, even if we don’t find evidence against it, a null hypothesis is not accepted.
A lack of evidence only means that you haven’t proven that something exists. It does not prove that something doesn’t exist.
It is risky to conclude that the null hypothesis is true merely because we did not find evidence to reject it. It is always possible that researchers elsewhere have disproved the null hypothesis, so we cannot accept it as true, but instead, we state that we failed to reject the null.
One can either reject the null hypothesis, or fail to reject it, but can never accept it.
Why Do We Use The Null Hypothesis?
We can never prove with 100% certainty that a hypothesis is true; We can only collect evidence that supports a theory. However, testing a hypothesis can set the stage for rejecting or accepting this hypothesis within a certain confidence level.
The null hypothesis is useful because it can tell us whether the results of our study are due to random chance or the manipulation of a variable (with a certain level of confidence).
A null hypothesis is rejected if the measured data is significantly unlikely to have occurred and a null hypothesis is accepted if the observed outcome is consistent with the position held by the null hypothesis.
Rejecting the null hypothesis sets the stage for further experimentation to see if a relationship between two variables exists.
Hypothesis testing is a critical part of the scientific method as it helps decide whether the results of a research study support a particular theory about a given population. Hypothesis testing is a systematic way of backing up researchers’ predictions with statistical analysis.
It helps provide sufficient statistical evidence that either favors or rejects a certain hypothesis about the population parameter.
Purpose of a Null Hypothesis
- The primary purpose of the null hypothesis is to disprove an assumption.
- Whether rejected or accepted, the null hypothesis can help further progress a theory in many scientific cases.
- A null hypothesis can be used to ascertain how consistent the outcomes of multiple studies are.
Do you always need both a Null Hypothesis and an Alternative Hypothesis?
The null (H0) and alternative (Ha or H1) hypotheses are two competing claims that describe the effect of the independent variable on the dependent variable. They are mutually exclusive, which means that only one of the two hypotheses can be true.
While the null hypothesis states that there is no effect in the population, an alternative hypothesis states that there is statistical significance between two variables.
The goal of hypothesis testing is to make inferences about a population based on a sample. In order to undertake hypothesis testing, you must express your research hypothesis as a null and alternative hypothesis. Both hypotheses are required to cover every possible outcome of the study.
What is the difference between a null hypothesis and an alternative hypothesis?
The alternative hypothesis is the complement to the null hypothesis. The null hypothesis states that there is no effect or no relationship between variables, while the alternative hypothesis claims that there is an effect or relationship in the population.
It is the claim that you expect or hope will be true. The null hypothesis and the alternative hypothesis are always mutually exclusive, meaning that only one can be true at a time.
What are some problems with the null hypothesis?
One major problem with the null hypothesis is that researchers typically will assume that accepting the null is a failure of the experiment. However, accepting or rejecting any hypothesis is a positive result. Even if the null is not refuted, the researchers will still learn something new.
Why can a null hypothesis not be accepted?
We can either reject or fail to reject a null hypothesis, but never accept it. If your test fails to detect an effect, this is not proof that the effect doesn’t exist. It just means that your sample did not have enough evidence to conclude that it exists.
We can’t accept a null hypothesis because a lack of evidence does not prove something that does not exist. Instead, we fail to reject it.
Failing to reject the null indicates that the sample did not provide sufficient enough evidence to conclude that an effect exists.
If the p-value is greater than the significance level, then you fail to reject the null hypothesis.
Is a null hypothesis directional or non-directional?
A hypothesis test can either contain an alternative directional hypothesis or a non-directional alternative hypothesis. A directional hypothesis is one that contains the less than (“<“) or greater than (“>”) sign.
A nondirectional hypothesis contains the not equal sign (“≠”). However, a null hypothesis is neither directional nor non-directional.
A null hypothesis is a prediction that there will be no change, relationship, or difference between two variables.
The directional hypothesis or nondirectional hypothesis would then be considered alternative hypotheses to the null hypothesis.
Gill, J. (1999). The insignificance of null hypothesis significance testing. Political research quarterly , 52 (3), 647-674.
Krueger, J. (2001). Null hypothesis significance testing: On the survival of a flawed method. American Psychologist , 56 (1), 16.
Masson, M. E. (2011). A tutorial on a practical Bayesian alternative to null-hypothesis significance testing. Behavior research methods , 43 , 679-690.
Nickerson, R. S. (2000). Null hypothesis significance testing: a review of an old and continuing controversy. Psychological methods , 5 (2), 241.
Rozeboom, W. W. (1960). The fallacy of the null-hypothesis significance test. Psychological bulletin , 57 (5), 416.
- PRO Courses Guides New Tech Help Pro Expert Videos About wikiHow Pro Upgrade Sign In
- EDIT Edit this Article
- EXPLORE Tech Help Pro About Us Random Article Quizzes Request a New Article Community Dashboard This Or That Game Happiness Hub Popular Categories Arts and Entertainment Artwork Books Movies Computers and Electronics Computers Phone Skills Technology Hacks Health Men's Health Mental Health Women's Health Relationships Dating Love Relationship Issues Hobbies and Crafts Crafts Drawing Games Education & Communication Communication Skills Personal Development Studying Personal Care and Style Fashion Hair Care Personal Hygiene Youth Personal Care School Stuff Dating All Categories Arts and Entertainment Finance and Business Home and Garden Relationship Quizzes Cars & Other Vehicles Food and Entertaining Personal Care and Style Sports and Fitness Computers and Electronics Health Pets and Animals Travel Education & Communication Hobbies and Crafts Philosophy and Religion Work World Family Life Holidays and Traditions Relationships Youth
- Browse Articles
- Learn Something New
- Quizzes Hot
- Happiness Hub
- This Or That Game
- Train Your Brain
- Explore More
- Support wikiHow
- About wikiHow
- Log in / Sign up
- Education and Communications
- College University and Postgraduate
- Academic Writing
Writing Null Hypotheses in Research and Statistics
Last Updated: September 2, 2024 Fact Checked
This article was co-authored by Joseph Quinones and by wikiHow staff writer, Jennifer Mueller, JD . Joseph Quinones is a Physics Teacher working at South Bronx Community Charter High School. Joseph specializes in astronomy and astrophysics and is interested in science education and science outreach, currently practicing ways to make physics accessible to more students with the goal of bringing more students of color into the STEM fields. He has experience working on Astrophysics research projects at the Museum of Natural History (AMNH). Joseph recieved his Bachelor's degree in Physics from Lehman College and his Masters in Physics Education from City College of New York (CCNY). He is also a member of a network called New York City Men Teach. There are 7 references cited in this article, which can be found at the bottom of the page. This article has been fact-checked, ensuring the accuracy of any cited facts and confirming the authority of its sources. This article has been viewed 30,116 times.
Are you working on a research project and struggling with how to write a null hypothesis? Well, you've come to the right place! Keep reading to learn everything you need to know about the null hypothesis, including a review of what it is, how it relates to your research question and your alternative hypothesis, as well as how to use it in different types of studies.
Things You Should Know
- Write a research null hypothesis as a statement that the studied variables have no relationship to each other, or that there's no difference between 2 groups.

- Adjust the format of your null hypothesis to match the statistical method you used to test it, such as using "mean" if you're comparing the mean between 2 groups.
What is a null hypothesis?

- Research hypothesis: States in plain language that there's no relationship between the 2 variables or there's no difference between the 2 groups being studied.
- Statistical hypothesis: States the predicted outcome of statistical analysis through a mathematical equation related to the statistical method you're using.
Examples of Null Hypotheses

Null Hypothesis vs. Alternative Hypothesis

- For example, your alternative hypothesis could state a positive correlation between 2 variables while your null hypothesis states there's no relationship. If there's a negative correlation, then both hypotheses are false.

- You need additional data or evidence to show that your alternative hypothesis is correct—proving the null hypothesis false is just the first step.
- In smaller studies, sometimes it's enough to show that there's some relationship and your hypothesis could be correct—you can leave the additional proof as an open question for other researchers to tackle.
How do I test a null hypothesis?

- Group means: Compare the mean of the variable in your sample with the mean of the variable in the general population. [6] X Research source
- Group proportions: Compare the proportion of the variable in your sample with the proportion of the variable in the general population. [7] X Research source
- Correlation: Correlation analysis looks at the relationship between 2 variables—specifically, whether they tend to happen together. [8] X Research source
- Regression: Regression analysis reveals the correlation between 2 variables while also controlling for the effect of other, interrelated variables. [9] X Research source
Templates for Null Hypotheses

- Research null hypothesis: There is no difference in the mean [dependent variable] between [group 1] and [group 2].

- Research null hypothesis: The proportion of [dependent variable] in [group 1] and [group 2] is the same.

- Research null hypothesis: There is no correlation between [independent variable] and [dependent variable] in the population.

- Research null hypothesis: There is no relationship between [independent variable] and [dependent variable] in the population.

Expert Q&A

You Might Also Like

Expert Interview

Thanks for reading our article! If you’d like to learn more about physics, check out our in-depth interview with Joseph Quinones .
- ↑ https://online.stat.psu.edu/stat100/lesson/10/10.1
- ↑ https://online.stat.psu.edu/stat501/lesson/2/2.12
- ↑ https://support.minitab.com/en-us/minitab/21/help-and-how-to/statistics/basic-statistics/supporting-topics/basics/null-and-alternative-hypotheses/
- ↑ https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5635437/
- ↑ https://online.stat.psu.edu/statprogram/reviews/statistical-concepts/hypothesis-testing
- ↑ https://education.arcus.chop.edu/null-hypothesis-testing/
- ↑ https://sphweb.bumc.bu.edu/otlt/mph-modules/bs/bs704_hypothesistest-means-proportions/bs704_hypothesistest-means-proportions_print.html
About This Article

- Send fan mail to authors
Reader Success Stories
Dec 3, 2022
Did this article help you?

Featured Articles

Trending Articles

Watch Articles

- Terms of Use
- Privacy Policy
- Do Not Sell or Share My Info
- Not Selling Info
Don’t miss out! Sign up for
wikiHow’s newsletter
Have a language expert improve your writing
Run a free plagiarism check in 10 minutes, generate accurate citations for free.
- Knowledge Base
Hypothesis Testing | A Step-by-Step Guide with Easy Examples
Published on November 8, 2019 by Rebecca Bevans . Revised on June 22, 2023.
Hypothesis testing is a formal procedure for investigating our ideas about the world using statistics . It is most often used by scientists to test specific predictions, called hypotheses, that arise from theories.
There are 5 main steps in hypothesis testing:
- State your research hypothesis as a null hypothesis and alternate hypothesis (H o ) and (H a or H 1 ).
- Collect data in a way designed to test the hypothesis.
- Perform an appropriate statistical test .
- Decide whether to reject or fail to reject your null hypothesis.
- Present the findings in your results and discussion section.
Though the specific details might vary, the procedure you will use when testing a hypothesis will always follow some version of these steps.
Table of contents
Step 1: state your null and alternate hypothesis, step 2: collect data, step 3: perform a statistical test, step 4: decide whether to reject or fail to reject your null hypothesis, step 5: present your findings, other interesting articles, frequently asked questions about hypothesis testing.
After developing your initial research hypothesis (the prediction that you want to investigate), it is important to restate it as a null (H o ) and alternate (H a ) hypothesis so that you can test it mathematically.
The alternate hypothesis is usually your initial hypothesis that predicts a relationship between variables. The null hypothesis is a prediction of no relationship between the variables you are interested in.
- H 0 : Men are, on average, not taller than women. H a : Men are, on average, taller than women.
Prevent plagiarism. Run a free check.
For a statistical test to be valid , it is important to perform sampling and collect data in a way that is designed to test your hypothesis. If your data are not representative, then you cannot make statistical inferences about the population you are interested in.
There are a variety of statistical tests available, but they are all based on the comparison of within-group variance (how spread out the data is within a category) versus between-group variance (how different the categories are from one another).
If the between-group variance is large enough that there is little or no overlap between groups, then your statistical test will reflect that by showing a low p -value . This means it is unlikely that the differences between these groups came about by chance.
Alternatively, if there is high within-group variance and low between-group variance, then your statistical test will reflect that with a high p -value. This means it is likely that any difference you measure between groups is due to chance.
Your choice of statistical test will be based on the type of variables and the level of measurement of your collected data .
- an estimate of the difference in average height between the two groups.
- a p -value showing how likely you are to see this difference if the null hypothesis of no difference is true.
Based on the outcome of your statistical test, you will have to decide whether to reject or fail to reject your null hypothesis.
In most cases you will use the p -value generated by your statistical test to guide your decision. And in most cases, your predetermined level of significance for rejecting the null hypothesis will be 0.05 – that is, when there is a less than 5% chance that you would see these results if the null hypothesis were true.
In some cases, researchers choose a more conservative level of significance, such as 0.01 (1%). This minimizes the risk of incorrectly rejecting the null hypothesis ( Type I error ).
The results of hypothesis testing will be presented in the results and discussion sections of your research paper , dissertation or thesis .
In the results section you should give a brief summary of the data and a summary of the results of your statistical test (for example, the estimated difference between group means and associated p -value). In the discussion , you can discuss whether your initial hypothesis was supported by your results or not.
In the formal language of hypothesis testing, we talk about rejecting or failing to reject the null hypothesis. You will probably be asked to do this in your statistics assignments.
However, when presenting research results in academic papers we rarely talk this way. Instead, we go back to our alternate hypothesis (in this case, the hypothesis that men are on average taller than women) and state whether the result of our test did or did not support the alternate hypothesis.
If your null hypothesis was rejected, this result is interpreted as “supported the alternate hypothesis.”
These are superficial differences; you can see that they mean the same thing.
You might notice that we don’t say that we reject or fail to reject the alternate hypothesis . This is because hypothesis testing is not designed to prove or disprove anything. It is only designed to test whether a pattern we measure could have arisen spuriously, or by chance.
If we reject the null hypothesis based on our research (i.e., we find that it is unlikely that the pattern arose by chance), then we can say our test lends support to our hypothesis . But if the pattern does not pass our decision rule, meaning that it could have arisen by chance, then we say the test is inconsistent with our hypothesis .
If you want to know more about statistics , methodology , or research bias , make sure to check out some of our other articles with explanations and examples.
- Normal distribution
- Descriptive statistics
- Measures of central tendency
- Correlation coefficient
Methodology
- Cluster sampling
- Stratified sampling
- Types of interviews
- Cohort study
- Thematic analysis
Research bias
- Implicit bias
- Cognitive bias
- Survivorship bias
- Availability heuristic
- Nonresponse bias
- Regression to the mean
Hypothesis testing is a formal procedure for investigating our ideas about the world using statistics. It is used by scientists to test specific predictions, called hypotheses , by calculating how likely it is that a pattern or relationship between variables could have arisen by chance.
A hypothesis states your predictions about what your research will find. It is a tentative answer to your research question that has not yet been tested. For some research projects, you might have to write several hypotheses that address different aspects of your research question.
A hypothesis is not just a guess — it should be based on existing theories and knowledge. It also has to be testable, which means you can support or refute it through scientific research methods (such as experiments, observations and statistical analysis of data).
Null and alternative hypotheses are used in statistical hypothesis testing . The null hypothesis of a test always predicts no effect or no relationship between variables, while the alternative hypothesis states your research prediction of an effect or relationship.
Cite this Scribbr article
If you want to cite this source, you can copy and paste the citation or click the “Cite this Scribbr article” button to automatically add the citation to our free Citation Generator.
Bevans, R. (2023, June 22). Hypothesis Testing | A Step-by-Step Guide with Easy Examples. Scribbr. Retrieved September 9, 2024, from https://www.scribbr.com/statistics/hypothesis-testing/
Is this article helpful?
Rebecca Bevans
Other students also liked, choosing the right statistical test | types & examples, understanding p values | definition and examples, what is your plagiarism score.
User Preferences
Content preview.
Arcu felis bibendum ut tristique et egestas quis:
- Ut enim ad minim veniam, quis nostrud exercitation ullamco laboris
- Duis aute irure dolor in reprehenderit in voluptate
- Excepteur sint occaecat cupidatat non proident
Keyboard Shortcuts
10.1 - setting the hypotheses: examples.
A significance test examines whether the null hypothesis provides a plausible explanation of the data. The null hypothesis itself does not involve the data. It is a statement about a parameter (a numerical characteristic of the population). These population values might be proportions or means or differences between means or proportions or correlations or odds ratios or any other numerical summary of the population. The alternative hypothesis is typically the research hypothesis of interest. Here are some examples.
Example 10.2: Hypotheses with One Sample of One Categorical Variable Section
About 10% of the human population is left-handed. Suppose a researcher at Penn State speculates that students in the College of Arts and Architecture are more likely to be left-handed than people found in the general population. We only have one sample since we will be comparing a population proportion based on a sample value to a known population value.
- Research Question : Are artists more likely to be left-handed than people found in the general population?
- Response Variable : Classification of the student as either right-handed or left-handed
State Null and Alternative Hypotheses
- Null Hypothesis : Students in the College of Arts and Architecture are no more likely to be left-handed than people in the general population (population percent of left-handed students in the College of Art and Architecture = 10% or p = .10).
- Alternative Hypothesis : Students in the College of Arts and Architecture are more likely to be left-handed than people in the general population (population percent of left-handed students in the College of Arts and Architecture > 10% or p > .10). This is a one-sided alternative hypothesis.
Example 10.3: Hypotheses with One Sample of One Measurement Variable Section

A generic brand of the anti-histamine Diphenhydramine markets a capsule with a 50 milligram dose. The manufacturer is worried that the machine that fills the capsules has come out of calibration and is no longer creating capsules with the appropriate dosage.
- Research Question : Does the data suggest that the population mean dosage of this brand is different than 50 mg?
- Response Variable : dosage of the active ingredient found by a chemical assay.
- Null Hypothesis : On the average, the dosage sold under this brand is 50 mg (population mean dosage = 50 mg).
- Alternative Hypothesis : On the average, the dosage sold under this brand is not 50 mg (population mean dosage ≠ 50 mg). This is a two-sided alternative hypothesis.
Example 10.4: Hypotheses with Two Samples of One Categorical Variable Section

Many people are starting to prefer vegetarian meals on a regular basis. Specifically, a researcher believes that females are more likely than males to eat vegetarian meals on a regular basis.
- Research Question : Does the data suggest that females are more likely than males to eat vegetarian meals on a regular basis?
- Response Variable : Classification of whether or not a person eats vegetarian meals on a regular basis
- Explanatory (Grouping) Variable: Sex
- Null Hypothesis : There is no sex effect regarding those who eat vegetarian meals on a regular basis (population percent of females who eat vegetarian meals on a regular basis = population percent of males who eat vegetarian meals on a regular basis or p females = p males ).
- Alternative Hypothesis : Females are more likely than males to eat vegetarian meals on a regular basis (population percent of females who eat vegetarian meals on a regular basis > population percent of males who eat vegetarian meals on a regular basis or p females > p males ). This is a one-sided alternative hypothesis.
Example 10.5: Hypotheses with Two Samples of One Measurement Variable Section

Obesity is a major health problem today. Research is starting to show that people may be able to lose more weight on a low carbohydrate diet than on a low fat diet.
- Research Question : Does the data suggest that, on the average, people are able to lose more weight on a low carbohydrate diet than on a low fat diet?
- Response Variable : Weight loss (pounds)
- Explanatory (Grouping) Variable : Type of diet
- Null Hypothesis : There is no difference in the mean amount of weight loss when comparing a low carbohydrate diet with a low fat diet (population mean weight loss on a low carbohydrate diet = population mean weight loss on a low fat diet).
- Alternative Hypothesis : The mean weight loss should be greater for those on a low carbohydrate diet when compared with those on a low fat diet (population mean weight loss on a low carbohydrate diet > population mean weight loss on a low fat diet). This is a one-sided alternative hypothesis.
Example 10.6: Hypotheses about the relationship between Two Categorical Variables Section
- Research Question : Do the odds of having a stroke increase if you inhale second hand smoke ? A case-control study of non-smoking stroke patients and controls of the same age and occupation are asked if someone in their household smokes.
- Variables : There are two different categorical variables (Stroke patient vs control and whether the subject lives in the same household as a smoker). Living with a smoker (or not) is the natural explanatory variable and having a stroke (or not) is the natural response variable in this situation.
- Null Hypothesis : There is no relationship between whether or not a person has a stroke and whether or not a person lives with a smoker (odds ratio between stroke and second-hand smoke situation is = 1).
- Alternative Hypothesis : There is a relationship between whether or not a person has a stroke and whether or not a person lives with a smoker (odds ratio between stroke and second-hand smoke situation is > 1). This is a one-tailed alternative.
This research question might also be addressed like example 11.4 by making the hypotheses about comparing the proportion of stroke patients that live with smokers to the proportion of controls that live with smokers.
Example 10.7: Hypotheses about the relationship between Two Measurement Variables Section
- Research Question : A financial analyst believes there might be a positive association between the change in a stock's price and the amount of the stock purchased by non-management employees the previous day (stock trading by management being under "insider-trading" regulatory restrictions).
- Variables : Daily price change information (the response variable) and previous day stock purchases by non-management employees (explanatory variable). These are two different measurement variables.
- Null Hypothesis : The correlation between the daily stock price change (\$) and the daily stock purchases by non-management employees (\$) = 0.
- Alternative Hypothesis : The correlation between the daily stock price change (\$) and the daily stock purchases by non-management employees (\$) > 0. This is a one-sided alternative hypothesis.
Example 10.8: Hypotheses about comparing the relationship between Two Measurement Variables in Two Samples Section
- Research Question : Is there a linear relationship between the amount of the bill (\$) at a restaurant and the tip (\$) that was left. Is the strength of this association different for family restaurants than for fine dining restaurants?
- Variables : There are two different measurement variables. The size of the tip would depend on the size of the bill so the amount of the bill would be the explanatory variable and the size of the tip would be the response variable.
- Null Hypothesis : The correlation between the amount of the bill (\$) at a restaurant and the tip (\$) that was left is the same at family restaurants as it is at fine dining restaurants.
- Alternative Hypothesis : The correlation between the amount of the bill (\$) at a restaurant and the tip (\$) that was left is the difference at family restaurants then it is at fine dining restaurants. This is a two-sided alternative hypothesis.
Module 9: Hypothesis Testing With One Sample
Null and alternative hypotheses, learning outcomes.
- Describe hypothesis testing in general and in practice
The actual test begins by considering two hypotheses . They are called the null hypothesis and the alternative hypothesis . These hypotheses contain opposing viewpoints.
H 0 : The null hypothesis: It is a statement about the population that either is believed to be true or is used to put forth an argument unless it can be shown to be incorrect beyond a reasonable doubt.
H a : The alternative hypothesis : It is a claim about the population that is contradictory to H 0 and what we conclude when we reject H 0 .
Since the null and alternative hypotheses are contradictory, you must examine evidence to decide if you have enough evidence to reject the null hypothesis or not. The evidence is in the form of sample data.
After you have determined which hypothesis the sample supports, you make adecision. There are two options for a decision . They are “reject H 0 ” if the sample information favors the alternative hypothesis or “do not reject H 0 ” or “decline to reject H 0 ” if the sample information is insufficient to reject the null hypothesis.
Mathematical Symbols Used in H 0 and H a :
| equal (=) | not equal (≠) greater than (>) less than (<) |
| greater than or equal to (≥) | less than (<) |
| less than or equal to (≤) | more than (>) |
H 0 always has a symbol with an equal in it. H a never has a symbol with an equal in it. The choice of symbol depends on the wording of the hypothesis test. However, be aware that many researchers (including one of the co-authors in research work) use = in the null hypothesis, even with > or < as the symbol in the alternative hypothesis. This practice is acceptable because we only make the decision to reject or not reject the null hypothesis.
H 0 : No more than 30% of the registered voters in Santa Clara County voted in the primary election. p ≤ 30
H a : More than 30% of the registered voters in Santa Clara County voted in the primary election. p > 30
A medical trial is conducted to test whether or not a new medicine reduces cholesterol by 25%. State the null and alternative hypotheses.
H 0 : The drug reduces cholesterol by 25%. p = 0.25
H a : The drug does not reduce cholesterol by 25%. p ≠ 0.25
We want to test whether the mean GPA of students in American colleges is different from 2.0 (out of 4.0). The null and alternative hypotheses are:
H 0 : μ = 2.0
H a : μ ≠ 2.0
We want to test whether the mean height of eighth graders is 66 inches. State the null and alternative hypotheses. Fill in the correct symbol (=, ≠, ≥, <, ≤, >) for the null and alternative hypotheses. H 0 : μ __ 66 H a : μ __ 66
- H 0 : μ = 66
- H a : μ ≠ 66
We want to test if college students take less than five years to graduate from college, on the average. The null and alternative hypotheses are:
H 0 : μ ≥ 5
H a : μ < 5
We want to test if it takes fewer than 45 minutes to teach a lesson plan. State the null and alternative hypotheses. Fill in the correct symbol ( =, ≠, ≥, <, ≤, >) for the null and alternative hypotheses. H 0 : μ __ 45 H a : μ __ 45
- H 0 : μ ≥ 45
- H a : μ < 45
In an issue of U.S. News and World Report , an article on school standards stated that about half of all students in France, Germany, and Israel take advanced placement exams and a third pass. The same article stated that 6.6% of U.S. students take advanced placement exams and 4.4% pass. Test if the percentage of U.S. students who take advanced placement exams is more than 6.6%. State the null and alternative hypotheses.
H 0 : p ≤ 0.066
H a : p > 0.066
On a state driver’s test, about 40% pass the test on the first try. We want to test if more than 40% pass on the first try. Fill in the correct symbol (=, ≠, ≥, <, ≤, >) for the null and alternative hypotheses. H 0 : p __ 0.40 H a : p __ 0.40
- H 0 : p = 0.40
- H a : p > 0.40
Concept Review
In a hypothesis test , sample data is evaluated in order to arrive at a decision about some type of claim. If certain conditions about the sample are satisfied, then the claim can be evaluated for a population. In a hypothesis test, we: Evaluate the null hypothesis , typically denoted with H 0 . The null is not rejected unless the hypothesis test shows otherwise. The null statement must always contain some form of equality (=, ≤ or ≥) Always write the alternative hypothesis , typically denoted with H a or H 1 , using less than, greater than, or not equals symbols, i.e., (≠, >, or <). If we reject the null hypothesis, then we can assume there is enough evidence to support the alternative hypothesis. Never state that a claim is proven true or false. Keep in mind the underlying fact that hypothesis testing is based on probability laws; therefore, we can talk only in terms of non-absolute certainties.
Formula Review
H 0 and H a are contradictory.
- OpenStax, Statistics, Null and Alternative Hypotheses. Provided by : OpenStax. Located at : http://cnx.org/contents/[email protected]:58/Introductory_Statistics . License : CC BY: Attribution
- Introductory Statistics . Authored by : Barbara Illowski, Susan Dean. Provided by : Open Stax. Located at : http://cnx.org/contents/[email protected] . License : CC BY: Attribution . License Terms : Download for free at http://cnx.org/contents/[email protected]
- Simple hypothesis testing | Probability and Statistics | Khan Academy. Authored by : Khan Academy. Located at : https://youtu.be/5D1gV37bKXY . License : All Rights Reserved . License Terms : Standard YouTube License
15 Null Hypothesis Examples

Chris Drew (PhD)
Dr. Chris Drew is the founder of the Helpful Professor. He holds a PhD in education and has published over 20 articles in scholarly journals. He is the former editor of the Journal of Learning Development in Higher Education. [Image Descriptor: Photo of Chris]
Learn about our Editorial Process

A null hypothesis is a general assertion or default position that there is no relationship or effect between two measured phenomena.
It’s a critical part of statistics, data analysis, and the scientific method . This concept forms the basis of testing statistical significance and allows researchers to be objective in their conclusions.
A null hypothesis helps to eliminate biases and ensures that the observed results are not due to chance. The rejection or failure to reject the null hypothesis helps in guiding the course of research.

Null Hypothesis Definition
The null hypothesis, often denoted as H 0 , is the hypothesis in a statistical test which proposes no statistical significance exists in a set of observed data.
It hypothesizes that any kind of difference or importance you see in a data set is due to chance.
Null hypotheses are typically proposed to be negated or disproved by statistical tests, paving way for the acceptance of an alternate hypothesis.
Importantly, a null hypothesis cannot be proven true; it can only be supported or rejected with confidence.
Should evidence – via statistical analysis – contradict the null hypothesis, it is rejected in favor of an alternative hypothesis. In essence, the null hypothesis is a tool to challenge and disprove that there is no effect or relationship between variables.
Video Explanation
I like to show this video to my students which outlines a null hypothesis really clearly and engagingly, using real life studies by research students! The into explains it really well:
“There’s an idea in science called the null hypothesis and it works like this: when you’re setting out to prove a theory, your default answer should be “it’s not going to work” and you have to convince the world otherwise through clear results”
Here’s the full video:
Null Hypothesis Examples
- Equality of Means: The null hypothesis posits that the average of group A does not differ from the average of group B. It suggests that any observed difference between the two group means is due to sampling or experimental error.
- No Correlation: The null hypothesis states there is no correlation between the variable X and variable Y in the population. It means that any correlation seen in sample data occurred by chance.
- Drug Effectiveness: The null hypothesis proposes that a new drug does not reduce the number of days to recover from a disease compared to a standard drug. Any observed difference is merely by chance and not due to the new drug.
- Classroom Teaching Method: The null hypothesis states that a new teaching method does not result in improved test scores compared to the traditional teaching method. Any improvement in scores can be attributed to chance or other unrelated factors.
- Smoking and Life Expectancy: The null hypothesis asserts that the average life expectancy of smokers is the same as that of non-smokers. Any perceived difference in life expectancy is due to random variation or other factors.
- Brand Preference: The null hypothesis suggests that the proportion of consumers preferring Brand A is the same as those preferring Brand B. Any observed preference in the sample is due to random selection.
- Vaccination Efficacy: The null hypothesis states that the efficacy of Vaccine A does not differ from that of Vaccine B. Any differences observed in a sample are due to chance or other confounding factors.
- Diet and Weight Loss: The null hypothesis proposes that following a specific diet does not result in more weight loss than not following the diet. Any weight loss observed among dieters is considered random or influenced by other factors.
- Exercise and Heart Rate: The null hypothesis states that regular exercise does not lower resting heart rate compared to no exercise. Any lower heart rates observed in exercisers could be due to chance or other unrelated factors.
- Climate Change: The null hypothesis asserts that the average global temperature this decade is not higher than the previous decade. Any observed temperature increase can be attributed to random variation or unaccounted factors.
- Gender Wage Gap: The null hypothesis posits that men and women earn the same average wage for the same job. Any observed wage disparity is due to chance or non-gender related factors.
- Psychotherapy Effectiveness: The null hypothesis states that patients undergoing psychotherapy do not show more improvement than those not undergoing therapy. Any improvement in the
- Energy Drink Consumption and Sleep: The null hypothesis proposes that consuming energy drinks does not affect the quantity of sleep. Any observed differences in sleep duration among energy drink consumers is due to random variation or other factors.
- Organic Food and Health: The null hypothesis asserts that consuming organic food does not lead to better health outcomes compared to consuming non-organic food. Any health differences observed in consumers of organic food are considered random or attributed to other confounding factors.
- Online Learning Effectiveness: The null hypothesis states that students learning online do not perform differently on exams than students learning in traditional classrooms. Any difference in performance can be attributed to chance or unrelated factors.
Null Hypothesis vs Alternative Hypothesis
An alternative hypothesis is the direct contrast to the null hypothesis. It posits that there is a statistically significant relationship or effect between the variables being observed.
If the null hypothesis is rejected based on the test data, the alternative hypothesis is accepted.
Importantly, while the null hypothesis is typically a statement of ‘no effect’ or ‘no difference,’ the alternative hypothesis states that there is an effect or difference.
Comprehension Checkpoint: How does the null hypothesis help to ensure that research is objective and unbiased?
| A statement of no effect or no relationship | A statement that suggests there is an effect or relationship | |
| H | H or H | |
| The average time to recover using Drug A is the same as with Drug B | The average time to recover using Drug A is less than with Drug B | |
| No statistical significance between observed data | Statistical significance exists between observed data | |
| The observed result is due to chance | The observed result is due to the effect or relationship |
Applications of the Null Hypothesis in Research
The null hypothesis plays a critical role in numerous research settings, promoting objectivity and ensuring findings aren’t due to random chance.
- Clinical Trials: Null hypothesis is used extensively in medical and pharmaceutical research. For example, when testing a new drug’s effectiveness, the null hypothesis might state that the drug has no effect on the disease. If data contradicts this, the null hypothesis is rejected, suggesting the drug might be effective.
- Business and Economics: Businesses use null hypotheses to make informed decisions. For instance, a company might use a null hypothesis to test if a new marketing strategy improves sales. If data suggests a significant increase in sales, the null hypothesis is rejected, and the new strategy may be implemented.
- Psychological Research: Psychologists use null hypotheses to test theories about behavior. For instance, a null hypothesis might state there’s no link between stress and sleep quality. Rejecting this hypothesis based on collected data could help establish a correlation between the two variables.
- Environmental Science: Null hypotheses are used to understand environmental changes. For instance, researchers might form a null hypothesis stating there is no significant difference in air quality before and after a policy change. If this hypothesis is rejected, it indicates the policy may have impacted air quality.
- Education: Educators and researchers often use null hypotheses to improve teaching methods. For example, a null hypothesis might propose a new teaching technique doesn’t enhance student performance. If data contradicts this, the technique may be beneficial.
In all these areas, the null hypothesis helps minimize bias, enabling researchers to support their findings with statistically significant data. It forms the backbone of many scientific research methodologies , promoting a disciplined approach to uncovering new knowledge.
See More Hypothesis Examples Here
The null hypothesis is a cornerstone of statistical analysis and empirical research. It serves as a starting point for investigations, providing a baseline premise that the observed effects are due to chance. By understanding and applying the concept of the null hypothesis, researchers can test the validity of their assumptions, making their findings more robust and reliable. In essence, the null hypothesis ensures that the scientific exploration remains objective, systematic, and free from unintended bias.

- Chris Drew (PhD) https://helpfulprofessor.com/author/chris-drew-phd-2/ 10 Reasons you’re Perpetually Single
- Chris Drew (PhD) https://helpfulprofessor.com/author/chris-drew-phd-2/ 20 Montessori Toddler Bedrooms (Design Inspiration)
- Chris Drew (PhD) https://helpfulprofessor.com/author/chris-drew-phd-2/ 21 Montessori Homeschool Setups
- Chris Drew (PhD) https://helpfulprofessor.com/author/chris-drew-phd-2/ 101 Hidden Talents Examples
Leave a Comment Cancel Reply
Your email address will not be published. Required fields are marked *
Null Hypothesis and Alternative Hypothesis
- Inferential Statistics
- Statistics Tutorials
- Probability & Games
- Descriptive Statistics
- Applications Of Statistics
- Math Tutorials
- Pre Algebra & Algebra
- Exponential Decay
- Worksheets By Grade
- Ph.D., Mathematics, Purdue University
- M.S., Mathematics, Purdue University
- B.A., Mathematics, Physics, and Chemistry, Anderson University
Hypothesis testing involves the careful construction of two statements: the null hypothesis and the alternative hypothesis. These hypotheses can look very similar but are actually different.
How do we know which hypothesis is the null and which one is the alternative? We will see that there are a few ways to tell the difference.
The Null Hypothesis
The null hypothesis reflects that there will be no observed effect in our experiment. In a mathematical formulation of the null hypothesis, there will typically be an equal sign. This hypothesis is denoted by H 0 .
The null hypothesis is what we attempt to find evidence against in our hypothesis test. We hope to obtain a small enough p-value that it is lower than our level of significance alpha and we are justified in rejecting the null hypothesis. If our p-value is greater than alpha, then we fail to reject the null hypothesis.
If the null hypothesis is not rejected, then we must be careful to say what this means. The thinking on this is similar to a legal verdict. Just because a person has been declared "not guilty", it does not mean that he is innocent. In the same way, just because we failed to reject a null hypothesis it does not mean that the statement is true.
For example, we may want to investigate the claim that despite what convention has told us, the mean adult body temperature is not the accepted value of 98.6 degrees Fahrenheit . The null hypothesis for an experiment to investigate this is “The mean adult body temperature for healthy individuals is 98.6 degrees Fahrenheit.” If we fail to reject the null hypothesis, then our working hypothesis remains that the average adult who is healthy has a temperature of 98.6 degrees. We do not prove that this is true.
If we are studying a new treatment, the null hypothesis is that our treatment will not change our subjects in any meaningful way. In other words, the treatment will not produce any effect in our subjects.
The Alternative Hypothesis
The alternative or experimental hypothesis reflects that there will be an observed effect for our experiment. In a mathematical formulation of the alternative hypothesis, there will typically be an inequality, or not equal to symbol. This hypothesis is denoted by either H a or by H 1 .
The alternative hypothesis is what we are attempting to demonstrate in an indirect way by the use of our hypothesis test. If the null hypothesis is rejected, then we accept the alternative hypothesis. If the null hypothesis is not rejected, then we do not accept the alternative hypothesis. Going back to the above example of mean human body temperature, the alternative hypothesis is “The average adult human body temperature is not 98.6 degrees Fahrenheit.”
If we are studying a new treatment, then the alternative hypothesis is that our treatment does, in fact, change our subjects in a meaningful and measurable way.
The following set of negations may help when you are forming your null and alternative hypotheses. Most technical papers rely on just the first formulation, even though you may see some of the others in a statistics textbook.
- Null hypothesis: “ x is equal to y .” Alternative hypothesis “ x is not equal to y .”
- Null hypothesis: “ x is at least y .” Alternative hypothesis “ x is less than y .”
- Null hypothesis: “ x is at most y .” Alternative hypothesis “ x is greater than y .”
- What 'Fail to Reject' Means in a Hypothesis Test
- Type I and Type II Errors in Statistics
- An Example of a Hypothesis Test
- The Runs Test for Random Sequences
- An Example of Chi-Square Test for a Multinomial Experiment
- The Difference Between Type I and Type II Errors in Hypothesis Testing
- What Level of Alpha Determines Statistical Significance?
- What Is the Difference Between Alpha and P-Values?
- What Is ANOVA?
- How to Find Critical Values with a Chi-Square Table
- Example of a Permutation Test
- Degrees of Freedom for Independence of Variables in Two-Way Table
- Example of an ANOVA Calculation
- How to Find Degrees of Freedom in Statistics
- How to Construct a Confidence Interval for a Population Proportion
- Degrees of Freedom in Statistics and Mathematics
- School Guide
- Mathematics
- Number System and Arithmetic
- Trigonometry
- Probability
- Mensuration
- Maths Formulas
- Integration Formulas
- Differentiation Formulas
- Trigonometry Formulas
- Algebra Formulas
- Mensuration Formula
- Statistics Formulas
- Trigonometric Table
Null Hypothesis
Null Hypothesis , often denoted as H 0, is a foundational concept in statistical hypothesis testing. It represents an assumption that no significant difference, effect, or relationship exists between variables within a population. It serves as a baseline assumption, positing no observed change or effect occurring. The null is t he truth or falsity of an idea in analysis.
In this article, we will discuss the null hypothesis in detail, along with some solved examples and questions on the null hypothesis.
Table of Content
What is Null Hypothesis?
Null hypothesis symbol, formula of null hypothesis, types of null hypothesis, null hypothesis examples, principle of null hypothesis, how do you find null hypothesis, null hypothesis in statistics, null hypothesis and alternative hypothesis, null hypothesis and alternative hypothesis examples, null hypothesis – practice problems.
Null Hypothesis in statistical analysis suggests the absence of statistical significance within a specific set of observed data. Hypothesis testing, using sample data, evaluates the validity of this hypothesis. Commonly denoted as H 0 or simply “null,” it plays an important role in quantitative analysis, examining theories related to markets, investment strategies, or economies to determine their validity.
Null Hypothesis Meaning
Null Hypothesis represents a default position, often suggesting no effect or difference, against which researchers compare their experimental results. The Null Hypothesis, often denoted as H 0 asserts a default assumption in statistical analysis. It posits no significant difference or effect, serving as a baseline for comparison in hypothesis testing.
The null Hypothesis is represented as H 0 , the Null Hypothesis symbolizes the absence of a measurable effect or difference in the variables under examination.
Certainly, a simple example would be asserting that the mean score of a group is equal to a specified value like stating that the average IQ of a population is 100.
The Null Hypothesis is typically formulated as a statement of equality or absence of a specific parameter in the population being studied. It provides a clear and testable prediction for comparison with the alternative hypothesis. The formulation of the Null Hypothesis typically follows a concise structure, stating the equality or absence of a specific parameter in the population.
Mean Comparison (Two-sample t-test)
H 0 : μ 1 = μ 2
This asserts that there is no significant difference between the means of two populations or groups.

Proportion Comparison
H 0 : p 1 − p 2 = 0
This suggests no significant difference in proportions between two populations or conditions.
Equality in Variance (F-test in ANOVA)
H 0 : σ 1 = σ 2
This states that there’s no significant difference in variances between groups or populations.
Independence (Chi-square Test of Independence):
H 0 : Variables are independent
This asserts that there’s no association or relationship between categorical variables.
Null Hypotheses vary including simple and composite forms, each tailored to the complexity of the research question. Understanding these types is pivotal for effective hypothesis testing.
Equality Null Hypothesis (Simple Null Hypothesis)
The Equality Null Hypothesis, also known as the Simple Null Hypothesis, is a fundamental concept in statistical hypothesis testing that assumes no difference, effect or relationship between groups, conditions or populations being compared.
Non-Inferiority Null Hypothesis
In some studies, the focus might be on demonstrating that a new treatment or method is not significantly worse than the standard or existing one.
Superiority Null Hypothesis
The concept of a superiority null hypothesis comes into play when a study aims to demonstrate that a new treatment, method, or intervention is significantly better than an existing or standard one.
Independence Null Hypothesis
In certain statistical tests, such as chi-square tests for independence, the null hypothesis assumes no association or independence between categorical variables.
Homogeneity Null Hypothesis
In tests like ANOVA (Analysis of Variance), the null hypothesis suggests that there’s no difference in population means across different groups.
- Medicine: Null Hypothesis: “No significant difference exists in blood pressure levels between patients given the experimental drug versus those given a placebo.”
- Education: Null Hypothesis: “There’s no significant variation in test scores between students using a new teaching method and those using traditional teaching.”
- Economics: Null Hypothesis: “There’s no significant change in consumer spending pre- and post-implementation of a new taxation policy.”
- Environmental Science: Null Hypothesis: “There’s no substantial difference in pollution levels before and after a water treatment plant’s establishment.”
The principle of the null hypothesis is a fundamental concept in statistical hypothesis testing. It involves making an assumption about the population parameter or the absence of an effect or relationship between variables.
In essence, the null hypothesis (H 0 ) proposes that there is no significant difference, effect, or relationship between variables. It serves as a starting point or a default assumption that there is no real change, no effect or no difference between groups or conditions.
The null hypothesis is usually formulated to be tested against an alternative hypothesis (H 1 or H [Tex]\alpha [/Tex] ) which suggests that there is an effect, difference or relationship present in the population.
Null Hypothesis Rejection
Rejecting the Null Hypothesis occurs when statistical evidence suggests a significant departure from the assumed baseline. It implies that there is enough evidence to support the alternative hypothesis, indicating a meaningful effect or difference. Null Hypothesis rejection occurs when statistical evidence suggests a deviation from the assumed baseline, prompting a reconsideration of the initial hypothesis.
Identifying the Null Hypothesis involves defining the status quotient, asserting no effect and formulating a statement suitable for statistical analysis.
When is Null Hypothesis Rejected?
The Null Hypothesis is rejected when statistical tests indicate a significant departure from the expected outcome, leading to the consideration of alternative hypotheses. It occurs when statistical evidence suggests a deviation from the assumed baseline, prompting a reconsideration of the initial hypothesis.
In statistical hypothesis testing, researchers begin by stating the null hypothesis, often based on theoretical considerations or previous research. The null hypothesis is then tested against an alternative hypothesis (Ha), which represents the researcher’s claim or the hypothesis they seek to support.
The process of hypothesis testing involves collecting sample data and using statistical methods to assess the likelihood of observing the data if the null hypothesis were true. This assessment is typically done by calculating a test statistic, which measures the difference between the observed data and what would be expected under the null hypothesis.
In the realm of hypothesis testing, the null hypothesis (H 0 ) and alternative hypothesis (H₁ or Ha) play critical roles. The null hypothesis generally assumes no difference, effect, or relationship between variables, suggesting that any observed change or effect is due to random chance. Its counterpart, the alternative hypothesis, asserts the presence of a significant difference, effect, or relationship between variables, challenging the null hypothesis. These hypotheses are formulated based on the research question and guide statistical analyses.
Difference Between Null Hypothesis and Alternative Hypothesis
The null hypothesis (H 0 ) serves as the baseline assumption in statistical testing, suggesting no significant effect, relationship, or difference within the data. It often proposes that any observed change or correlation is merely due to chance or random variation. Conversely, the alternative hypothesis (H 1 or Ha) contradicts the null hypothesis, positing the existence of a genuine effect, relationship or difference in the data. It represents the researcher’s intended focus, seeking to provide evidence against the null hypothesis and support for a specific outcome or theory. These hypotheses form the crux of hypothesis testing, guiding the assessment of data to draw conclusions about the population being studied.
Criteria | Null Hypothesis | Alternative Hypothesis |
|---|---|---|
Definition | Assumes no effect or difference | Asserts a specific effect or difference |
Symbol | H | H (or Ha) |
Formulation | States equality or absence of parameter | States a specific value or relationship |
Testing Outcome | Rejected if evidence of a significant effect | Accepted if evidence supports the hypothesis |
Let’s envision a scenario where a researcher aims to examine the impact of a new medication on reducing blood pressure among patients. In this context:
Null Hypothesis (H 0 ): “The new medication does not produce a significant effect in reducing blood pressure levels among patients.”
Alternative Hypothesis (H 1 or Ha): “The new medication yields a significant effect in reducing blood pressure levels among patients.”
The null hypothesis implies that any observed alterations in blood pressure subsequent to the medication’s administration are a result of random fluctuations rather than a consequence of the medication itself. Conversely, the alternative hypothesis contends that the medication does indeed generate a meaningful alteration in blood pressure levels, distinct from what might naturally occur or by random chance.
People Also Read:
Mathematics Maths Formulas Probability and Statistics
Example 1: A researcher claims that the average time students spend on homework is 2 hours per night.
Null Hypothesis (H 0 ): The average time students spend on homework is equal to 2 hours per night. Data: A random sample of 30 students has an average homework time of 1.8 hours with a standard deviation of 0.5 hours. Test Statistic and Decision: Using a t-test, if the calculated t-statistic falls within the acceptance region, we fail to reject the null hypothesis. If it falls in the rejection region, we reject the null hypothesis. Conclusion: Based on the statistical analysis, we fail to reject the null hypothesis, suggesting that there is not enough evidence to dispute the claim of the average homework time being 2 hours per night.
Example 2: A company asserts that the error rate in its production process is less than 1%.
Null Hypothesis (H 0 ): The error rate in the production process is 1% or higher. Data: A sample of 500 products shows an error rate of 0.8%. Test Statistic and Decision: Using a z-test, if the calculated z-statistic falls within the acceptance region, we fail to reject the null hypothesis. If it falls in the rejection region, we reject the null hypothesis. Conclusion: The statistical analysis supports rejecting the null hypothesis, indicating that there is enough evidence to dispute the company’s claim of an error rate of 1% or higher.
Q1. A researcher claims that the average time spent by students on homework is less than 2 hours per day. Formulate the null hypothesis for this claim?
Q2. A manufacturing company states that their new machine produces widgets with a defect rate of less than 5%. Write the null hypothesis to test this claim?
Q3. An educational institute believes that their online course completion rate is at least 60%. Develop the null hypothesis to validate this assertion?
Q4. A restaurant claims that the waiting time for customers during peak hours is not more than 15 minutes. Formulate the null hypothesis for this claim?
Q5. A study suggests that the mean weight loss after following a specific diet plan for a month is more than 8 pounds. Construct the null hypothesis to evaluate this statement?
Summary – Null Hypothesis and Alternative Hypothesis
The null hypothesis (H 0 ) and alternative hypothesis (H a ) are fundamental concepts in statistical hypothesis testing. The null hypothesis represents the default assumption, stating that there is no significant effect, difference, or relationship between variables. It serves as the baseline against which the alternative hypothesis is tested. In contrast, the alternative hypothesis represents the researcher’s hypothesis or the claim to be tested, suggesting that there is a significant effect, difference, or relationship between variables. The relationship between the null and alternative hypotheses is such that they are complementary, and statistical tests are conducted to determine whether the evidence from the data is strong enough to reject the null hypothesis in favor of the alternative hypothesis. This decision is based on the strength of the evidence and the chosen level of significance. Ultimately, the choice between the null and alternative hypotheses depends on the specific research question and the direction of the effect being investigated.
FAQs on Null Hypothesis
What does null hypothesis stands for.
The null hypothesis, denoted as H 0 , is a fundamental concept in statistics used for hypothesis testing. It represents the statement that there is no effect or no difference, and it is the hypothesis that the researcher typically aims to provide evidence against.
How to Form a Null Hypothesis?
A null hypothesis is formed based on the assumption that there is no significant difference or effect between the groups being compared or no association between variables being tested. It often involves stating that there is no relationship, no change, or no effect in the population being studied.
When Do we reject the Null Hypothesis?
In statistical hypothesis testing, if the p-value (the probability of obtaining the observed results) is lower than the chosen significance level (commonly 0.05), we reject the null hypothesis. This suggests that the data provides enough evidence to refute the assumption made in the null hypothesis.
What is a Null Hypothesis in Research?
In research, the null hypothesis represents the default assumption or position that there is no significant difference or effect. Researchers often try to test this hypothesis by collecting data and performing statistical analyses to see if the observed results contradict the assumption.
What Are Alternative and Null Hypotheses?
The null hypothesis (H0) is the default assumption that there is no significant difference or effect. The alternative hypothesis (H1 or Ha) is the opposite, suggesting there is a significant difference, effect or relationship.
What Does it Mean to Reject the Null Hypothesis?
Rejecting the null hypothesis implies that there is enough evidence in the data to support the alternative hypothesis. In simpler terms, it suggests that there might be a significant difference, effect or relationship between the groups or variables being studied.
How to Find Null Hypothesis?
Formulating a null hypothesis often involves considering the research question and assuming that no difference or effect exists. It should be a statement that can be tested through data collection and statistical analysis, typically stating no relationship or no change between variables or groups.
How is Null Hypothesis denoted?
The null hypothesis is commonly symbolized as H 0 in statistical notation.
What is the Purpose of the Null hypothesis in Statistical Analysis?
The null hypothesis serves as a starting point for hypothesis testing, enabling researchers to assess if there’s enough evidence to reject it in favor of an alternative hypothesis.
What happens if we Reject the Null hypothesis?
Rejecting the null hypothesis implies that there is sufficient evidence to support an alternative hypothesis, suggesting a significant effect or relationship between variables.
What are Test for Null Hypothesis?
Various statistical tests, such as t-tests or chi-square tests, are employed to evaluate the validity of the Null Hypothesis in different scenarios.
Please Login to comment...
Similar reads.
- Geeks Premier League
- School Learning
- Geeks Premier League 2023
- Math-Concepts
- 105 Funny Things to Do to Make Someone Laugh
- Best PS5 SSDs in 2024: Top Picks for Expanding Your Storage
- Best Nintendo Switch Controllers in 2024
- Xbox Game Pass Ultimate: Features, Benefits, and Pricing in 2024
- #geekstreak2024 – 21 Days POTD Challenge Powered By Deutsche Bank
Improve your Coding Skills with Practice
What kind of Experience do you want to share?
9.1 Null and Alternative Hypotheses
The actual test begins by considering two hypotheses . They are called the null hypothesis and the alternative hypothesis . These hypotheses contain opposing viewpoints.
H 0 : The null hypothesis: It is a statement of no difference between the variables—they are not related. This can often be considered the status quo and as a result if you cannot accept the null it requires some action.
H a : The alternative hypothesis: It is a claim about the population that is contradictory to H 0 and what we conclude when we reject H 0 . This is usually what the researcher is trying to prove.
Since the null and alternative hypotheses are contradictory, you must examine evidence to decide if you have enough evidence to reject the null hypothesis or not. The evidence is in the form of sample data.
After you have determined which hypothesis the sample supports, you make a decision. There are two options for a decision. They are "reject H 0 " if the sample information favors the alternative hypothesis or "do not reject H 0 " or "decline to reject H 0 " if the sample information is insufficient to reject the null hypothesis.
Mathematical Symbols Used in H 0 and H a :
| equal (=) | not equal (≠) greater than (>) less than (<) |
| greater than or equal to (≥) | less than (<) |
| less than or equal to (≤) | more than (>) |
H 0 always has a symbol with an equal in it. H a never has a symbol with an equal in it. The choice of symbol depends on the wording of the hypothesis test. However, be aware that many researchers (including one of the co-authors in research work) use = in the null hypothesis, even with > or < as the symbol in the alternative hypothesis. This practice is acceptable because we only make the decision to reject or not reject the null hypothesis.
Example 9.1
H 0 : No more than 30% of the registered voters in Santa Clara County voted in the primary election. p ≤ .30 H a : More than 30% of the registered voters in Santa Clara County voted in the primary election. p > 30
A medical trial is conducted to test whether or not a new medicine reduces cholesterol by 25%. State the null and alternative hypotheses.
Example 9.2
We want to test whether the mean GPA of students in American colleges is different from 2.0 (out of 4.0). The null and alternative hypotheses are: H 0 : μ = 2.0 H a : μ ≠ 2.0
We want to test whether the mean height of eighth graders is 66 inches. State the null and alternative hypotheses. Fill in the correct symbol (=, ≠, ≥, <, ≤, >) for the null and alternative hypotheses.
- H 0 : μ __ 66
- H a : μ __ 66
Example 9.3
We want to test if college students take less than five years to graduate from college, on the average. The null and alternative hypotheses are: H 0 : μ ≥ 5 H a : μ < 5
We want to test if it takes fewer than 45 minutes to teach a lesson plan. State the null and alternative hypotheses. Fill in the correct symbol ( =, ≠, ≥, <, ≤, >) for the null and alternative hypotheses.
- H 0 : μ __ 45
- H a : μ __ 45
Example 9.4
In an issue of U. S. News and World Report , an article on school standards stated that about half of all students in France, Germany, and Israel take advanced placement exams and a third pass. The same article stated that 6.6% of U.S. students take advanced placement exams and 4.4% pass. Test if the percentage of U.S. students who take advanced placement exams is more than 6.6%. State the null and alternative hypotheses. H 0 : p ≤ 0.066 H a : p > 0.066
On a state driver’s test, about 40% pass the test on the first try. We want to test if more than 40% pass on the first try. Fill in the correct symbol (=, ≠, ≥, <, ≤, >) for the null and alternative hypotheses.
- H 0 : p __ 0.40
- H a : p __ 0.40
Collaborative Exercise
Bring to class a newspaper, some news magazines, and some Internet articles . In groups, find articles from which your group can write null and alternative hypotheses. Discuss your hypotheses with the rest of the class.
This book may not be used in the training of large language models or otherwise be ingested into large language models or generative AI offerings without OpenStax's permission.
Want to cite, share, or modify this book? This book uses the Creative Commons Attribution License and you must attribute OpenStax.
Access for free at https://openstax.org/books/introductory-statistics-2e/pages/1-introduction
- Authors: Barbara Illowsky, Susan Dean
- Publisher/website: OpenStax
- Book title: Introductory Statistics 2e
- Publication date: Dec 13, 2023
- Location: Houston, Texas
- Book URL: https://openstax.org/books/introductory-statistics-2e/pages/1-introduction
- Section URL: https://openstax.org/books/introductory-statistics-2e/pages/9-1-null-and-alternative-hypotheses
© Jul 18, 2024 OpenStax. Textbook content produced by OpenStax is licensed under a Creative Commons Attribution License . The OpenStax name, OpenStax logo, OpenStax book covers, OpenStax CNX name, and OpenStax CNX logo are not subject to the Creative Commons license and may not be reproduced without the prior and express written consent of Rice University.
Thank you for visiting nature.com. You are using a browser version with limited support for CSS. To obtain the best experience, we recommend you use a more up to date browser (or turn off compatibility mode in Internet Explorer). In the meantime, to ensure continued support, we are displaying the site without styles and JavaScript.
- View all journals
- Explore content
- About the journal
- Publish with us
- Sign up for alerts
- Open access
- Published: 11 September 2024
Brain-wide dynamics linking sensation to action during decision-making
- Andrei Khilkevich 1 na1 ,
- Michael Lohse ORCID: orcid.org/0000-0001-8864-0704 1 na1 ,
- Ryan Low 1 ,
- Ivana Orsolic 1 ,
- Tadej Bozic 1 ,
- Paige Windmill ORCID: orcid.org/0009-0005-6410-3564 1 &
- Thomas D. Mrsic-Flogel ORCID: orcid.org/0000-0002-8947-408X 1
Nature ( 2024 ) Cite this article
Metrics details
- Sensory processing
- Short-term memory
Perceptual decisions rely on learned associations between sensory evidence and appropriate actions, involving the filtering and integration of relevant inputs to prepare and execute timely responses 1 , 2 . Despite the distributed nature of task-relevant representations 3 , 4 , 5 , 6 , 7 , 8 , 9 , 10 , it remains unclear how transformations between sensory input, evidence integration, motor planning and execution are orchestrated across brain areas and dimensions of neural activity. Here we addressed this question by recording brain-wide neural activity in mice learning to report changes in ambiguous visual input. After learning, evidence integration emerged across most brain areas in sparse neural populations that drive movement-preparatory activity. Visual responses evolved from transient activations in sensory areas to sustained representations in frontal-motor cortex, thalamus, basal ganglia, midbrain and cerebellum, enabling parallel evidence accumulation. In areas that accumulate evidence, shared population activity patterns encode visual evidence and movement preparation, distinct from movement-execution dynamics. Activity in movement-preparatory subspace is driven by neurons integrating evidence, which collapses at movement onset, allowing the integration process to reset. Across premotor regions, evidence-integration timescales were independent of intrinsic regional dynamics, and thus depended on task experience. In summary, learning aligns evidence accumulation to action preparation in activity dynamics across dozens of brain regions. This leads to highly distributed and parallelized sensorimotor transformations during decision-making. Our work unifies concepts from decision-making and motor control fields into a brain-wide framework for understanding how sensory evidence controls actions.
To link external events to beneficial actions, the brain must learn to transform relevant sensory input to drive the neural dynamics that underlie movement preparation and execution 1 , 11 . Where and how these transformations occur in the brain remain unclear.
When individuals make decisions based on ambiguous sensory information over time, the brain is thought to gradually accumulate the relevant input into an integrated neural representation that determines the upcoming choice 1 . Neural activity reflecting the integration of sensory evidence has been reported in several brain areas 1 , 8 , 12 , 13 , 14 , 15 , 16 , 17 , 18 , 19 , 20 , 21 , 22 , most prominently in cortical areas such as frontal-premotor cortex 8 , 13 , 14 , 22 and posterior parietal cortex 15 , 16 , 17 , 18 , and their immediate downstream targets such as the striatum 19 , 20 , 21 . However, recent studies have uncovered a broader encoding of sensory inputs, choice and actions throughout the brains of trained animals 3 , 5 , 6 , 9 , raising questions about where sensory input is transformed into integrated task-relevant representations that guide action, and how widely distributed these representations are. It also remains unclear whether specific brain areas specialize in integration of sensory evidence owing to their inherent properties 8 , 23 , 24 , 25 , 26 , or whether learning shapes the nature of this computation.
Here we address how integrated sensory evidence is converted to a choice and ultimately action. Action initiation is preceded by a build-up of preparatory activity that is observed in many brain areas 4 , 14 , 27 , 28 , 29 , 30 , 31 , 32 (also referred as choice-related activity), which in motor and premotor regions appears distinct from and orthogonal to the pattern of population activity that drives movement execution 4 , 33 , 34 , 35 (see ref. 36 for debate). Although evidence integration has been reported to modulate the preparatory activity of individual neurons in certain brain regions 14 , 18 , 37 , 38 , 39 , 40 , 41 , the effect of evidence integration on the evolving neural dynamics surrounding movement 33 , 34 , as well as the brain regions involved 42 , 43 , 44 , remain to be understood on a brain-wide scale. In particular, it is unclear how segregated or parallelized the transformations between evidence integration, movement preparation and execution are across brain areas as well as across dimensions of neural activity.
To understand the brain-wide transformation of sensory input into choice and action, it is necessary to use tasks that can distinguish sensory and decision-related processes from action signals that dominate global brain activity 3 , 5 , 6 , 9 . Such tasks, pioneered in non-human primates 1 , 16 , 45 , 46 , 47 , have recently been adapted for rodents 3 , 14 , 48 , 49 , 50 , enabling greater access to interrogate the underlying circuit mechanisms as well as unbiased, brain-wide measurements with dense electrode recordings 3 , 5 , 51 .
In this study, we describe how sensory evidence propagates and is transformed across the brain as mice engage in a task that requires temporal integration of visual input, designed to separate the influence of sensory evidence and movement on neural responses 14 . Our results reveal that ambiguous sensory input becomes integrated within widely distributed multi-regional premotor circuits in a learning-dependent manner, driving the preparatory phase of movement-related neural dynamics that eventually trigger the initiation of appropriate actions.
To study how relevant sensory input is transformed across the brain prior to a decision, we trained food-restricted, head-fixed mice on a visual change detection task designed to dissociate ongoing visual evidence observation from movement-related activity 14 . Mice were trained to be stationary on a running wheel while observing a drifting grating stimulus, whose speed fluctuated noisily every 50 ms around a geometric mean temporal frequency (TF) of 1 Hz ( σ = 0.25 octaves), and to report a sustained increase in its speed by licking a reward spout (Fig. 1a ). The mice were motivated to react promptly upon detecting a change by limiting the time in which the reward was accessible ( Methods ). Since changes in speed were often ambiguous, their timing unpredictable and the change in magnitude was randomized, mice had to continuously track the sensory stimulus for a prolonged duration (3–15.5 s) prior to the change. To ensure mice remained still during this time, any licking or movement on the running wheel prior to the stimulus change caused the trial to be aborted ( Methods ).

a , Schematic of the visual change detection task for head-fixed mice. b , Psychometric and reaction-time curves (mean and 95% confidence interval; two-sided Student’s t -test; n = 114 sessions, 15 mice). c , Mean stimulus TF (with 95% confidence interval) preceding early licks during the baseline period. Dashed lines indicate linear mean (1.016 Hz) of baseline stimulus TF. d , Number of units recorded per recording session. e , Brain map of number of units recorded per area across all recording sessions of trained mice. f , Example time series across two trials (a rewarded trial and an early lick trial) of stimulus TF, spike times across simultaneously recorded neurons (two probes), face motion energy (from videography), pupil size and running wheel movement. HPC, hippocampus; TH, thalamus. g , Schematic of single-trial Poisson GLM. Prep., preparation. h , Mean firing rate around early licks (left), and mean response to fast and slow TF pulses during baseline period (right) for an example neuron in MOs and trigeminal motor nucleus (V), together with GLM predicted (on 10% held-out data) mean activity (dashed lines, with 95% confidence interval). Exec., execution; PSTH, peristimulus time histogram. i , Mean (with 95% confidence interval) face motion energy (from videography ( Methods )) around early licks, and around fast and slow TF pulses. j , Brain maps with labelled brain regions. See Supplementary Table 2 for definitions of abbreviations. k , Brain maps of percentage of units encoding lick execution (top row), lick preparation (middle row) and stimulus TF fluctuations during the baseline period in the absence of movement (bottom row). l , Percentage of units encoding lick execution, lick preparation and stimulus TF fluctuations during baseline across all brain regions with more than 40 units recorded. Resp., response. See Supplementary Table 1 for number of units recorded in each brain area and Supplementary Table 2 for definitions of brain region abbreviations. * P < 0.05, ** P < 0.01, *** P < 0.001.
Source Data
The detection performance of the mice improved with the size of the change in stimulus TF (Fig. 1b ). At the same time, their reaction times were hundreds of milliseconds faster for large stimulus changes (Fig. 1b ), similar to other reaction-time tasks requiring temporal integration 1 . Furthermore, the average stimulus speed preceding ‘early licks’ (Fig. 1c ), which occasionally occur during the baseline stimulus prior to change, was increased during approximately 0.3 to 1 s before early lick (Fig. 1c ). This suggests that at least some early licks are triggered by fluctuations in the baseline stimulus and that sensory information influences the mouse’s judgments on the timescale of hundreds of milliseconds.
Thus, by encouraging mice to continuously monitor ambiguous sensory evidence while controlling for their movement, this task enables us to examine how the brain processes sensory evidence and transforms it into action commands.
Brain-wide encoding of sensory input
To understand how the brain of trained mice transforms visual stimulus speed into goal-directed licking in this task, we performed dense silicon electrode recordings (Neuropixels probes 51 ) from 15,406 units spanning 51 brain regions (that is, 12,772 units from regions with more than 40 manually curated, good and stable units; Extended Data Fig. 1 , Supplementary Table 1 and Methods ) distributed across the cortex, basal ganglia, hippocampus, thalamus, midbrain, cerebellum and hindbrain (Fig. 1d–f , 15 mice, 114 recording sessions, 167 probe insertions and 50,997 trials), while capturing high-speed videos of the face and pupil as well as movements of the running wheel (Fig. 1f ).
To identify which neurons encode visual evidence (stimulus TF), lick preparation and lick execution, we utilized single-cell Poisson generalized linear models (GLMs) that fit trial-to-trial neural activity from task-related events, stimuli and behaviour (Fig. 1g and Extended Data Fig. 2 ). By using a cross-validated nested test (that is, holding out a predictor of interest to assess its contribution to neural activity), we identified the neurons that significantly encode different variables of interest while accounting for variance captured by other predictors ( Methods ).
In agreement with the prevalence of motor-related signals in the brain 3 , 4 , 5 , 6 , 9 , lick execution was encoded globally with the activity of at least 50% of neurons recorded encoding this action (Fig. 1k,l and Extended Data Fig. 3a ). Using videography to establish the onset of lick execution, we also identified a smaller, yet substantial fraction of neurons encoding lick preparatory activity (that is, modulation of activity within 1.25 s leading up to a lick), also distributed globally (Fig. 1h,k,l ). A sparser fraction of neurons encoded subtle fluctuations in stimulus TF during the baseline period on trials devoid of mouse movements (5–45%; referred to as TF-responsive units; Methods ). These neurons were distributed across the majority of brain areas. Although the largest contingent of TF-responsive units were found in the visual system (visual cortex, visual thalamus and superficial superior colliculus), significant fractions (5–25%) were also observed in most areas outside the visual system, including regions of the frontal cortex (secondary motor cortex (MOs), anterior cingulate cortex (ACA), medial prefrontal cortex (mPFC), frontal pole (FRP), orbitofrontal cortex (ORB) and primary motor cortex (MOp)), basal ganglia (striatum (caudoputamen; CP), globus pallidus external segment (GPe) and sibstantia nigra reticular part (SNr)), hippocampus (dentate gyrus (DG), CA1, CA3 and subiculum (SUB)), midbrain (midbrain reticular nucleus (MRN), anterior pretectal nucleus (APN), multimodal and motor superior colliculus (SCm) and nucleus of the posterior commisure (NPC)) and cerebellum (lobules 4/5 (Lob4/5), simplex lobule (SIM), central lobule 3 (CENT3), CRUS1/2 and deep cerebellar nuclei (DCN)). Of note, these multi-regional responses to visual input could not be explained by other variables that might correlate with fluctuations in stimulus TF because fast or slow TF pulses did not trigger consistent movements of the face or running wheel (Fig. 1i ), there was an absence of TF-responsive cells in the medulla and orofacial motor/premotor nuclei whose activity reflects movements of the mouth and tongue (Fig. 1h,k,l ), and the GLM was unable to predict responses to TF fluctuations without the stimulus TF as predictor (Extended Data Fig. 2f,g ).
Together, these results show that sensory evidence representations are surprisingly widespread, with a sparse subpopulation of neurons tracking behaviourally subthreshold fluctuations of relevant sensory input in almost all brain areas, but excluding the nuclei controlling orofacial movements which become engaged when mice report their decision. These sparse, distributed representations of visual evidence ultimately give rise to the initiation of movement which itself recruits activity in more than half of neurons across the brain.
Timescales of sensory responses across the brain
To determine how sensory evidence propagates in activity across the brain, we quantified neural responses to momentary samples of stimulus TF during baseline period when mice did not lick or move. We aligned neural responses to fast TF pulses (50 ms stimulus samples 1× s.d. above baseline TF of 1 Hz; Fig. 2a–c and Methods ), and quantified their peak time (Fig. 2d ) and duration (full width at half peak value; Fig. 2f ), which closely matched those estimated by the GLM (Fig. 2e,g and Extended Data Fig. 5a–d ). As expected, brain regions in early visual system (dorsal lateral geniculate complex (LGd), primary visual cortex (VISp) and superficial superior colliculus (SCs)) responded earliest to fast TF pulses with brief responses that faithfully tracked the stimulus TF (Fig. 2b,d–i ). By contrast, brain regions outside the visual system containing TF-responsive units responded significantly more delayed to fast TF pulses (Fig. 2b–e,h ) and exhibited more prolonged responses than neurons in visual areas (Fig. 2b,c,f,g,i and Extended Data Fig. 4 ). Specifically, neurons in frontal motor cortex, basal ganglia, cerebellum and some regions of the midbrain and thalamus maintained the representation of sensory evidence for several hundred milliseconds beyond the duration of the stimulus sample that triggered the response (Fig. 2b,c,f ).
a , Schematic of identification of fast (TF pulse > 1 s.d.) and slow (TF pulse < –1 s.d.) TF pulses fluctuating around the mean baseline stimulus TF. b , Single-neuron examples of fast and slow TF pulse responses from selected areas across the brain (mean with 95% confidence interval). FR, firing rate. c , Fast TF pulse responses of all TF-responsive neurons in all brain areas with ten or more TF-responsive units. d , Distribution of response peak times estimated from fast TF pulse responses for each brain area with ten or more TF-responsive units (grey line and circles indicate median peak time per area). e , Comparison of median peak times estimated from fast TF pulse responses (left column) and GLM weights tracking TF fluctuations (GLM TF kernels; see Extended Data Fig. 2 for example kernels; Methods ) for each area (right column). f , Distribution of fast TF pulse response half-peak widths (estimated from fast TF pulse responses) for each area with ten or more TF-responsive units (grey line and circles indicate median peak time per area). g , Median fast TF pulse response half-peak widths compared with half-peak widths of the GLM TF kernel. h , Fast TF pulse response peak times across major brain area groupings (median and 95% confidence interval; brain areas in each group are listed in Supplementary Table 1 ). i , Fast TF pulse response half-peak widths across major brain area groupings (median and 95% confidence interval). Wilcoxon rank sum test. Values of n for each brain area grouping are presented in Supplementary Table 1 and definitions of brain area abbreviations can be found in Supplementary Table 2 . NS, not significant.
Parallel sensory integration in premotor areas
The longer timescales of neural responses to fast TF pulses outside the visual system suggests that these areas can integrate multiple samples of behaviourally relevant visual input. Indeed, previous modelling of mouse behaviour in this task shows that mice are guided by TF fluctuations unfolding over several hundred milliseconds 14 . Although this suggests that mice use temporal integration of stimulus TF to detect changes, they may also respond to outliers in stimulus to guide their lick responses. To disambiguate between these behavioural strategies (integration versus outlier detection), we applied a combination of analytical and modelling approaches to mouse behaviour to show that mice indeed do use integration of evidence over a timescale of around 0.25 s. First, the decay time ( τ ) of the early lick-triggered stimulus average (psychophysical kernel; see ref. 52 ) is 0.27 s, a time course significantly longer than predicted by an artificial agent relying solely on an outlier detection strategy (Fig 3a,b and Methods ). Second, mice are more likely to lick when two fast pulses occur within 0.25 s of each other than would be predicted by the joint independent effect of two fast pulses (Fig 3d and Extended Data Fig. 6e–i ). Moreover, the independent effect of two fast pulses fully explained the data of the outlier-detection agent (Extended Data Fig. 6h,i ). Finally, a simple leaky-integrator model with a 0.25 s decay time ( τ ) better predicts early lick times and single-trial hit reaction times than when this model is not allowed to integrate evidence (Extended Data Fig. 7b–h ).
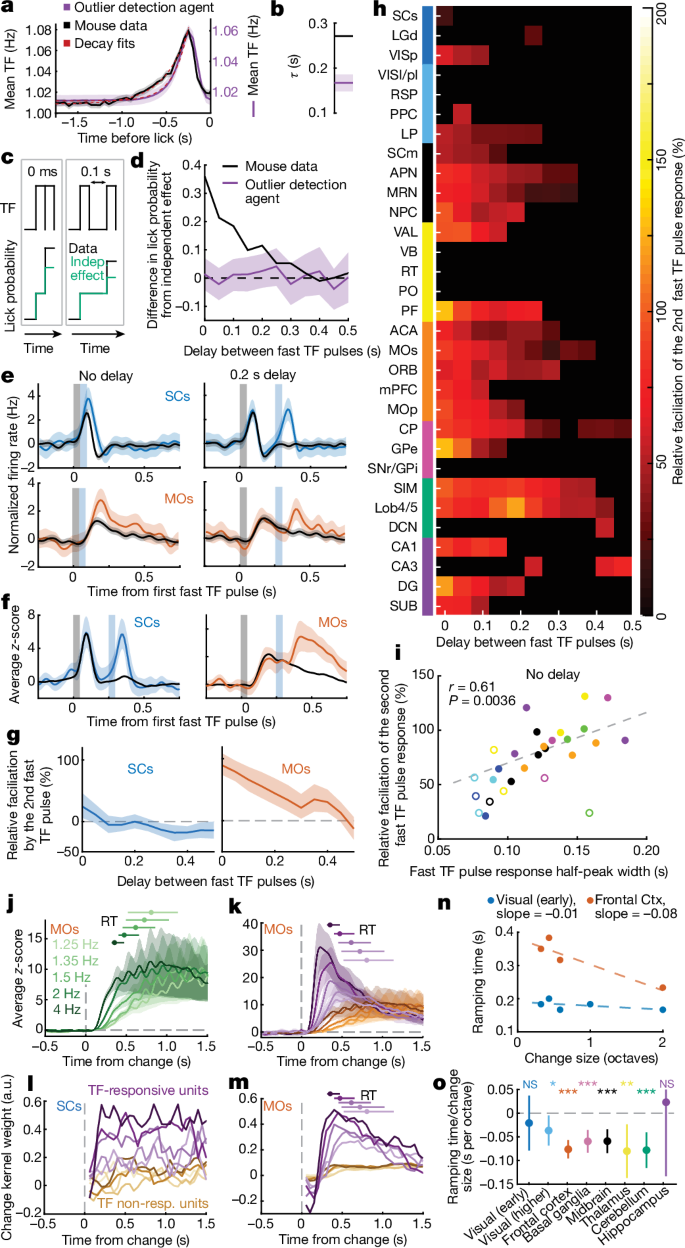
a , Mean stimulus TF preceding early licks in mouse data and outlier-detection agent. Red dashed lines show exponential decay fits. b , Decay time of the exponential fits in a . c , Schematic showing how lick probability is affected by two fast TF pulses that either integrate temporally (black) or act independently (indep.; green). d , Difference between observed early lick probability after two sequential fast TF pulses and the one predicted from their independent effect (Extended Data Fig. 6e–g ), normalized by the probability from independent effect, shown as a function of delay between pulses. Data are mean with 95% confidence intervals. e , Responses to a single fast TF pulse (black) or a sequence of two fast pulses separated by 0 s (left) or 0.2 s (right) in example neurons from SCs and MOs. f , Average response to a sequence of two fast TF pulses separated by 0.2 s delay from all TF-responsive neurons in SCs (left) and MOs (right). g , Facilitation of response to the second fast TF pulse as a function of delay between two pulses for TF-responsive units in SCs and MOs. h , Same as g , but for all brain regions with at least ten TF-responsive units. Only time points with 95% confidence interval above zero (bootstrap test) are shown. i , Pearson correlation between second fast TF pulse facilitation and the median half-peak width of response to fast TF pulse across brain regions ( P value based on t -statistic). Correlation excludes brain regions without significant facilitation, shown as open circles. j , Average activity of MOs units aligned to TF change onset on hit trials, split by change magnitude. Reaction times (RTs) per magnitude are shown as median (dots) with ranges between 25th and 75th percentiles. k , Same as j , but with the MOs population split into TF-responsive (shades of purple) and TF non-responsive (shades of orange) units. Darker colours correspond to larger change magnitudes. l , m , Mean GLM weights tracking activity after change (change kernels) from SCs ( l ) and MOs ( m ) units, derived from activity during change periods. Kernels shown for TF-responsive and non-responsive units, across different change magnitudes. Colour coding as in k . Reaction times shown as in j . a.u., arbitrary units. n , Each dot is the time to 50% of the peak value (ramping time) of the average change kernel across TF-responsive units in early visual areas and frontal cortex (Ctx), shown per change magnitude. o , Scaling of ramping time in activity with change size: each point represents a slope (in seconds per octave) of the linear fit to the dependence shown in m , for each group of brain regions. Bootstrap test. Values of n for each brain region and brain region group are presented in Supplementary Table 1 and definitions of brain area abbreviations can be found in Supplementary Table 2 . In all panels, shaded regions or error bars indicate 95% confidence intervals.
Given that lick responses depend on integrating the stimulus TF over several hundred milliseconds, we next determined the neural correlates of this integration process. We reasoned that a prolonged response to a fast TF pulse serves as a neural substrate for temporal integration of multiple fast TF pulses, by allowing responses to successive fast TF pulses to build on each other. By finding instances during the baseline period when two fast TF pulses occurred at a given delay from each other (Fig. 3e and Methods ), we calculated the average response across all TF-responsive units in a brain region to those pulses, and measured the amount of response facilitation to the second fast pulse relative to the first fast pulse response (Fig. 3f,g , and Fig. 3e for single-neuron examples; Methods ). The majority of early and higher-order visual areas did not show facilitated responses to the second fast pulse even at a 0.1 s interval between the pulses (Fig. 3g,h and Extended Data Fig. 5e ), whereas thalamic lateral posterior nucleus (LP) and hippocampal regions showed facilitation of up to 0.2–0.3 s inter-pulse delay. Across non-visual thalamus, facilitation was observed only in ventral anterior-lateral complex (VAL) and parafascicular nucleus (PF), the key nodes in cortico-cerebellar and cortico-basal-ganglia loops, respectively 29 , 53 , 54 . Most regions in frontal cortex, basal ganglia, cerebellum and midbrain exhibited significant facilitation around 0.2–0.4 s from the first fast pulse (Fig. 3g,h ), resembling the behavioural integration timescales (Fig. 3b,d ). The amount of relative facilitation to the second fast TF pulse correlated with response duration to a single fast TF pulse across brain regions (Fig. 3i ), highlighting that one is a prerequisite for the other.
Thus far, we had isolated the sensory evidence representations by studying them in the absence of movement (that is, baseline period of the trial). Typically, however, neural representations of sensory integration are studied by examining neural responses during presentation of stimuli that trigger the learned response, when there is an overlap of multiple correlated signals related to sensory integration, movement preparation and execution 1 , 37 , 55 . There, evidence integration is inferred by the ramping of neural responses that scale with stimulus strength 37 . Similarly, we found that in regions that integrate pulses of sensory evidence during the baseline period (Fig. 3e–h ), such as MOs, the slopes of ramping activity in the change period scaled with the magnitude of the TF change (Fig. 3j ). Notably, the TF-responsive subpopulation responded more strongly than the rest (Fig. 3k ), with its ramping activity starting and peaking considerably earlier.
To account for the influence of the mouse’s movement on these response profiles, we used the visual response components of the GLM fitted separately to neural responses for each change magnitude (Fig. 3l,m and Methods ). In most areas outside of the visual system and hippocampus, the visual response components of TF-responsive neurons showed ramp-like activity that steepened with increasing change magnitudes, suggesting that these neural populations implement temporal integration of sensory evidence as mice report the change (Fig. 3l–o , Extended Data Fig. 7n,o and Methods ). Moreover, for comparison, early visual areas, such as SCs, exhibited step-like, sustained responses to different change magnitudes (Fig. 3l–o and Extended Data Fig. 7n,o ), thus signalling the change in stimulus TF, but without integration. This is consistent with the early visual system faithfully tracking the fluctuations in sensory input, whereas downstream structures have the capacity to integrate the stimulus stream, essentially denoising it, thus making sensory change detection easier (Extended Data Fig. 7k–m ).
These results reveal that temporal integration of sensory evidence is a parallel, distributed, multi-regional computation—implemented by transforming transient responses to sensory input in visual areas into prolonged representations of integrated sensory evidence in frontal cortex, basal ganglia, cerebellum, thalamus and midbrain structures—which does not propagate to motor execution nuclei in the medulla.
Learning enables widespread sensory integration
We next tested whether the encoding of sensory evidence outside the visual system is intrinsic to the brain regions themselves or a result of learning the relevant stimulus–reward associations. We recorded neural activity in untrained mice (6,215 units, 45 sessions, 6 mice) that had been exposed to the same stimuli but given random rewards (Fig. 4a,b and Methods ), thus never associated changes in stimulus TF with reward. As expected, we found significant fractions of neurons encoding fluctuations in stimulus TF in the visual system (SCs, LGd, LP and VISp) and parts of the midbrain (APN and SCm) in untrained mice. However, we did not find cells with prominent TF responses in frontal-motor cortex, cerebellum, striatum or MRN—regions that in trained mice respond to TF fluctuations (Fig. 4c–e and Extended Data Fig. 8a ). This demonstrates that encoding of sensory evidence in regions outside the visual system—where the sensory evidence is integrated—to a large degree, emerges with learning.
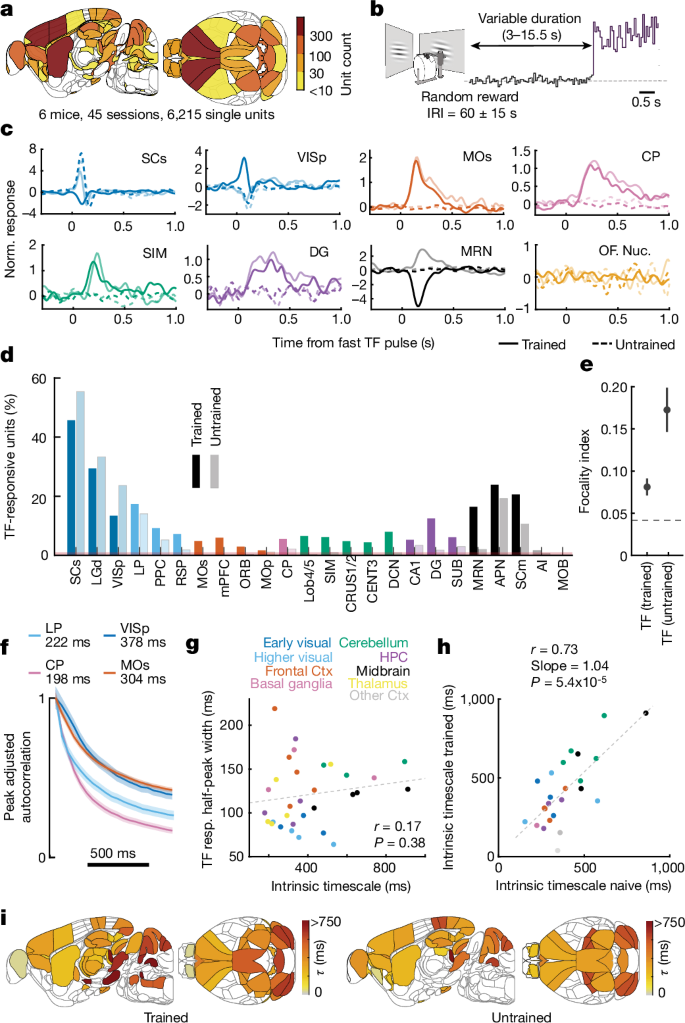
a , Schematic of stimulus presentation with random reward delivery used for recordings in untrained mice ( Methods ). b , Brain maps of unit counts recorded from untrained mice. IRI, inter-reward interval. c , Examples of top two (lowest P value) fast TF-responsive neurons in trained mice (solid lines) or untrained mice (dashed lines) in SCs, VISp, MOs, CP, SIM, DG, MRN and in the orofacial motor nucleus. Norm., normalized. d , Percentage TF-responsive units in all brain areas with more than 40 neurons recorded in both trained and untrained mice. e , Focality index of distribution of TF-responsive units across areas with more than 40 neurons recorded in both untrained and trained mice. In untrained mice, TF-responsive units were confined to a much more limited set of brain regions, compared to trained mice, leading to a significantly higher focality index ( n = 24 overlapping brain regions; P < 0.001, bootstrap test ( Methods )). Error bars show 95% confidence intervals ( Methods ). f , Examples of autocorrelation functions from which intrinsic timescales are estimated (that is, τ of decay of autocorrelation function). Error bars are 95% bootstrapped confidence intervals. g , Pearson correlation ( P value based on t -statistic) between intrinsic timescales and median half-peak width of responses to a fast TF pulse for all TF-responsive neurons across the brain of trained mice. h , Pearson correlation ( P value based on t -statistic) between intrinsic timescales in untrained mice and trained mice. i , Brain maps of intrinsic timescales of trained mice (left) and untrained mice (right). See Supplementary Table 2 for definitions of brain region abbreviations.
To test whether the integrative properties of neurons in non-visual areas are shaped by learning, we assessed whether stimulus integration can be predicted from intrinsic timescales of neural firing of each area. Intrinsic timescales of activity in cortical areas in non-human primates and rodents, defined as the time constant of autocorrelation function of each neuron’s activity, have been suggested to determine duration of task-relevant responses 8 , 23 . However, we did not find intrinsic timescales of neural activity (measured in the inter-trial periods devoid of visual stimuli and movement) to correlate with the duration of fast TF pulse responses across different brain regions (Fig. 4f,g ) or in individual neurons (Extended Data Fig. 8b–e ). Notably, the intrinsic timescales of individual brain regions were similar in trained and untrained mice, indicating that they are an intrinsic property of each area that is unaffected by learning (Fig. 4h,i ). Together, these results imply that representation and integration of sensory evidence emerge with learning in most association and premotor areas outside of the visual system.
Evidence-encoding cells initiate preparatory activity
We next explored how the integrated sensory evidence is transformed into preparation of an action that reports the decision. Preparatory activity before action initiation has been observed in multiple brain areas during motor planning and in decision-making tasks 4 , 15 , 28 , 30 , including our task (Figs. 1j and 5a ). Given that neurons downstream of the visual system encode both sensory evidence and lick preparation (Fig. 1j and Extended Data Fig. 3d ), we tested whether evidence integration and preparatory activity engage similar patterns of activity in these brain regions. We computed the alignment of population vectors between responses to a single fast TF pulse (Fig. 5a,b , left) and preparatory activity before the early lick onset (Fig. 5a,b , right) of TF-responsive subpopulations in different brain regions. In MOs (Fig. 5c ) and other areas outside of the visual system capable of integrating sensory evidence—including frontal cortex, cerebellum, midbrain and basal ganglia—these population vectors were significantly aligned (Fig. 5d ), whereby neurons that increase their firing to fast TF pulses also increase their activity prior to lick initiation, and vice versa (Fig. 5c ). By contrast, no such relationship was observed in areas that do not integrate sensory evidence (Fig. 5d ), such as SCs (Fig. 5c ). These results imply a widespread coupling between integration of sensory evidence and movement preparation, as previously observed in monkey lateral intraparietal area (LIP) and frontal cortex 22 , 37 , but which we find to be far more widespread across sparse subpopulations of frontal cortex, basal ganglia, cerebellum, thalamus and midbrain.
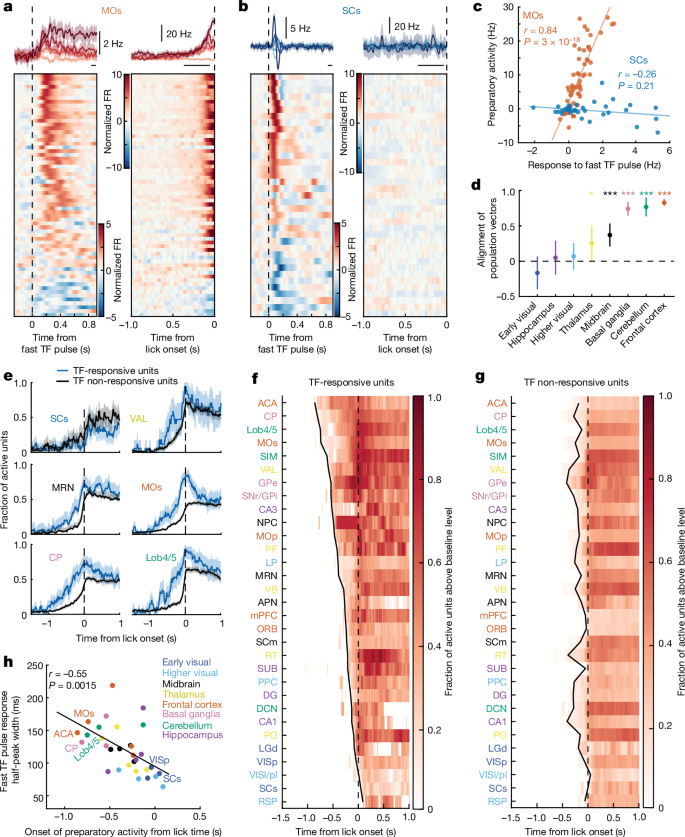
a , Left, mean responses to a fast TF pulse of five example TF-responsive units in MOs (top) and responses to a fast TF pulse for all TF-responsive units in MOs ( z -scored firing rate) (bottom). Right, activity of the same neurons aligned to early lick onset. b , Same as a , but for TF-responsive units in SCs. Horizontal black lines indicate windows of activity used to calculate the alignment of population vectors in c . c , Alignment (Pearson correlation; P value based on t -statistic) between responses (baseline subtracted) of TF-responsive MOs or SCs units to a fast TF pulse and their preparatory activity before the early lick. d , Mean alignment of population vectors (correlation in c ) for each group of brain regions (bootstrap test). See Supplementary Table 1 for n of each brain region group. e , Fraction of significantly active units ( P < 0.01, z -test) as a function of time, shown separately for TF-responsive and TF non-responsive units for six example brain regions. Values of n for each brain region are presented in Supplementary Table 1 . f , Fraction of active TF-responsive units (thresholded by lower 95% confidence interval greater than zero, bootstrap test) as a function of time from the hit-lick onset, shown for each brain region. Brain regions are sorted according to the time of the earliest, significantly active fraction (black line; Methods ). g , Same as f , but for the TF non-responsive subpopulation. h , Relationship between the onset of preparatory activity in TF-responsive units and their median response duration to a fast TF pulse across brain regions. Pearson correlation and corresponding P value from t -statistic are shown on top. In all panels, shaded regions and error bars indicate 95% confidence interval. See Supplementary Table 2 for definitions of brain region abbreviations.
If accumulation of evidence contributes to the build-up of preparatory activity, we would expect the neural subpopulations that integrate evidence to be recruited first prior to a decision to a lick, and that brain regions with longer timescales of integration would have an earlier onset of preparatory activity. Indeed, prior to hit-lick onset during the change period, the TF-responsive populations were recruited significantly earlier than the TF non-responsive populations in areas integrating sensory evidence, including the frontal cortex, basal ganglia, cerebellum and midbrain (Fig. 5e–g , Extended Data Fig. 9b,c and Methods ). The earliest differences in activation were observed across several brain subdivisions, including ACA, MOs, striatum (CP) and Lob4/5 (Extended Data Fig. 9b,c ). Moreover, the onset of preparatory activity of the TF-responsive subpopulation scaled with the duration of response to a fast TF pulse (Fig. 5h and Extended Data Fig. 9d ), revealing that the longer timescales of integration lead to an earlier onset of preparatory activity. Together, these results demonstrate that accumulation of evidence contributes to the build-up of preparatory activity in multiple brain regions downstream of the visual system.
Brain-wide orthogonal dynamics surrounding action
Previous studies have found that population activity in motor cortex transitions between orthogonal sets of dimensions (subspaces) before and after movement onset 33 , 34 . Following movement onset, activity occupies a ‘movement’ subspace, in which projections of activity closely resemble the muscle activity during movement execution. Prior to movement onset, the patterns of activity are different and confined to an orthogonal subspace (‘movement-null’), wherein activity builds up or persists, but does not drive the movement itself. To understand the neural dynamics during the transition between movement preparation and execution in our task, we applied the same analysis framework to each brain region population activity on hit-lick trials, by decomposing population activity into projections onto movement and movement-null dimensions ( Methods ). We defined the movement dimensions as those that captured the best similarity with the activity of orofacial motor and premotor nuclei that drive licking 56 , 57 (Extended Data Fig. 10b,c ), and a set of movement-null dimensions orthogonal to them, wherein activity can reside without directly affecting licking.
We first tested whether the preparatory activity occupies a movement subspace or is orthogonal to it, as previously demonstrated in primary and premotor cortex 33 , 34 , 35 (Fig. 6a , orthogonal modes hypothesis). Figure 6b–d shows MOs activity aligned to hit-lick onset and projected onto the first movement and movement-null dimensions (see also Extended Data Fig. 10b–d ). Relative occupancy of these subspaces around lick onset (Fig. 6e,f and Methods ) revealed that pre-lick activity in MOs predominantly resided within the movement-null subspace (Fig. 6e , and was largely one-dimensional (Extended Data Fig. 10c )), and then transitioned into the movement subspace after the lick onset. Of note, preparatory activity was confined to the movement-null subspace across all other brain regions (Fig. 6f and Extended Data Fig. 11a,b ).
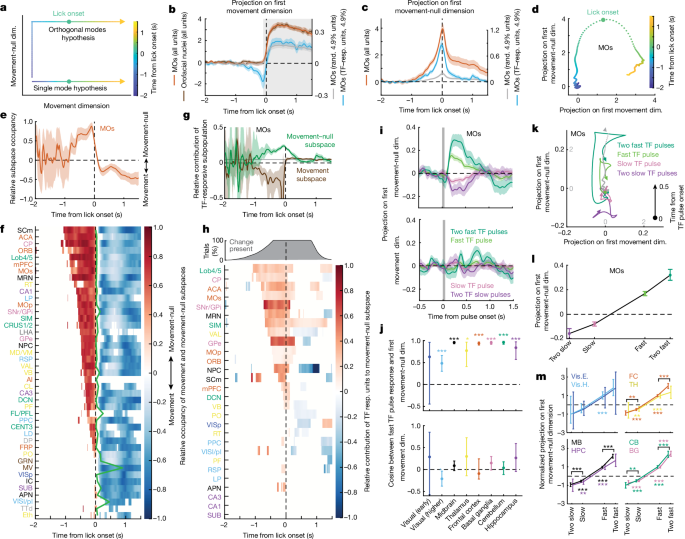
a , Schematic of two hypothetical ways population activity can transition from movement preparation to execution. Preparatory activity and action execution proceed either along the same mode of activity (single mode hypothesis) or are orthogonal to each other (orthogonal modes hypothesis). Dim., dimension. b , Mean projection of all MOs neuron activities around lick on hit trials onto the first movement dimension, defined by activity in orofacial nuclei in the time window around lick (grey; see Methods ). Projection of activity of TF-responsive subpopulation of MOs is shown in blue ( Methods ; scale on the right); projection from a random (rand.) sample of MOs neurons (grey; matched to number of TF-responsive neurons; scale on the right). c , Projection of MOs activity onto the first movement-null dimension during hit trials. d , Same as b , c , but shown in a state-space formed from first movement and movement-null dimensions. Dots correspond to the state of MOs activity in 10-ms bins. Time relative to lick onset is indicated by colour. e , Relative occupancy of MOs activity in movement versus movement-null subspaces as a function of time ( Methods ). f , Same as e , but across brain regions (excluding brain regions with poor goodness of fit ( R 2 < 0.8) to activity in orofacial nuclei; Extended Data Fig. 10d ). Only time points with relative occupancy significantly different from zero ( P < 0.05, bootstrap test) are shown (also for h ). Brain regions are sorted according to the earliest latency of significant relative occupancy. Time of peak occupancy in movement-null subspace is shown by the green line. g , Relative contribution of TF-responsive subpopulation to movement-null and movement subspaces. The grey line indicates the value expected from a random sample of neurons from MOs (matched to number of TF-responsive neurons). h , Same as g , but shown across brain regions sorted by latency of significant contribution of TF-responsive subpopulation. Top, fraction of trials with ongoing change epoch. i , Projections of MOs population responses to pulses of sensory evidence onto the first movement-null (top) and movement (bottom) dimensions. j , Cosine of the angle between population response to a fast TF pulse and first movement-null (top) and movement (bottom) dimensions. Data pooled across grouped brain regions (mean ± 95% confidence interval; bootstrap test). k , MOs population responses to pulses of sensory evidence (0–0.5 s after the pulse onset), shown in state-space formed by first movement and movement-null dimensions. Overlaid, MOs preparatory activity (grey) up to 100 ms before hit-lick onset (note the different scale). l , Peak value of projections of MOs responses to a slow or fast TF pulse, or two sequential fast or two sequential slow TF pulses, onto the first movement-null dimension. m , Same as l , but for groups of brain regions (bootstrap test). BG, basal ganglia; CB, cerebellum; FC, frontal cortex; MB, midbrain; Vis.E., visual (early); Vis.H., visual (higher). In all panels, shaded regions or error bars indicate bootstrapped 95% confidence intervals ( Methods ). Values of n for each brain region or brain region group are presented in Supplementary Table 1 and definitions of brain area abbreviations can be found in Supplementary Table 2 .
Shortly following lick onset, population activity transitioned from movement-null into the movement subspace, almost concurrently throughout the brain. This state transition could result only from an increase in activity within movement subspace (Extended Data Fig. 11a ) or also from a decrease in activity within the moment-null subspace following lick onset. Consistent with the latter, activity within movement-null subspace peaked and then sharply decreased immediately after the lick onset in most brain regions that had preparatory activity (Fig. 6f , green line, and Extended Data Fig. 11b, c ).
Together, these results reveal that the abrupt transitions in neural dynamics between orthogonal movement-null and movement subspaces at movement onset is a general computational feature observed in most association and premotor brain areas.
Linking evidence integration and motor dynamics
If accumulation of visual evidence drives preparatory activity, which resides in movement-null subspace, one would expect TF-responsive units to have a disproportionate contribution to activity in movement-null subspace. To test this, we decomposed projections onto movement and movement-null dimensions into a sum of contributions from TF-responsive units and the rest of the population (see Methods ). For example, in MOs, we observed a disproportional contribution from TF-responsive subpopulation to the preparatory activity within the movement-null subspace (Fig. 6c,g ). Applying this analysis across all brain regions, we found that the TF-responsive subpopulation contributed disproportionately to the preparatory activity in a more restricted subset of areas (Fig. 6h and Extended Data Fig. 11d,e ): frontal cortex (ACA, MOs, MOp, ORB and mPFC), cerebellum (Lob4/5, SIM and DCN), basal ganglia (CP, SNr/globus pallidus internal segment (GPi) and GPe), as well as some regions of the midbrain (MRN, NPC and SCm) and thalamus (VAL and ventrobasal complex (VB)). Notably, these predominantly premotor areas integrated evidence over longer timescales (Extended Data Fig. 11f ; see also Fig. 5h ), emphasizing the link between evidence accumulation and preparatory activity.
Sensory evidence should no longer be informative of choice once the animal has committed to its decision. Accordingly, the contribution of TF-responsive units to preparatory activity in movement-null subspace collapsed to chance level after lick onset in most premotor areas in which TF-responsive units disproportionately drove preparatory activity (Fig. 6h ; see Extended Data Fig. 11g for a comparable analysis in movement subspace). This collapse is consistent with the cessation of evidence accumulation despite the continuous presence of the change stimulus (see also Fig. 3j–l ).
Consistent with the observations that preparatory activity and responses to pulses of sensory evidence are aligned within TF-responsive population of neurons (Fig. 5c,d ) and that the preparatory activity of the entire population is confined to the movement-null subspace (Fig. 6f ), we found that a response to TF pulse is aligned with the dimension that captures the most variance of the preparatory activity (first movement-null dimension) in most regions beyond the early visual system (Fig. 6i,j , top, k and Extended Data Fig. 12a ). By contrast, responses to fast TF pulses were not positively aligned with the first movement dimension in any brain region group (Fig. 6i, j , bottom, k ). Consequently, outside of the early visual system, we find that the integration of sequential pulses of evidence primarily takes place along the first movement-null dimension (Fig. 6k–m and Extended Data Fig. 12b ). This provides an explanation for how sensory evidence can recruit activity across the majority of brain regions without directly driving the movement.
Here we describe the brain-wide neural implementation of evidence integration, movement preparation and execution—the key processes underpinning decision-making—revealing a global mechanism for transforming ambiguous sensory evidence into goal-directed actions. We show that evidence integration is a widespread phenomenon that emerges with learning and is implemented in a sparse population of neurons across most premotor areas. In these neurons, the timescales of integration are independent of intrinsic regional dynamics, suggesting that they are shaped by task experience. Notably, evidence integration and movement preparation are encoded in the same subspace of population activity across the brain, orthogonal to movement-related dynamics. Activity in this subspace was driven by neurons integrating evidence and collapsed at movement onset, allowing the integration process to reset, whereupon activity transitioned into a different subspace for movement execution concurrently across the brain. Our work links evidence accumulation onto motor dynamics on a brain-wide scale, unifying concepts from motor control and decision-making fields into a common framework for understanding how sensory evidence controls actions through global neural mechanisms.
Our finding that only expert mice exhibited robust encoding of visual input in almost all brain areas outside the visual system is consistent with previous reports of learning increasing the connectivity and correlations between cortical and subcortical regions 58 , 59 , 60 , which may explain the distributed encoding of task variables across cortical and subcortical structures in trained animals 3 , 4 , 14 . We now show that these learning-induced multi-regional representations of task-relevant stimuli are not simply a distributed echo of the sensory input, but a transformed and integrated representation explicitly used to guide decisions. In association and premotor areas, such as frontal-premotor cortex, basal ganglia, cerebellum, parts of midbrain and thalamus, the prolonged responses to individual samples of evidence enabled their integration on a timescale of several hundred milliseconds, consistent with timescales of behavioural integration (Fig. 3 and Extended Data Fig. 6 ). This is a key distinction from visual areas, such as VISp and SCs (and primate middle temporal visual area 37 (MT)), where neurons do not integrate evidence (Fig. 3 ). Consequently, the integration of ambiguous task-relevant stimuli becomes a multi-regional distributed process implemented in a sparse population of neurons, and one that emerges with training as mice learn the value of the relevant stimulus feature. Notably, in our task both neural and behavioural evidence integration is ‘leaky’, consistent with the idea that in dynamic sensory environments perfect integration is not an optimal behavioural strategy 52 . Instead, leaky integration of a noisy stimulus stream is beneficial as it increases the signal relative to noise by temporally smoothing the input (Extended Data Fig. 7k–m ).
We found that the timescales of integration are as diverse across the entire brain as has been shown across cortex 8 , 14 . However, evidence integration times were not explained by the intrinsic timescales within each area, previously suggested to be predictive of response duration and ability to integrate stimuli in cortex of non-human primates and mice, respectively 8 , 23 , 24 , 25 , 26 . A possible reason for this discrepancy may be that our task allows estimation of both the intrinsic timescales and stimulus integration times in the absence of potentially confounding movement signals. In this study, we found that intrinsic timescales remain stable with learning, confirming they are an inherent property of each area. In fact, decoupling of intrinsic timescales from integration times may be advantageous because it allows task demands to sculpt the timescales of integration 26 , 61 . This decoupling may be implemented by learning mechanisms 59 , 62 that shape the activity propagating in multi-regional long-range loops involving cortex, basal ganglia, cerebellum, thalamus and midbrain, as observed during motor planning 28 , 29 , 35 , 53 , 54 .
To understand how evidence integration leads to action, we adopted a framework developed for understanding the neural dynamics of movement generation, which identifies the relationship between modes of population activity that precede and follow action onset 33 , 34 , 63 . Using this framework, we demonstrate that neural dynamics of lick preparation and lick execution occupy distinct, orthogonal subspaces in most subdivisions of the brain, as previously shown in primate primary and premotor cortex during arm movements 33 , 34 and more recently in the mouse brain during memory-guided movements 4 . Of note, the subpopulations of neurons capable of integrating sensory evidence initiated and dominated preparatory activity in movement-null subspace. We found preparatory activity to originate earliest in regions with the longest integration timescales, such as frontal cortex, basal ganglia and cerebellum, and then transition abruptly into an orthogonal subspace upon movement initiation almost instantaneously in all brain regions investigated. This demonstrates that the transformation of accumulated evidence into movement planning and execution takes place within and across subspaces of neural activity that are shared across multi-regional circuits, rather than proceeding successively across a subset of specialized brain areas. Future research should determine the degree to which the principles of brain-wide neural dynamics observed in our study generalize to tasks involving multiple sensorimotor contingencies.
A clear advantage of orthogonalizing neural dynamics during decision-making is that it allows computations such as evidence accumulation, movement preparation or movement execution to proceed within the same population of neurons 64 . Our results highlight a particular advantage of occupying the movement-null subspace as it allows evidence integration to take place without directly causing movement. Accordingly, the lack of responses to visual evidence in the orofacial nuclei in medulla, which become active only upon lick initiation, demonstrates that brain-wide preparatory activity patterns driven by sensory evidence are incapable of driving the activity in motor circuits that control mouth and tongue movements.
The transition of population activity from movement-null to movement subspace is thought to proceed via a brief release of activity occupying movement-null subspace as an input to the movement subspace 34 , which triggers the action. In a delayed response task using an explicit auditory Go cue, a trigger signal in premotor cortex depends on a pedunculopontine nucleus (PPN)/MRN–thalamic circuit 35 . Our task, however, requires an internally generated trigger when sufficient evidence is accumulated. Future work is needed to elucidate the regions that generate the trigger signal, with likely candidates receiving information from areas with early onsets of preparatory activity such as ACA, MOs, CP and Lob4/5. Conversely, an action initiation signal may propagate to the movement-null subspace, since the contribution of evidence-accumulating neurons to the movement-null subspace collapsed shortly following action onset, even though the change stimulus was still present, thus allowing the integration process to reset. This observation suggests that evidence-integrating neurons perform this function only when it is relevant and before the mouse has committed to an action. These findings imply that activity in one orthogonal subspace can influence the activity in the other subspace, highlighting the dynamic interplay between movement-null and movement-related neural dynamics.
In summary, we demonstrate that learning recruits a neural subpopulation that is widely distributed across the brain, which concurrently integrates evidence and drives movement preparation, allowing sensory evidence to control global neural dynamics required for generation of behavioural responses.
All experiments were performed under the UK Animals (Scientific Procedures) Act of 1986 (PPL: PD867676F) following local ethical approval by the Sainsbury Wellcome Centre Animal Welfare Ethical Review Body. A total of 21 C57BL/6 J male mice (age = 34.5 ± 15.8 weeks (mean ± s.d.)) were used for electrophysiological recordings. Fifteen mice first underwent head-fixed behavioural training prior to acute electrophysiological recordings (see ‘Task and training stages’), and six mice (untrained mice) only underwent habituation to the recording setup prior to acute electrophysiological recording.
Prior to behavioural training and recordings, all mice were implanted with a head-fixation bar under approximately 1.5% isoflurane and administration of Meloxicam (5 mg kg −1 ) to allow for head-fixation during behavioural training and electrophysiological recordings.
During training, mice were co-housed with littermates in individually vented cages. After implantation of the recording chamber, mice were singly housed to protect the implant. Mice were housed in reversed day–night cycle lighting conditions, with the ambient temperature and humidity set to 23 °C and 56% relative humidity, respectively.
Behavioural task
The design of the behavioural task was as previously described in ref. 14 . In brief, mice were head-fixed and placed on a polystyrene wheel. Two monitors (21.5 inch, 1,920 × 1,080, 60 Hz) were placed on each side of the mouse at approximately 20 cm from the mouse head. The monitors were gamma corrected to 40 cd m −2 of maximum luminance using custom MATLAB scripts utilizing PsychToolbox-3. The stimulus presentation was controlled by custom written software in MATLAB utilizing PsychToolbox-3. The visual stimulus was a sinusoidal grating with the spatial frequency of 0.04 cycles per degree resulting in 3 grating periods shown on a screen. Each trial began with a presentation of a grey texture covering both screens. After a randomized delay (at least 3 s plus a random sample from an exponential distribution with the mean of 0.5 s), the baseline stimulus appeared. The TF of the grating was drawn every 50 ms (3 monitor frames) from a lognormal distribution, such that log 2 -transformed TF had the mean of 0 and s.d. of 0.25 octaves and the geometric mean of 1 Hz. The direction of drift was randomized trial to trial between upward or downward drift. The sustained increase in TF, referred to in the text as change period, occurred after a randomized delay (3–15.5 s) from the start of baseline period and lasted for 2.15 s. For early and late blocks training (stage 8), change period times were sampled between [3, 8] s and [10.5, 15.5] s, respectively, with the delay from the earliest allowed change period sampled from an exponential distribution with a mean of 4 s. Random 15% of trials were assigned as no-change trials and did not have a change period. For stage 8 training, 10% of trials were designated to be probe trials and had a change time drown from the distribution of the other block type. Because there were no qualitative differences in neural TF pulse response between early and late blocks (data not shown) we have combined data from both block types for analyses throughout this manuscript. Findings related to stage 8 (early and late blocks) will be presented in an upcoming paper.
Mice were trained to report sustained increases in TF by licking the spout to trigger reward delivery (drop of soy milk). Licks that occurred outside of the change period are referred in the text as early licks. If mice moved on the wheel (movement exceeding 2.5 mm in a 50-ms window) in either direction, the trial was aborted (stages 7 and 8). If mice did not lick within 2.15 s from the change onset, the trial was considered a miss trial.
Training stages
Following the implantation of the headplate, mice were allowed to recover for a week. After that, mice went through several stages of training:
Mice were handled for 3 to 7 days, until mice were comfortable with being handled by the experimenter. During this stage mice were also habituated to being restrained by being placed into a soft cloth for a short period of time. After the brief restraints they were given a small amount of soy milk as reward.
Next, mice were put on food restriction. Mouse weight was monitored daily with the amount of food given adjusted per mouse to keep them sufficiently motivated for getting rewards and keep their weight no lower than 85% of the original weight prior to food restriction.
Next, mice were head-fixed and placed on the running wheel of the behavioural training setup with the monitors turned off. Mice were allowed to freely run on the wheel, but not encouraged to. Typically, there were 3 habituation sessions, with the duration progressive increasing from 15 to 45 min.
Next, mice were introduced to the visual stimuli used in the task. Mice were initially shown only trials with two largest changes of TF (2 and 4 Hz, lasting 2.15 s), followed by a reward auto-delivery 1.5 s after the change onset. After mice started to robustly make licks during the change period that preceded the reward auto-delivery, they were transitioned to the next stage.
Here only hit trials were rewarded, early licks and running did not result in termination of the trial.
After mice robustly detected strong changes in the previous step, we introduced trials with weaker changes in TF (1.25 Hz, 1.35 Hz and 1.5 Hz). Additionally, a consequence of an early lick outside of the change period was a mild air-puff to the mouse’s right cheek and a termination of the trial.
After mice detected weaker changes as well (assessed as higher hit rate compared to no-change trials), they were transitioned to the next stage where in order to initiate the trial start (start of the baseline stimulus), mice were required to remain stationary on the running wheel for at least 3 s plus a random sample from an exponential distribution with the mean of 0.5 s. Additionally, after the trial start, a trial was aborted as a consequence of a movement on the wheel.
Finally, after mice reached sufficient proficiency at the previous stage, early and late blocks were introduced. During the session start, a block type was randomly chosen. A block was defined as a period of the session during which a mouse completed 30 hit trials. After completion of a block of trials, the block type was switched to the other block type (early to late or vice versa).
Six mice that were used in the untrained control experiment (Fig. 4e–h ) went through training stages 1–3 above. Following that, they were shown the same stimuli as the trained mice, with the difference that their movements on the wheel or licking the spout did not terminate a trial nor trigger reward. Instead, they were given rewards at random times with inter-reward intervals drawn from the uniform distribution of 60 ± 15 s.
Behavioural setup and data acquisition
Reward delivery (soya milk) was controlled by a solenoid pinch valve (161P011, NResearch) and delivered to the mouse via a spout positioned in front of it. Mouse licking the spout was measured by a piezo element (TDK PS1550L40N) coupled to the spout and amplified with a custom-made amplifier system. Running wheel movement was measured with a rotary encoder (model Kübler) that was connected to the wheel axle. All behavioural data and events, such as piezo signal voltage trace, valve or change period on/off state, etc., were acquired via analogue and digital channels of PXIe-6341 acquisition card (National Instruments) with SpikeGLX ( https://github.com/billkarsh/SpikeGLX ) at 8,474 Hz.
Behavioural data analysis
Psychometric performance, reaction times and lick-triggered stimulus average.
Psychometric curves were calculated per session by counting the amount of hits relative to all trials where mice did not early lick nor abort. Mean hit rates (performance) and parametric 95% confidence intervals (s.e.m. × 1.96) of hit rates were calculated across sessions ( n = 114) per change size. Mean reaction times and parametric 95% confidence intervals were calculated across sessions ( n = 114) per change size, and p -values were estimated from t -tests.
Lick-triggered stimulus average was estimated by extracting the TF pulses from −1.5 to 0 s preceding early licks and averaged across all trials, revealing mean stimulus TF prior to early licks. Parametric 95% confidence intervals were estimated by calculating the s.e.m. of TF values at each 50 ms bin (TF pulse resolution) prior to an early lick and multiplying the s.e.m. by 1.96.
Simple behavioural leaky-integrator model
In order to formally test if mice behaviourally integrated stimulus evidence (TF pulses) over time in our task, we constructed a simple behavioural leaky-integrator model with two adjustable parameters: decay time ( τ ), and threshold. We fitted these two parameters by estimating which decay time and threshold predicted most early lick times (from 2 s after trial start, to exclude trial onset licks) correctly for each mouse and then determined the average best-fit decay time and threshold values across mice. For each early lick trial, we calculated the integrated log-scaled TF with decay across the entire trial up until the early lick.
For each early lick trial, we then estimated whether a threshold crossing of the integrated TF had been predicted within a second preceding an actual early lick onset. If this was the case, we considered the model to have predicted the early lick time. If not, we considered the model to not have correctly predicted that trial. We did this for all early lick trials, using a 58 × 151 parameter space: 58 possible decay times spanning from 0.05 s decay time (that is, no integration) to 1,000 s decay time (that is, perfect integration): (50 log-spaced decay times spanning 0.050–3 s, as well as 8 additional very long decay times: 4, 5, 6, 7, 8, 9, 20, 1,000 s), and 151 linearly spaced thresholds spanning [0.01–0.16]. Significance testing of best decay time across mice (that is, larger than no integration (0.05 s)) was done with a t -test.
We also tested if the best-fit decay/integration time parameter estimated from predicting early lick times also outperformed a model with no integration when predicting single-trial hit reaction times (that is, a trial type which the parameters were not optimized on). We did this by comparing actual and predicted reaction times per change size, and calculated Pearson’s correlation between actual reaction times and predicted reaction times per change size. We calculated this by either looking at all reaction times, or only including a subset of trials with reaction times under a defined value (that is, reaction-time cut-off). This was done to better detect if any of the models specifically struggled to predict very late reaction times which may be modulated by non-sensory factors such as such as inattention or lack of engagement. Finally, for significance testing (that is, paired t -test) of whether a model with no integration (decay time = 0.05 s) versus a model with the best-fit decay/integration time (estimated from early lick trials as described above), were significantly different at predicting single-trial reaction times, we z-scored actual and predicted reaction times per change size (to account for change size mean reaction-time differences), and calculated the correlation between all actual reaction times (1 s reaction-time cut-off) and all predicted reaction times of a model with or without integration per mouse, and performed a paired t -test (across mice) of the correlation values from integration versus no integration models.
Outlier detection agent
To test whether mice accumulate evidence over time or merely respond to the instantaneous stimulus, we formulated a null model where behavioural responses are produced via a stochastic outlier detection strategy. Here, an internal decision occurs when a noisy sensory representation of the stimulus crosses a decision boundary, and a response occurs after a stochastic delay. The response is triggered by a single, instantaneous value of the stimulus. However, owing to the stochastic delay, responses may show a gradually decaying statistical dependence on the stimulus history, and may even mimic evidence accumulation strategies such as integration 42 .
Model. According to the outlier detection model, behavioural responses are generated independently for each trial as follows. Let s i be the stimulus amplitude (log TF) at each time point t i . We chose time points to correspond with video frames of the stimulus, which were presented at 60 Hz (3 frames per TF pulse). At each time point, a noisy sensory representation Z i is formed as the sum of the stimulus amplitude and independent and identically distributed (i.i.d.) Gaussian sensory noise ε i (with mean zero and variance σ 2 ):
An internal decision to respond occurs at time D , given by the first time point where the sensory representation exceeds a decision bound b (or ∞ if the bound is not crossed before the stimulus ends):
The hazard function of the decision time is thus:
where Φ is the standard normal cumulative density function (CDF).
A motor response begins at time R , given by the decision time plus an independent, nonnegative stochastic delay ∆ representing the duration of nondecision processes (for example, decision to motor delays):
The delay has a shifted log-logistic distribution with location α , scale β and shape γ , and can be obtained by exponentiating a logistic random variable and then adding a constant. We constrained the location ( α > 0) and shape ( γ > 1) to give the distribution nonnegative support and a bump-like density that decreases on both sides of the mode. The delay time probability density function (PDF) and CDF are:
Because the decision and delay times are independent, the marginal response time distribution is given by the convolution of the decision and delay time distributions. The marginal PDF and CDF of the response time are:
where the decision time probability mass function (PMF) p D can be computed from the hazard function H D above. Because delays are nonnegative, p Δ ( r − d ) = F Δ ( r − d ) = 0 for all d > r , so the above sums need only be computed over time steps up to the given response time.
The outlier detection model was implemented using custom Python software using the NumPy, SciPy, and PyTorch libraries. All computations involving probabilities were performed in log space, using functions designed to avoid numerical under/overflow.
Fitting. A separate model was fit for each mouse in two stages. We first fit the delay time distribution using only trials with the largest change magnitude, then fit the remaining decision parameters using the entire dataset (excluding the abort trials). This two-stage approach relies on the assumption that delays are identically distributed across trials. In return, it allows more direct estimation of the delay time distribution, providing better ability to distinguish between outlier detection and longer-timescale strategies such as integration.
For each trial i , let n ( i ) be the number of time points, \({s}^{(i)}=\{{s}_{1}^{(i)},\ldots ,{s}_{{n}^{(i)}}^{(i)}\}\) be the stimulus amplitudes, and c ( i ) be the time of the change point. For trials where a response occurred, let r ( i ) be the response time, measured as the onset of facial movement (see ‘Motion onset time estimation’ section) and ℓ ( i ) be the subsequent lick time (measured at the reward spout).
Fitting the delay time distribution. We assumed that the greatest change magnitude (geometric mean TF 4 Hz) was large enough to trigger an immediate decision at or near the change point. Under this assumption, the delay time on large-change hit trials can be approximated by the reaction time, which can be directly measured as the time elapsed between the change point and the onset of facial movement. Thus, we fit the delay time distribution (shifted log-logistic distribution) to reaction times on large-change hit trials (denoted \({{\mathcal{T}}}_{{\rm{bighit}}}\) ) by maximum likelihood, subject to the constraints described above:
This approach is conservative for our use of outlier detection as a null model. If the largest changes were not immediately followed by a decision, then delays would tend to be overestimated, causing the fitted outlier detection model to display longer-timescale dependencies that are typically associated with evidence accumulation strategies such as integration. Thus, the risk of falsely rejecting this null model would not increase.
For the largest change magnitude, miss trials predominantly reflected task disengagement rather than typical sensory/motor delays, and were therefore excluded when fitting the delay time distribution. According to a hidden Markov model, disengagement was the a posteriori most probable state for the majority of large-change miss trials (95.2% of large-change misses were during a disengaged state).
Fitting decision parameters. The decision parameters (sensory noise variance and decision threshold) were subsequently fit using the entire dataset, holding the delay time distribution fixed. Here in the general case, the decision and delay times cannot be directly observed, and were marginalized out as latent variables. The decision parameters were chosen to maximize the log marginal likelihood of the observed response data:
For hit and early lick trials (denoted \({{\mathcal{T}}}_{{\rm{nonmiss}}}\) ), the likelihood is given by the marginal probability density of a response at the observed movement onset time. For miss trials (denoted \({{\mathcal{T}}}_{{\rm{miss}}}\) ), the response time is treated as right-censored; its precise value is unknown, but is known to exceed the last time point in the trial. The likelihood for miss trials is thus given by the marginal probability mass lying beyond this point.
Sampling. To statistically compare mouse behaviour to the outlier detection null model, we sampled 10,000 synthetic datasets from the model fitted for each mouse. For every quantity of interest, the value computed from the real data was compared to values computed from each synthetic dataset, comprising 10,000 samples from the null distribution. Synthetic datasets were generated for each mouse as follows.
Each trial used the same change point and stimulus amplitudes presented in the real data. The real stimulus ended after the lick on trials where mice responded, leaving unknown future values that would have been presented had a lick not occurred. Such missing stimulus values were filled in by sampling from the same distribution used to produce the original stimuli (independently for each synthetic dataset).
Given the stimulus, a decision time and delay time were sampled from the distributions p D and p Δ described above. The sum of these quantities yielded a synthetic response time, representing movement onset.
To generate synthetic lick times, we assumed that the additional delay between movement onset and licking was i.i.d. across trials. We therefore sampled with replacement from the measured movement-to-lick delays in the real data. Synthetic lick times were obtained by adding sampled movement-to-lick delays to synthetic movement onset times.
Synthetic lick times were used to determine trial outcomes (hit, early lick, miss). Each trial was classified as a: hit if the lick occurred during the change period; early lick if the lick occurred before the change point; or miss if no lick occurred before the end of the change period.
Effect of magnitude and timing of TF pulses on probability of early licks
For analyses of the effect of TF pulses on probability of early licks we used the training data of the same 15 mice used for Neuropixels recordings. Here we only used sessions where mice reached robust proficiency of the task and were at the final training protocol (mean of 77.5 sessions per mouse). Note that here the time of lick onset was measured from the registration of lick by the spout as opposed to the videography analysis on Neuropixels recording sessions elsewhere in the manuscript. We used only trials where early licks happened at least 2 s after the baseline onset to decrease the influence of impulsive licks on results.
To empirically validate that mice use multiple pulses of sensory evidence to influence their decision to lick during the baseline period, we analysed how early lick probability is influenced by magnitudes and timing of preceding TF pulses. First, we tested whether the deviation of a single TF pulse relative to the mean baseline 1 Hz makes mice correspondingly more or less likely to make an early lick within the subsequent 0.2–1.0 s. For that we separated TF pulses by magnitude (in octaves) into 15 bins such that each bin contained approximately equal number of TF pulses. To calculate the conditional probability of early lick at a certain time after a TF pulse of a given magnitude, we found instances of such events (pulled across all sessions with robust performance for each mouse) and divided them by the total amount of early licks (Extended Data Fig. 6c ). To calculate an overall influence of a TF pulse on early lick probability, we summed conditional probabilities within a [−1, −0.2] s window relative to early lick onset (Extended Data Fig. 6d ):
which can also be written as: P (L|TF) = P 0 + Δ P (L|TF), where
And can be thought as a chance level of making a lick without a deviation of stimulus TF from the mean baseline TF value.
The empirical effect of two TF pulses on lick probability was calculated from behavioural data in a similar way. To compare the measured effect of two TF pulses with their expected effect if they influenced the lick probability independently, we calculated their cumulative independent effect on early lick probability based on empirically measured effect of a single TF pulse on early lick probability. The independent effect of two TF pulses with a delay of Δ t s between them can then be written as follows:
A deviation of lick probability after two TF pulses from the probability predicted by the independent effect of two TF pulses would indicate an interactive effect between pulses, which should be expected if mice utilize integration of sensory evidence. To measure the relative difference between the behavioural result and the expected independent effect of two fast TF pulses (Fig. 3d and Extended Data Fig. 6i ), we calculated:
When applying this analysis to the outlier detection agent data, we used data only from trials that resulted in early licks, meaning that the model made a decision to initiate a lick during the baseline period and before the TF change epoch. For outlier detection agent model that was fitted to a particular mouse data, we sampled the same number of early lick trials across 4,000 synthetic datasets (see section above) as there were present across all behavioural sessions of that mouse. The data was then pulled across all models corresponding to different mice and analysis steps were applied to the combined dataset as described above for the mice data. This procedure was repeated 4,000 times to estimate non-parametric 95% confidence intervals of results from the outlier detection agent.
Electrophysiological recordings
Prior to acute electrophysiological recordings, we habituated mice to the electrophysiological recording setup for 2–7 days (depending on the performance of the mouse in the electrophysiological recording setup), to allow mice to perform optimally during electrophysiological recording sessions.
Once mice were habituated to the recording setup, we implanted a recording chamber with one or two 3 mm craniotomies inside, together with a stainless-steel grounding wire in the contralateral hemisphere, under 1.5% isoflurane together with administration of meloxicam (5 mg kg −1 ) and dexamethasone (2–3 mg kg −1 ). During surgery a kapton disk (Laser Micromachining Limited) was placed on top of the dura inside each craniotomy. The disk had 19 holes with 0.5 mm diameter, arranged in a honeycomb shape, for keeping track of probe insertions. The craniotomy and disk were covered with DuraGel (Cambridge NeuroTech) to protect the brain. A 1–2 mm tall plastic enclosure was then positioned around craniotomies and sealed around the edges with bone cement. Finally, we covered the plastic enclosure with a removable plastic cover, to create a rigid physical barrier over the DuraGel sealed craniotomy, to provide robust protection of the recording preparation between recording sessions. The mice were allowed to recover for 24 h before the first recording session took place.
Electrophysiological data collection was done using Neuropixels 1.0 probes (IMEC, Belgium) and collected with a PXI based system (National Instruments), and saved using SpikeGLX ( https://github.com/billkarsh/SpikeGLX ). For trained mice, we recorded up to 13 sessions per mouse (167 probe insertion from 114 sessions total (15 mice)). For untrained mice, we recorded up to 9 sessions per mouse (89 probe insertions from 45 sessions total (6 mice)). Probes were dipped in CM-DiI (Sigma-Aldrich) prior to insertion. In each session, we inserted up to 2 probes at a time. The probes were always inserted at the same angle within the coronal plane (10° and −15° relative to the vertical axis) to aid subsequent histological probe tract tracing.
At the beginning of each session, we removed the plastic lid above the recording chamber exposing the DuraGel covered craniotomy, and inserted the probe(s) through the DuraGel using microcontrollers (Sensapex) at 5–10 μm s −1 . The probe(s) was allowed to settle for 20 min, to increase stability throughout the recording session. At the end of the session probes were removed (at 15 μm s −1 ) and the plastic cover over the recording chamber was reattached for protection of recording preparation.
The setup for presenting stimuli and monitoring behaviour were identical to the setups in which mice had been trained (see ‘Behavioural task’).
Pre-processing and spike sorting of electrophysiological data
Electrophysiological data was first filtered using CatGT ( https://billkarsh.github.io/SpikeGLX/#catgt ) with modified form of common average referencing (-dlbdmx flag).
Spike sorting. We spike-sorted electrophysiological data from each probe in each session using KiloSort2.0 65 ( https://github.com/MouseLand/Kilosort ). For initial selection of units undergoing further curation, we only selected units designated as ‘good’ (based on cross-correlogram contamination) by KiloSort2.0.
Quality checks. For our electrophysiological recordings of trained mice, we manually inspected and curated, in Phy2.0 ( https://github.com/kwikteam/phy ), every unit which KiloSort2.0 had designated as ‘good’. For our recordings in trained mice this left 44,288 units to be manually inspected and curated, and 15,406 units were kept for analysis after manual curation. Based on the manual curation data from trained mouse recordings (see ‘Manual curation of spike-sorted units from trained mice’), we established a series of heuristics for creating automatic curation of units (see ‘Automatic curation of spike-sorted units from untrained mice’) and used these for recordings from untrained mice.
Manual curation of spike-sorted units from trained mice. We manually inspected and curated all units which KiloSort2.0 had designated as good, based on cross-correlogram contamination. In Phy2.0, we first inspected and merged units that clearly belonged to the same cluster, but had been split by KiloSort2.0, or split the noise from signal in units with clearly separatable noise contamination. We then designated each unit into one of five categories:
Perfect, or almost perfect, with no/very minimal noise, drifting, cutting in/out for the full duration of recording.
Usable and good signal with some noise that cannot be extracted that lasts for the full duration of the recording.
Some drift, but possible physiological change in signal. Clear signal for most of duration of the recording.
Drifting/sudden loss, but otherwise usable/close to perfect. Clear signal for over 50% of the duration of the recording but requires only using a subset of the session.
Noise/useless. Spike shape is not physiological.
Our goal was to remove from analyses units that had large contamination with multi-unit activity, were not recorded throughout the full duration of a session, or were a result of artifacts in recorded signals. We therefore used units designated as category 1–3 above for all further analysis from trained mice.
Automatic curation of spike-sorted units from untrained mice. We next used the manual designations of units to establish a set of criteria for automatic detection of units we would include with manual curation. Based on the manual curation data above we established the following 7 criteria for considering a unit good for analysis:
Firing rate criteria:
Mean firing rate must be above 0.5 Hz.
Rolling 20-min average firing rate cannot drop below 30% (that is, 70% drop from mean) of its mean firing rate.
Rolling 10-min average firing rate cannot drop below 20% (that is, 80% drop from mean) of its mean firing rate.
Rolling 5-min average firing rate cannot drop below 10% (that is, 90% drop from mean) of its mean firing rate.
Inter-spike interval (ISI) violations. Absolute refractory period needs to have <20% estimated contamination rate from other neurons (this is what Kilosort2.0 calls ‘good’).
If there are some spikes in the refractory period, the ISI peak in the first 5 ms cannot be within the first 2 ms.
ISI histograms cannot have sudden large spikes in their shape (that is, peak of ISI cannot be 4 times larger than the second highest peak—that is usually its immediate neighbour).
These criteria selected approximately 90% of units we would have designated with categories 1 (perfect, or almost perfect) or 2 (usable and good signal with some noise) with manual curation, and excluded approximately 85% we would have designated as 4 (drifting/sudden loss) or 5 (noise) with manual curation.
This automatic selection of units was used to select units for analysis from untrained mice recordings and yielded 6,215 units out of 20,292 ‘good’ KiloSort2.0 units.
Clock-drift correction. A shared 1 Hz square wave signal was recorded on the clock of each headstage and National Instruments (NI) acquisition card using a SYNC option in SpikeGLX. Clock drift between spike times from different probes and behavioural events extracted from NI acquisition card recording was corrected post-hoc via TPrime ( https://billkarsh.github.io/SpikeGLX/#tprime ) using the shared square wave signal.
Videography
Acquisition.
High-speed videography of front (100 frames s −1 , 640 × 512 pixels) and side view (50 frames s −1 , 976 × 1,024 pixels) of the mouse face was acquired using two Chameleon3 cameras (CM3-U3-13Y3M-CS, FLIR) with infrared illumination. The videos were acquired in an 8 bit greyscale format. Cameras were configured to send a TTL signal to the National Instruments PXIe board at the start of exposure of every acquired frame. These TTL signals were used to align frame times to the time of behavioural events and spike times.
In order to estimate the pupil size, we trained DeepLabCut 66 to track the pupil size and position using videos acquired with the side camera. The model was trained to track 12 points surrounding the mouse pupil. In order to assess the model performance, after the training the model was tested on videos from sessions not used for training. Pupil size was estimated as an area of an ellipsoidal best fit to the tracked 12 points surrounding the pupil.
Motion energy
For calculation of motion energy, we primarily used videos acquired with the front camera to access a finer temporal resolution (with the exception of 2 sessions where for technical reasons we used a lower jaw ROI from side camera video). To estimate motion onset times, we used ROI centred around the mouse’s face, though nearly identical results were obtained with lower jaw or whisker pad ROIs from the side camera (data not shown). Motion energy was defined as a square root of the sum of squared frame-to-frame pixel value differences, divided by the number of pixels within the ROI.
Movement onset time estimation
In order to find the onset times of orofacial movements, we wanted to estimate the typical noise level of the motion energy signal and find the time points where the signal significantly deviated from the noise-band level. As a first step, we calculated the distribution of motion energy values in a 2-s window centred around the lick registration times. We next fitted a mixture of Gaussian distributions with the goal to capture both contribution of the variance of motion energy values during the lick as well as due to noise. The mixture of three Gaussian distributions worked well to fit the data across all sessions and mice. The threshold for the presence of movement was defined as the mean plus two standard deviations of the Gaussian with the lowest value of the mean from the Gaussian mixture.
Finally, to find the time of motion onset time, we looked backwards in time from the time of lick registration by the piezo signal. The time point preceding the first instance of motion energy going below the threshold value defined above was considered the onset time of the orofacial movement.
For histological identification of the location of the recording probes and allocation of unit location in the mouse brain, we followed a protocol similar to ref. 67 .
Serial 2-photon tomography for Neuropixels probe tract tracing
Following a terminal administration of pentobarbital, mice were perfused with a phosphate buffer solution (PBS) followed by 4% paraformaldehyde (PFA) solution. We post-fixed the brain in the 4% paraformaldehyde for a minimum of 24 h at approximately 5 C. Following fixation, brains were moved to PBS for a minimum of 12 h prior to imaging. For imaging, brains were embedded in 5% agarose gel and mounted onto a vibratome cutting stage under the microscope objective. The brains were imaged using serial section two-photon microscopy 68 . The microscope was controlled with ScanImage Basic (Vidrio Technologies), and custom software (BakingTray ( https://github.com/SainsburyWellcomeCentre/BakingTray )). Images were stitched into a full 3D rendering of the brain using custom software (StitchIt ( https://github.com/SainsburyWellcomeCentre/StitchIt )). We imaged the entire brain (from the olfactory bulb to the beginning of the spinal cord) with a resolution of x : approximately 2 μm, y : approximately 2 μm, z : 20 μm, with a 920 nm two-photon laser (100–150 mW power at sample). We sliced the brain in 40-μm sections, and imaged 2 z -planes (around 25 μm and around 45 μm from the tissue surface) into the remaining tissue following each 40-μm section. Two PMTs, one for capturing green (bandpass filter ET525/50 m) and red (bandpass filter ET570lp) fluorescence acquired the 2 channels of data subsequently used for analysis.
Neuropixels probe tract alignment to the Allen Common Coordinate Framework atlas and estimation of unit location
Prior to image processing, we downsampled microscopy images to 10-μm voxels and registered the brain to the standardized Allen Common Coordinate Framework (Allen CCF 69 ) using custom software (BrainRegister ( https://github.com/stevenjwest/brainregister )). We then manually traced each neuropixels probe tract through the brain in 3D using custom software (Lasagna ( https://github.com/SainsburyWellcomeCentre/lasagna )). Finally, we assessed the overall firing rates and LFP spectra of individual Neuropixels channels and compared it to atlas positions. Where needed, we manually adjusted the scaling of brain regions along the probe track to align responses on channels with features associated with anatomical locations using custom software (Ephys alignment tool ( https://github.com/int-brain-lab/iblapps/tree/master/atlaselectrophysiology ) 70 ). Unit location was estimated from the location of the channel that had the largest absolute peak value of the mean waveform. For all analyses, we combined units across all subdivisions of a brain region (layers of cerebral cortex, dorsal and ventral divisions as ACAd and ACAv and in some cases functionally similar brain regions—see Supplementary Tables 1 and 2 ).
Neural data analysis
Only brain regions with at least 40 units were analysed. Analyses specific to TF-responsive units were done only for brain regions with ≥10 such units. No further sample size calculations were performed. Manual curation of units’ quality and stability was done without the knowledge of brain regions from which recordings were made. The subsequent analyses pipeline was applied in the same manner to data from all applicable brain regions, but the custom nature of analyses prevented investigators to remain blind to the identity of brain regions or dataset type (trained versus naive mice).
GLM of neural activity
Model. We binned neural activity in 50-ms bins (matching the duration of each TF pulse) aligned to trial start. We then fitted a Poisson generalized linear model to predict trial-to-trial neural activity as a function of a set of temporally unfolded task-related predictors that were present during a trial. Each predictor was extended temporally prior and/or post the timing of the predictor in 50-ms discretized steps (matching neural activity binning), with an independent weight estimated for each time step around the predictor. We predicted neural activity using 19 task-related predictors:
(1) TF fluctuations during baseline period (kernel length: 0–1.5 s); (2) Trial start (0–1 s); (3) Time since baseline start (from 1 s from trial start to change onset); (4–9) Six change onsets (a separate predictor for each change size (0–2 s)); (10) Lick preparation (−1.25–0 s prior to lick); (11) Lick execution (0–0.5 s post lick); (12) Air-puff (0–0.25 s); (13) Reward (0–0.4); (14) Abort (−1.25–0.25); (15) Phase of grating for upwards drift (12 phase bins from 0–360°); (16) Phase of grating for downwards drift (12 phase bins from 0–360°); (17) Video motion energy (−0.05–0.8 s); (18) Running wheel movement (−0.05–0.8 s); (19) Pupil diameter (−0.75–0.75 s).
We fit the model with L2 (ridge) regularization, optimized with cyclical coordinate descent as implemented in GLMnet 71 ( α = 0). We trained a model for each neuron on 90% of the data, and cross-validated on 10% of the data, and iterated the predictions over a tenfold cross-validation. Within the training dataset we tuned the L2 regularization term using tenfold cross-validation.
Identification of units encoding TF, lick preparatory activity and/or lick execution activity. To identify which cells significantly responded to a predictor of interest (that is TF fluctuations during baseline, lick preparation epoch, or lick execution epoch), we first re-fitted reduced models similar to the full model on 90% of the data, with 10-fold cross-validation, except we removed a predictor(s) of interest: (1) For identification of TF-responsive units, we estimated a model where we removed the predictor estimating the responses to TF fluctuations during baseline. (2) For identification of units with lick preparation activity, we estimated a model where we removed the predictor estimating the activity leading up to a lick. (3) For identification of units responding to lick execution, we estimated a model where we removed the predictor estimating activity during lick execution, the predictor estimating activity modulation by motion energy captured by videography, and the predictor estimating activity modulation by running wheel movement.
For each 10% test set, for each neuron we then calculated the mean actual peri-event time histogram (PETH) as well as the mean predicted PETH of both the full model and the reduced model for the following types of events: (1) −0.15 to 0.75 s around fast and slow TF pulses (that is, TF values 0.5 s.d. from the mean TF during baseline); (2) −1.5 to 0 s prior to early lick onsets; and (3) 0 to 0.4 s post lick onset.
A unit was considered significantly encoding TF pulses during the baseline period if two criteria were satisfied: (1) The mean Pearson’s correlation prediction of the full model (across k-folds) from the combined mean fast and slow TF pulse response (that is, mean fast TF pulse and mean slow TF pulse responses subtracted from each other) was >0.2; and (2) if the cross-validated prediction of the TF response after subtracting the predicted TF response of the reduced model with no TF fluctuation predictor—that is, residual prediction—was significant ( P < 0.01 ( t -test), n = 10 independent cross-validations). A unit was considered significantly encoding lick preparation if (1) the mean Pearson’s correlation prediction of the full model (across k -folds) of the mean activity leading up to a lick (−1.25 to 0 s) was >0.2; and (2) if the cross-validated prediction of the mean activity after subtracting the predicted mean activity of the reduced model with no lick preparation kernel—that is, residual prediction—was significant ( P < 0.01 ( t -test), n = 10 independent cross-validations). Finally, a unit was considered significantly encoding lick execution if (1) the mean Pearson’s correlation prediction of the full model (across k -folds) of the mean activity following a lick (0 to 0.25 s) was >0.2; and (2) if the cross-validated prediction of the mean activity after subtracting the predicted mean activity of the reduced model with no lick preparation kernel—that is, residual prediction—was significant ( P < 0.01 ( t -test), n = 10 independent cross-validations).
Focality index. To assess how distributed TF encoding was across brain areas, before and after learning, we computed a focality index ( F ) (similar to Steinmetz et al. 3 ) of the TF encoding:
where p a is the proportion of neurons in an area that is encoding stimulus TF during the baseline period. If all TF encoding neurons were confined to a single area, this measure would take on the value of 1. If encoding was perfectly distributed across all areas recorded this measure would take on the value 1/ N areas . In order to compare between untrained and trained mice, we identified the common areas which had more than 40 units recorded in both trained and untrained mice. This left N = 24 areas from which to estimate the focality index. We estimated 95% confidence intervals and P values by bootstrapping the neurons included in the estimation 10,000 times with replacement.
Peak time and width of GLM estimated TF kernels for TF-responsive neurons. To investigate the peak time and width of the GLM estimated TF kernel for assessing how sustained responses to TF fluctuations were based on GLM weights, we first identified the absolute peak value of the TF kernel; because the GLM was based on 50 ms binning of spike counts, peak times for the GLM TF kernel was in 50 ms resolution. In cases where the absolute peak position within 1 s was a negative weight, we flipped the kernel in order to calculate the width. We then estimated the full width at half maximum (FWHM) of each TF kernel around its peak using findpeaks in MATLAB. For each area, we calculated the median peak time and median FWHM across all TF-responsive units.
Ramping differences in GLM change kernels. To test how neurons accumulated evidence when they were presented with a rewarding sustained change in stimulus speed, we tested how the slope of the visual evoked ramping activity following a change onset was dependent on the amount of evidence (change size) being presented. To isolate the visual component of the activity following change onset, we used the GLM kernel which fits the activity following change onset until change offset, while linearly taking into account other variables which may contribute to activity such as pupil size, preparatory activity and movement-related activity (see Model).
We estimated the mean change kernel for each change size for TF-responsive and non-TF-responsive units separately for each area. In cases where responses to fast TF pulses were negative, we flipped the change kernel so every unit had responses aligned to positive fast TF pulses—this allowed the mean to capture the visual evidence activity ramp irrespective of sign. We then identified the time point for each change size where the change kernel reached 50% of its maximum weight (To control for noise fluctuations in kernel weights, we approximated the 50th percentile crossing by taking the mean time point of the 33.33rd percentile, 50th percentile and 66.66th percentile crossing). We then calculated the degree to which activity ramping time scaled with change size, by regressing the 50th percentile crossing against change size. We estimated the non-parametric 95% confidence intervals and P values of the relationship between change size and 50th percentile crossing (that is, ramping time/change size) by bootstrapping with replacement (10,000 times) the neurons went into the mean change kernels, and then estimating the slope of the regression for each bootstrapped mean change kernels.
Propagation and widening of TF pulse evoked activity
Identification of TF pulse outlier events. Fast TF pulse was defined as TF fluctuations larger than 1 s.d. of baseline TF fluctuations (in log 2 scale) above the mean TF value (TF > 1.19 Hz). Similarly, slow TF pulse was defined as TF fluctuations below 0.84 Hz.
For calculation of average response to TF outlier events, we considered only TF outlier events satisfying the following criteria:
Later than 1 s from the baseline onset.
Earlier than 2 s + post pulse analysis window from the motion onset time on early lick or abort trials.
Excluding the change period plus a post pulse analysis window.
The aim of these criteria was to exclude the influence of baseline onset, movement, or preparatory activity on the response to TF pulses.
Estimation of peak time and width of TF pulse evoked activity. For each unit defined as TF-responsive by the GLM analysis described above, we calculated a mean response to a fast pulse using outlier events that occurred during the baseline period and satisfied the criteria outlined above. Additionally, we calculated a mean response to TF pulses within [−0.5, 0.5] s.d. of the baseline TF fluctuations. The goal of this procedure was to capture continuous ramps of activity that some units exhibited and exclude their influence on the shape of response to a TF pulse. We applied the subtraction of this baseline response for all TF pulse response analysis unless explicitly stated.
Next, for the baseline subtracted mean response to a fast TF pulse, we calculated its peak time, as the time of the largest absolute change in firing rate within 1 s from the pulse onset, and a corresponding half-peak width.
Integration of multiple TF pulses. Because the noise in TF fluctuations is random, by chance there are occurrences of two fast pulses separated by a certain delay. To study the integration of TF pulses, we found such instances of events where two fast pulses occurred at a given delay between the offset of the first and the onset of the second, additionally also satisfying the exclusion criteria outlined above. The mean response aligned to such events was considered a response to a sequence of two fast pulses.
For computing the mean response across all TF-responsive units within a brain region, in order to avoid averaging across responses with different signs, we flipped the sign of response for units that showed decreases in activity after a single fast pulse. For computing a z -score of response, the mean and s.d. were estimated from 0.5 s preceding the first pulse onset.
Facilitation by the second fast pulse. First, we measured an average of z -scored responses across the population of TF-responsive units within a brain region to a single fast TF pulse. We then computed the peak value of that response ( r 1fast ), and a corresponding peak time. To find the size of response to a sequence of two fast pulses ( r 2fast ), we found a time point at the same delay from the onset of the second fast pulse as the peak time of response to a single fast pulse and found a peak value of response within 100 ms centred around that time point. The relative facilitation to a sequence of fast pulses was defined as \(\varDelta \,=\,\frac{{r}_{2{\rm{fast}}}\,-\,{r}_{1{\rm{fast}}}}{{r}_{1{\rm{fast}}}}\) .
To determine the confidence intervals for the results of this analysis, we bootstrapped with replacement (2,000 times) across TF-responsive neurons and repeated the analysis described above for each sample of neurons. Shaded regions indicate 2.5 and 97.5 percentiles of the resulting distribution.
Preparatory activity before the lick onset
To study change-aligned (Fig. 3 ) or hit lick-aligned (Fig. 5 ) activity, we computed z -score of mean PETH for each unit. z -Scoring was done using the mean and s.d. estimated from activity during 2 s before the change onset.
For analysis shown on Fig. 5 , for each brain region the fraction of significantly active units within a group (that is, TF-responsive) was measured by calculating at every time point a fraction of units with the absolute value of z -score larger than the significance threshold of 2.576 (corresponding to P < 0.01). Additionally, we subtracted the ‘baseline’ level of activity calculated within [−2, −1.8] s before hit-lick onset, which for a few brain regions was larger than chance level likely due to non-normal distribution of firing rates or a small number of events used for estimation of the mean and s.d. The confidence intervals were estimated by bootstrapping with replacement (5,000 times) across TF-responsive (or TF non-responsive) neurons and repeating the estimation of fraction of significantly active neurons for each sample of neurons.
The latency of activation of TF-responsive or TF non-responsive populations was defined as the earliest time point following which within a 100-ms window for at least 80 ms: (1) the lower 95% confidence interval of fraction of active units was above zero; and (2) the mean fraction of active units was above 0.1.
The latency of significant difference in activation between TF-responsive and TF non-responsive populations was estimated as the first time point where within a 100-ms window for at least 80 ms the confidence intervals of the difference in activation were above zero.
The latency of significant difference in activation across all units in each brain region (Extended Data Fig. 9a ) was estimated as the first time point where within a 100-ms window: (1) the lower 95% confidence interval of fraction of active units was above zero; and (2) the mean fraction of active units was above 0.05.
Intrinsic timescales
We binned the neural activity into 50-ms bins (same binning was used in ref. 23 ). We then calculated the temporal autocorrelation (20 lags = 1 s) of spike counts using Pearson’s correlation in the inter-trial intervals between −2.5 s to −0.5 s prior to trial onset for each neuron (in this period mice were seeing a grey screen, and trained mice had to remain stationary for at least 3 s for the trial to begin).
To determine the intrinsic timescale for each area, we fit an exponential decay function to the mean autocorrelation function of all the units recorded in the area. For single-neuron analysis of relationship between intrinsic timescales and TF width, we estimated the autocorrelation for each TF-responsive neuron separately. For areas or neurons with autocorrelation functions with non-monotonic decay, we fit the exponential decay from the part of the autocorrelation where monotonic decay was happening (in a subset of areas this would mean offsetting the fit 1–3 time bins). Finally, we calculated the τ (that is, the intrinsic timescale value) of the exponential decay (accounting for offset where necessary).
Population analysis
Similarity of tf pulse responses and lick preparation activity in tf-responsive populations.
We assessed the similarity of TF responses and lick preparation activity across TF-responsive populations in each area by estimating the Pearson’s correlation of mean firing rates (within a 50-ms window around the mean activity peak time across neurons within the each area) following fast TF pulses (that is, >1 s.d. TF value) and their mean activity prior to early lick [−0.3 to 0 s] (after normalizing firing rates by subtracting baseline firing rates from both TF responses and lick preparation activity). We estimated the non-parametric 95% confidence intervals and P values by bootstrapping with replacement (10,000 times) the neurons going into the correlation.
Pre-processing steps
For all units located within a given brain region, but not necessary simultaneously recorded, we first computed the mean neural responses across a given trial type (for results shown on Fig. 6b–h : hit trials during weak TF changes (1.25 and 1.35 Hz) aligned to the lick onset times, [−2, 1.5] s time window). Only trials with hit-lick onset times larger or equal 0.4 s from change onset were used. Neurons from sessions with less than 10 trials of a given type were excluded from this analysis. Firing rates were calculated as spike counts averaged in 10 ms bins and smoothened by convolution with two-sided Gaussian with 30 ms s.d. The mean neural responses were combined into a firing rate matrix (but also see cross-validation section) with dimensions of Neurons × Time.
Neural data was pre-processed in the following way: first, to limit the dominant influence of high-firing units, we applied soft-normalization to each neuron’s firing rate, such that the neurons with strong responses had close to unity range of responses \({r}^{/}\,=\,\frac{r}{7+(\max (r)-\min (r))}\) . The constant 7 was chosen as the roughly 20th percentile value of the firing rate range across all units. Second, the neural responses were mean-centred by subtracting the mean of each neuron’s activity across time and the mean activity across all neurons at every time point.
Definitions of movement and movement-null subspaces
We used the approach first utilized in ref. 33 . There, the authors formalized a method to find a linear mapping between low-dimensional representation of activity in PMd/M1 and the muscles EMG data, which defines a movement subspace. A null-space relative to that subspace forms an orthogonal set of dimensions which activity can occupy without directly affecting the movement execution. To extend this analysis on our data, we used combined recordings of orofacial motor and premotor nuclei (V, IRN, SPVI and SPVO) as a proxy for activity of orofacial muscles involved in execution of a lick. While recordings from GRN could have also been included into this group, we kept it separate to allow the population analysis to be applied to that region because (1) we had a large number of units recorded from that region alone; and (2) it was the only nucleus in medulla with above-chance number of TF-responsive units, warranting a separate analysis.
We considered a possible mapping onto the movement subspace for each brain region. Our rationale was the following: there exist several parallel neural pathways that can drive the activity of orofacial nuclei neurons–from primary motor cortex, basal ganglia, cerebellar or midbrain output regions 56 , 57 , 72 . Thus, the modes of activity within these regions that map onto the movement subspace may have a causal role for the execution of licks. In general, however, these signals can also be caused by movement afference that is broadcasted globally 3 , 5 , 9 (Fig. 1k,l ). It is impossible to differentiate between these two possibilities from our data alone and thus the existence of mapping of activity onto the movement subspace does not necessarily imply that the brain region is causally involved in execution of the lick. With that said, we did not find a good mapping onto a movement subspace for most of the early visual areas, olfactory regions and hippocampal input regions (Extended Data Fig. 10a ), suggesting that existence of mapping onto the movement subspace is not possible across all brain regions.
The mapping onto movement subspace was defined as:
where \(\widetilde{M}\) and \(\widetilde{N}\) are low-dimensional representations of activity (projections onto main the principal components, the latter found via svd Matlab function) of neurons within the orofacial nuclei group and the target brain region, respectively, and W is a linear mapping operator onto the movement subspace.
Before finding a linear mapping, we also zeroed the initial state across projections on principal components by subtracting from each projection the mean value within [−2, −1.5] s from lick onset. This step avoided the need for using intercept in the linear fit and simplified the visualization of projections on principal components and movement/movement-null dimensions. Linear mapping was found using only the time-period containing movement-related activity of orofacial nuclei [−0.1, 1.5] s around lick onset. This way we did not preclude the presence of preparatory activity on movement dimensions from the definition of the linear fit itself. A linear mapping to movement dimensions was found using linear regression with the Matlab function lsqnonlin.
Correspondingly, W null was a null-space of W and was found using the Matlab function null. We used two top principal components of orofacial nuclei activity (which captured 61% of the total cross-validated variance; Extended Data Fig. 10a,b ) and 4 top principal components of activity in a target brain region to find W and W null operators (see Extended Data Fig. 10b–d ). This choice resulted in both movement and movement-null subspaces being two-dimensional. We additionally ensured that norms of these operator are equal \(\parallel {W}_{{\rm{null}}}\parallel =\parallel W\parallel \) in order to make the comparison between the movement and movement-null subspaces fair.
Since the definition of specific dimensions in movement-null subspace is to a degree arbitrary, we defined the first movement-null dimension by finding a rotation within the movement-null subspace that maximized the amount of variance captured by that dimension prior to lick onset. The second movement-null dimension was then simply orthogonal to the first dimension in movement-null subspace. This was used mainly to simplify visualization, with all subspace-related analyses done using both dimensions in each subspace.
The positive direction of movement dimensions was chosen such that the mean value of projection of orofacial nuclei activity within [−2, 0.5] s around lick onset was positive. The positive direction for movement-null dimensions was chosen such that the mean value of projection of activity within [−2, 0] s around lick onset was positive.
Subspace occupancy
Relative subspace occupancy at a moment of time t was defined as
where E null ( t ) and E m ( t ) are Euclidean distances within movement-null and movement subspaces, measured between the neural state at the current moment of time t and the initial time point (the mean across 2 and 1.5 s before the lick onset). Values close to zero signify equal occupancy between subspaces and positive values indicate a preferential occupancy of the movement-null subspace. The peak-normalized occupancy (Extended Data Fig. 11a,b ) was defined as \(O(t)=\frac{E(t)}{\max (E)}\)
Decomposition of projections onto contributions from TF-responsive and TF non-responsive units
We decomposed the projections on main principal components into a sum of contributions from TF-responsive and the TF non-responsive units. For that, we used the knowledge of identity of each unit as TF-responsive or TF non-responsive and wrote down the principal components U (from the singular value decomposition (SVD) of the firing rate matrix N = USV T ) as a sum of two parts as:
where U i is a loading of the i th unit.
With that, projections on principal components can be written as:
Substituting equation ( 3 ) into equation ( 1 ) gives projections onto movement dimensions as:
and, correspondingly, projections on movement-null dimensions are written as:
The relative contribution of TF-responsive units within movement and movement-null subspaces at the moment of time t was then defined as following:
where the second multiplicative term ensures that the sign of contribution is relative to the defined positive direction (see above) of dimensions within each subspace.
In order to test whether the contribution of TF-responsive units is larger than what is expected from a uniform contribution of the full population, we repeatedly randomly selected (2,000 times) the same number of units as there were TF-responsive ones from the whole population and computed their contribution to projections on movement and movement-null dimensions as described above.
In addition to the analysis described above, we have also checked whether the above-chance contribution of TF-responsive units is a consequence of their level of activity, despite the normalization method that we used, or does it reflect a better correspondence of their activity to the population modes of activity within the movement-null subspace. For that we looked at the distribution of loadings along the first movement-null dimension–that captured the majority of preparatory activity there. We found that the majority of brain regions where TF-responsive units had above-chance contribution to the preparatory activity also had larger absolute values of loadings along that dimension than the rest of the population (Extended Data Fig. 11d,e ).
Cross-validation
Since our analyses were focused on characterizing the mean neural responses, the cross-validation procedure that we used was designed to test the stability of the mean neural responses and their corresponding low-dimensional representations across trials. For that, we split trials into two randomly assigned and equally sized groups (fit and test trials) and calculated the mean neural response per unit across each group of trials. We next combined firing rates of neurons from the same brain region(s) (but not necessarily simultaneously recorded) into a joint matrix. After applying the pre-processing steps outlined above, we had two firing rate matrices from fit and test trials.
For cross-validated PCA (Extended Data Fig. 10a ), we applied SVD on the first (fit) matrix and measured how well the remaining (test) matrix is predicted by the reconstruction from SVD components found from the first matrix. Similarly, the projections of activity on main principal components (Extended Data Fig. 13 ) were done using the test data, projected onto principal components found from the fit data.
For further analyses utilizing movement and movement-null subspaces, we applied SVD separately on each matrix and found their projections on first four main principal components. We then used low-dimensional representation of fit trials data to find linear mapping W and W null onto the movement and moment-null subspaces. Finally, we applied W and W null found from the fit data to the low-dimensional representation of the test data. This procedure was repeated 2,000 times, the 95% confidence intervals shown in Fig. 6 illustrate the 2.5 and 97.5 percentiles across projections of the test data. Because the sign of projection is arbitrary defined, we additionally applied a potential flipping of the sign of eigenvectors from each draw based on which direction had better alignment with the eigenvectors computed from the full firing rate matrix without the split into fit and test trials.
Responses to TF pulses
For each brain region, we constructed a firing rate matrix of all units responses to a fast TF pulse (or concatenating in time responses of each unit to different types of TF pulses for analysis shown in Fig. 6i,k–m ), and used the same pre-processing steps as described above. The projections onto the movement and movement-null dimensions were done using loadings found from the analysis of hit licks activity described above (using the full firing rate matrix of hit-lick responses without the split into fit and test trials). Cross-validation of consistency of projections was done by randomly selecting half of TF outlier events, computing the mean firing rate across those events for each unit, applying the steps above to find the projections, and repeating this procedure 2,000 times. For analyses where different brain regions were combined into a common group, all units from those brain regions were combined into a joined firing rate matrix and the steps described above were applied.
Alignment of fast TF pulse response with a given dimension in movement or movement-null subspace was calculated as a cosine of an angle between the projection onto a target dimension and a 4-dimensional vector of TF pulse response (2 movement and 2 movement-null dimensions) at a time of the maximum Euclidean distance from the initial state across 4 dimensions within a 0.75-s window from the pulse onset. Similarly, for calculating the scaling of responses to different TF pulses along the first movement-null dimension, we found the sizes of projections at times of maximal Euclidean distance from the initial state within a 0.75-s window from the first TF pulse onset.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The data that support the findings of this study are available from the corresponding authors upon reasonable request. Source data are provided with this paper.
Code availability
Custom acquisition, post-processing and analysis code is available at https://github.com/BaselLaserMouse/Khilkevich_Lohse_2024 .
Gold, J. I. & Shadlen, M. N. The neural basis of decision making. Annu. Rev. Neurosci. 30 , 535–574 (2007).
Article CAS PubMed Google Scholar
Hanks, T. D. & Summerfield, C. Perceptual decision making in rodents, monkeys, and humans. Neuron 93 , 15–31 (2017).
Steinmetz, N. A., Zatka-Haas, P., Carandini, M. & Harris, K. D. Distributed coding of choice, action and engagement across the mouse brain. Nature 576 , 266–273 (2019).
Article CAS PubMed PubMed Central Google Scholar
Chen, S. et al. Brain-wide neural activity underlying memory-guided movement. Cell 187 , 676–691 (2024).
Allen, W. E. et al. Thirst regulates motivated behavior through modulation of brainwide neural population dynamics. Science 364 , 253 (2019).
Article ADS PubMed PubMed Central Google Scholar
Musall, S., Kaufman, M. T., Juavinett, A. L., Gluf, S. & Churchland, A. K. Single-trial neural dynamics are dominated by richly varied movements. Nat. Neurosci. 22 , 1677–1686 (2019).
International Brain Laboratory. A brain-wide map of neural activity during complex behaviour. Preprint at bioRxiv https://doi.org/10.1101/2023.07.04.547681 (2023).
Pinto, L., Tank, D. W. & Brody, C. D. Multiple timescales of sensory-evidence accumulation across the dorsal cortex. eLife 11 , e70263 (2022).
Stringer, C. et al. Spontaneous behaviors drive multidimensional, brainwide activity. Science 364 , 255 (2019).
Siegel, M., Buschman, T. J. & Miller, E. K. Cortical information flow during flexible sensorimotor decisions. Science 348 , 1352–1355 (2015).
Article ADS CAS PubMed PubMed Central Google Scholar
Inagaki, H. K. et al. Neural algorithms and circuits for motor planning. Annu. Rev. Neurosci. 45 , 249–271 (2022).
Deverett, B., Koay, S.A., Oostland, M. & Wang, S. H.-H. Cerebellar involvement in an evidence-accumulation decision-making task. eLife 7 , e36781 (2018).
Article PubMed PubMed Central Google Scholar
Mante, V., Sussillo, D., Shenoy, K. V. & Newsome, W. T. Context-dependent computation by recurrent dynamics in prefrontal cortex. Nature 503 , 78–84 (2013).
Orsolic, I., Rio, M., Mrsic-Flogel, T. D. & Znamenskiy, P. Mesoscale cortical dynamics reflect the interaction of sensory evidence and temporal expectation during perceptual decision-making. Neuron 109 , 1861–1875.e10 (2021).
Shadlen, M. N. & Newsome, W. T. Neural basis of a perceptual decision in the parietal cortex (area LIP) of the rhesus monkey. J. Neurophysiol. 86 , 1916–1936 (2001).
Huk, A. C. & Shadlen, M. N. Neural activity in macaque parietal cortex reflects temporal integration of visual motion signals during perceptual decision making. J. Neurosci. 25 , 10420–10436 (2005).
Yao, J. D., Gimoto, J., Constantinople, C. M. & Sanes, D. H. Parietal cortex is required for the integration of acoustic evidence. Curr. Biol. 30 , 3293–3303.e4 (2020).
Hanks, T. D. et al. Distinct relationships of parietal and prefrontal cortices to evidence accumulation. Nature 520 , 220–223 (2015).
Ding, L. & Gold, J. I. Caudate encodes multiple computations for perceptual decisions. J. Neurosci. 30 , 15747–15759 (2010).
Yartsev, M. M., Hanks, T. D., Yoon, A. M. & Brody, C. D. Causal contribution and dynamical encoding in the striatum during evidence accumulation. eLife 7 , e34929 (2018).
Bolkan, S. S. et al. Opponent control of behavior by dorsomedial striatal pathways depends on task demands and internal state. Nat Neurosci 25 , 345–357 (2022).
Gold, J. I. & Shadlen, M. N. Representation of a perceptual decision in developing oculomotor commands. Nature 404 , 390–394 (2000).
Article ADS CAS PubMed Google Scholar
Murray, J. D. et al. A hierarchy of intrinsic timescales across primate cortex. Nat Neurosci 17 , 1661–1663 (2014).
Runyan, C. A., Piasini, E., Panzeri, S. & Harvey, C. D. Distinct timescales of population coding across cortex. Nature 548 , 92–96 (2017).
Cavanagh, S. E., Hunt, L. T. & Kennerley, S. W. A diversity of intrinsic timescales underlie neural computations. Front. Neural Circuits https://doi.org/10.3389/fncir.2020.615626 (2020).
Ossmy, O. et al. The timescale of perceptual evidence integration can be adapted to the environment. Curr. Biol. 23 , 981–986 (2013).
Tanji, J. & Evarts, E. V. Anticipatory activity of motor cortex neurons in relation to direction of an intended movement. J. Neurophysiol. 39 , 1062–1068 (1976).
Guo, Z. V. et al. Maintenance of persistent activity in a frontal thalamocortical loop. Nature 545 , 181–186 (2017).
Chabrol, F. P., Blot, A. & Mrsic-Flogel, T. D. Cerebellar contribution to preparatory activity in motor neocortex. Neuron 103 , 506–519.e4 (2019).
Weinrich, M., Wise, S. P. & Mauritz, K.-H. A neurophysiological study of the premotor cortex in rhesus monkey. Brain 107 , 385–414 (1984).
Article PubMed Google Scholar
Kornhuber, H. H. & Deecke, L. Hirnpotentialänderungen bei Willkürbewegungen und passiven Bewegungen des Menschen: Bereitschaftspotential u. reafferente Potentiale. Pflugers Arch. Gesamte Physiol. Menschen Tiere 284 , 1–17 (1965).
Wu, Z. et al. Context-dependent decision making in a premotor circuit. Neuron 106 , 316–328.e6 (2020).
Kaufman, M. T., Churchland, M. M., Ryu, S. I. & Shenoy, K. V. Cortical activity in the null space: permitting preparation without movement. Nat. Neurosci. 17 , 440–448 (2014).
Elsayed, G. F., Lara, A. H., Kaufman, M. T., Churchland, M. M. & Cunningham, J. P. Reorganization between preparatory and movement population responses in motor cortex. Nat. Commun. 7 , 13239 (2016).
Inagaki, H. K. et al. A midbrain–thalamus–cortex circuit reorganizes cortical dynamics to initiate movement. Cell 185 , 1065–1081.e23 (2022).
Darlington, T. R. & Lisberger, S. G. Mechanisms that allow cortical preparatory activity without inappropriate movement. eLife 9 , e50962 (2020).
Roitman, J. D. & Shadlen, M. N. Response of neurons in the lateral intraparietal area during a combined visual discrimination reaction time task. J. Neurosci. 22 , 9475–9489 (2002).
Britten, K. H., Newsome, W. T., Shadlen, M. N., Celebrini, S. & Movshon, J. A. A relationship between behavioral choice and the visual responses of neurons in macaque MT. Vis. Neurosci. 13 , 87–100 (1996).
Boyd-Meredith, J. T., Piet, A. T., Dennis, E. J., El Hady, A. & Brody, C. D. Stable choice coding in rat frontal orienting fields across model-predicted changes of mind. Nat. Commun. 13 , 3235 (2022).
Bennur, S. & Gold, J. I. Distinct representations of a perceptual decision and the associated oculomotor plan in the monkey lateral intraparietal area. J. Neurosci. 31 , 913–921 (2011).
Fitzgerald, J. K., Freedman, D. J. & Assad, J. A. Generalized associative representations in parietal cortex. Nat. Neurosci. 14 , 1075–1079 (2011).
Stine, G. M., Trautmann, E. M., Jeurissen, D. & Shadlen, M. N. A neural mechanism for terminating decisions. Neuron 111 , 2601–2613.e5 (2023).
Ding, L. & Gold, J. I. Neural correlates of perceptual decision making before, during, and after decision commitment in monkey frontal eye field. Cereb. Cortex 22 , 1052–1067 (2012).
Duan, C. A. et al. A cortico-collicular pathway for motor planning in a memory-dependent perceptual decision task. Nat. Commun. 12 , 2727 (2021).
Newsome, W. T. & Park, E. B. A selective impairment of motion perception following lesions of the middle temporal visual area (MT). J. Neurosci. 8 , 2201–2211 (1988).
Mountcastle, V. B., Steinmetz, M. A. & Romoa, R. Frequency discrimination in the sense of flutter: psychophysical measurements correlated with postcentral events in behaving monkeys. J. Neurosci. 10 , 3032–3044 (1990).
Romo, R. & de Lafuente, V. Conversion of sensory signals into perceptual decisions. Prog. Neurobiol. 103 , 41–75 (2013).
Brunton, B. W., Botvinick, M. M. & WangBrody, C. D. Rats and humans can optimally accumulate evidence for decision-making. Science 340 , 95–97 (2013).
Pinto, L. et al. An accumulation-of-evidence task using visual pulses for mice navigating in virtual reality. Front. Behav. Neurosci. 12 , 36 (2018).
Akrami, A., Kopec, C. D., Diamond, M. E. & Brody, C. D. Posterior parietal cortex represents sensory history and mediates its effects on behaviour. Nature 554 , 368–372 (2018).
Jun, J. J. et al. Fully integrated silicon probes for high-density recording of neural activity. Nature 551 , 232–236 (2017).
Ruesseler, M., Weber, L. A., Marshall, T. R., O’Reilly, J. & Hunt, L. T. Quantifying decision-making in dynamic, continuously evolving environments. eLife 12 , e82823 (2023).
Gao, Z. et al. A cortico-cerebellar loop for motor planning. Nature 563 , 113–116 (2018).
Wang, Y. et al. A cortico-basal ganglia-thalamo-cortical channel underlying short-term memory. Neuron 109 , 3486–3499.e7 (2021).
Park, I. M., Meister, M. L. R., Huk, A. C. & Pillow, J. W. Encoding and decoding in parietal cortex during sensorimotor decision-making. Nat. Neurosci. 17 , 1395–1403 (2014).
Takatoh, J. et al. Constructing an adult orofacial premotor atlas in Allen mouse CCF. eLife 10 , e67291 (2021).
Guo, H. et al. Whole-brain monosynaptic inputs to hypoglossal motor neurons in mice. Neurosci. Bull. 36 , 585–597 (2020).
Lemke, S. M., Ramanathan, D. S., Guo, L., Won, S. J. & Ganguly, K. Emergent modular neural control drives coordinated motor actions. Nat. Neurosci. 22 , 1122–1131 (2019).
Xiong, Q., Znamenskiy, P. & Zador, A. M. Selective corticostriatal plasticity during acquisition of an auditory discrimination task. Nature 521 , 348–351 (2015).
Peters, A. J., Fabre, J. M. J., Steinmetz, N. A., Harris, K. D. & Carandini, M. Striatal activity topographically reflects cortical activity. Nature 591 , 420–425 (2021).
Piet, A. T., El Hady, A. & Brody, C. D. Rats adopt the optimal timescale for evidence integration in a dynamic environment. Nat. Commun. 9 , 4265 (2018).
Herzfeld, D. J., Kojima, Y., Soetedjo, R. & Shadmehr, R. Encoding of error and learning to correct that error by the Purkinje cells of the cerebellum. Nat. Neurosci. 21 , 736–743 (2018).
Churchland, M. M., Cunningham, J. P., Kaufman, M. T., Ryu, S. I. & Shenoy, K. V. Cortical preparatory activity: representation of movement or first cog in a dynamical machine? Neuron 68 , 387–400 (2010).
Aoi, M. C., Mante, V. & Pillow, J. W. Prefrontal cortex exhibits multidimensional dynamic encoding during decision-making. Nat. Neurosci. 23 , 1410–1420 (2020).
Pachitariu, M., Steinmetz, N., Kadir, S., Carandini, M. & Harris, K. D. Kilosort: realtime spike-sorting for extracellular electrophysiology with hundreds of channels. Preprint at bioRxiv https://doi.org/10.1101/061481 (2016).
Mathis, A. et al. DeepLabCut: markerless pose estimation of user-defined body parts with deep learning. Nat. Neurosci. 21 , 1281–1289 (2018).
International Brain Laboratory. Reproducibility of in-vivo electrophysiological measurements in mice. Preprint at bioRxiv https://doi.org/10.1101/2022.05.09.491042 (2022).
Amato, S. P., Pan, F., Schwartz, J. & Ragan, T. M. Whole brain imaging with serial two-photon tomography. Front. Neuroanat. 10 , 31 (2016).
Wang, Q. et al. The Allen Mouse Brain Common Coordinate Framework: a 3D reference atlas. Cell 181 , 936–953.e20 (2020).
Liu, L. D. et al. Accurate localization of linear probe electrode arrays across multiple brains. eNeuro https://doi.org/10.1523/ENEURO.0241-21.2021 (2021).
Friedman, J., Hastie, T. & Tibshirani, R. Regularization paths for generalized linear models via coordinate descent. J. Stat. Softw. 33 , 1–22 (2010).
Dempsey, B. et al. A medullary centre for lapping in mice. Nat. Commun. 12 , 6307 (2021).
Download references
Acknowledgements
The authors thank M. Hamada, D. Gupta and S. Hofer for comments on the manuscript; M. Sahani for helpful discussions regarding population analyses; R. Campbell for help with histology, microscopy and image processing; M. Faulkner for providing access to ephys-atlas alignment tools implemented by the International Brain Laboratory; the staff at the SWC Neurobiological Research Facility for mouse husbandry; the SWC Fabrication Laboratory for help with machining; M. Skretowska for help with training mice; members of the Mrsic-Flogel, Hofer and other SWC laboratories for insightful discussion and advice. This work was supported by Wellcome awards to T.D.M.-F. (217211/Z/19/Z) and M.L. (224121/Z/21/Z), by the Sainsbury Wellcome Centre’s core provided by Wellcome (219627/Z/19/Z) and the Gatsby Charitable Foundation (GAT3755).
Author information
These authors contributed equally: Andrei Khilkevich, Michael Lohse
Authors and Affiliations
Sainsbury Wellcome Centre for Neural Circuits and Behaviour, University College London, London, UK
Andrei Khilkevich, Michael Lohse, Ryan Low, Ivana Orsolic, Tadej Bozic, Paige Windmill & Thomas D. Mrsic-Flogel
You can also search for this author in PubMed Google Scholar
Contributions
A.K., M.L., I.O. and T.D.M.-F. conceived the project. A.K., M.L. and T.D.M.-F. designed the experiments. A.K. and M.L. collected and analysed the data with supervision from T.D.M.-F. T.B. trained the mice with guidance from A.K., M.L. and I.O. R.L. developed the outlier detection agent for decision-making. P.W. manually curated spike-sorted extracellular data with guidance from A.K. and M.L. A.K., M.L. and T.D.M.-F. wrote the paper with input from all authors.
Corresponding authors
Correspondence to Andrei Khilkevich , Michael Lohse or Thomas D. Mrsic-Flogel .
Ethics declarations
Competing interests.
The authors declare no competing interests
Peer review
Peer review information.
Nature thanks the anonymous reviewer(s) for their contribution to the peer review of this work. Peer review reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended data fig. 1 summary of recordings in trained mice..
a , Number of cells recorded from trained mice in each Allen Brain Atlas designated region. b-f , Locations of all well-isolated and stable units, shown within a 3D rendering of Allen Common Coordinate Framework from five perspectives.
Extended Data Fig. 2 GLM Performance.
a , Schematic of Poisson GLM. b , Cross-validated model prediction performance of single trial spike counts with full GLM model ( r ). c , Cross-validated model prediction performance of mean PSTH following fast and slow pulses ( r ). d , Cross-validated model prediction performance of mean PSTH leading up to an early lick (Lick preparation) ( r ). e , Cross-validated model prediction performance of mean PSTH after early lick (Lick execution) ( r ). f , GLM predictions on example neuron recorded in MOs. Top: GLM kernels which the predictions are made from. Bottom: Real vs full GLM predicted vs reduced GLM (without key predictor in model) PSTHs. g , GLM predictions on example neuron recorded in SCs. Top: GLM kernels which the predictions are made from. Bottom: Real vs full GLM predicted vs reduced GLM (without key predictor in model) PSTHs. h , Mean TF kernels across all areas with 10 or more TF-responsive units recorded (for averaging kernels are flipped when needed to always have a positive response). i , Mean lick preparation and lick execution kernels across all areas with 10 or more lick preparation neurons responsive units recorded (for averaging kernels are flipped when needed to always have a positive response).
Extended Data Fig. 3 Encoding of temporal frequency fluctuations, lick preparation and lick execution across brain areas.
a-c , Percentage of units encoding temporal frequency fluctuations during baseline, lick preparation, or lick execution in major area groupings with 95% binomial confidence intervals. a , Percentage lick execution units: All areas: p < 0.001 (Binomial test). b , Percentage lick preparation units: Early visual, Higher visual, Basal ganglia, Frontal cortex, Olfactory nuclei (OLF), Thalamus, Midbrain, Hippocampus, Cerebellum, Lateral hypothalamus (LHA), GRN (Medulla*), Medulla: p < 0.001 (Binomial test), Medulla: p < 0.01 (Binomial test). c , Percentage TF Responsive units: Early visual, Higher visual, Basal ganglia, Frontal cortex, Thalamus, Midbrain, Hippocampus, Cerebellum, GRN (Medulla*): p < 0.001 (Binomial test), Olfactory nuclei (OLF), Lateral hypothalamus (LHA), and Medulla: p > 0.05 (Binomial test). Error bars in panels a-c are 95% binomial confidence intervals. Red areas designate chance level. See Supplementary Table 1 for n of each brain area grouping. d , Percentage overlap of encoding (estimated from GLM) of TF, lick preparation, and lick execution, in all areas with more than 40 units recorded. y-axis is the source population (i.e., all TF responsive neurons, all lick preparation neurons, or all lick execution neurons).
Extended Data Fig. 4 Responses of TF responsive neurons across the brain to fast or slow TF pulses and early licks.
Activity (z-scored) of individual neurons around fast TF pulses ( left ), slow TF pulses ( middle ) and early licks ( right ) for all TF responsive units from all areas with 10 or more TF responsive units recorded. Major subdivisions of the brain grouped by colour. Each line represents one neuron.
Extended Data Fig. 5 Properties of responses to a single fast TF pulse from PSTHs and GLM + Relative facilitation by the second fast TF pulse as a function of delay from the first one.
a-d , Comparison of peak time and response width of PSTHs following a fast TF pulse vs GLM TF kernels. a , Median peak time of response to a fast TF pulse estimated from PSTH (red) and median peak time of GLM TF kernel (blue), shown for each brain region. b , Correlation across brain regions between median peak time estimated from PSTH and median peak time of GLM TF kernel. c-d , Same as a-b, but for fast TF pulse response half-peak width. e , Relative facilitation by the second fast TF pulse, normalized by the response to a single fast TF pulse, shown as a function of delay between two fast TF pulses for each brain region with at least 10 TF responsive units (mean and 95% confidence intervals, bootstrap test (see Methods )). Values close to zero imply no facilitation (same size of response to the second fast TF pulse as to the first one), while values close to 100% imply doubling of the response size.
Extended Data Fig. 6 Effect of magnitude and timing of TF pulses on probability of early licks.
a , Mean performance (psychometric curves) for mice data (dashed black line, n = 15 mice) and outlier detection agent (purple). b , Mean reaction times per change magnitude for outlier detection agent (purple) and mice data (dashed black line, n = 15 mice). Error bars indicate 95% confidence intervals across 4000 synthetic datasets of the model (see Methods ). c , Conditional probability of early lick at a specific time after a TF pulse of given magnitude. Here and later early lick probability is shown relative to the probability at the mean baseline TF (1 Hz). d , Probability of early lick after a TF pulse of given magnitude (here and later cumulatively within [0.2, 1] s window). Mice data is shown in black, outlier detection agent – in purple (mean and non-parametric 95% confidence intervals, see Methods ). e , Upper panel: probability of early lick after two sequential TF pulses of given magnitudes; middle panel: expected effect if both pulses influence early lick probability independently; lower panel: difference from the independent effect of TF pulses. f-g , The same format as in e , but for two TF pulses with 100 ms or 500 ms delay between them. h , The same format as in c , but shown for data generated by the outlier detection agent (for two sequential TF pulses). i , Difference in probability of early lick relative to the independent effect after a sequence of two fast TF pulses (top right corner in lower panels e-g ), normalized by the expected probability from the effect of independent pulses and shown as a function of delay between fast TF pulses. The results of the same analysis applied to the outlier detection agent data are shown in purple (mean and non-parametric 95% confidence intervals, see Methods ).
Extended Data Fig. 7 A simple two parameter leaky integrator model supports behavioural evidence integration + GLM change kernels across individual areas and large area groupings.
a , Schematic of the leaky-integrator model. b , Parameter search grid identifying which values the integration time and threshold best predicts early licks (i.e., correct predictions of early lick times (on single trials). c , Lick triggered stimulus average of early licks detected by the leaky integrator model, and early licks not detected by the model. d , Best-fit integration decay time of leaky-integrator model, shown per mouse (black dots) and mean across animals ( n = 15 mice, error bar is 95% confidence intervals). *** p < 0.001, two sided t -test. e , Relationship between real reaction time and predicted reaction time from leaky integrator model (tau: 0.25 s) for change size 1.25 Hz of example mouse 12. Correlation is calculated across all reaction times. f , Same as f but for change size 1.35 Hz. g , Correlation between observed and predicted reaction times during the change period for outlier detection agent (no integration, top ) and leaky-integrator model ( bottom ). Threshold parameters corresponding to best-fit were used for each model. The colour along each row corresponds to the correlation value between predicted hit lick reaction times and actual hit lick reaction times on trials with that change magnitude, conditioned by the maximum RT included for this analysis (cutoff time). h , Summary of panel g with results shown per mouse and RT combined across all change magnitudes (RT cutoff equal to 1 second from change onset). n = 15 mice, *** p < 0.001, two sided t -test. i , Mean decision value (integrated TF) after filtering stimulus though a leaky integrator model with a tau of 0.25 s. j , Mean reaction time curve for leaky integrator model. k , Example trials around change onset when model has no integration. Note the similarity to change kernels of TF responsive units in the SCs in Fig 3l . l , Example trials around change onset when model has leaky integration (0.25 s tau). Note the similarity to change kernels of to TF responsive units in the MOs in Fig 3l . m , Leaky evidence integration smooths and denoises the noisy sensory input so that the signal-to-noise ratio (S/σ) is considerably larger 0.5 s after change onset, compared to no integration–- making detection of noisy changes easier. n , Change size specific GLM change kernels for all areas recorded with 10 or more TF responsive units. o , Change size specific change kernels for major area groupings. Dotted line indicates the 50% response crossing for each change size.
Extended Data Fig. 8 Intrinsic vs learned TF pulse response properties.
a , Percentage of units encoding temporal frequency fluctuation during baseline in major area groupings with 95% binomial confidences in untrained and trained mice. Stars designate significance of difference (binomial test) in fractions between naïve and trained mice: n.s.: Not significant, ** p < 0.01, *** p < 0.001, binomial tests. Error bars are 95% binomial confidence intervals. OLF: Olfactory nuclei, Ctx: Cortex. See Supplementary Table 1 for n of each brain area grouping. b , Intrinsic timescales (tau) estimated for each TF responsive unit across the brain vs the TF response width for those units. Intrinsic times scales do not correlate with TF response width at a single cell level (p > 0.05, Pearson correlation, p-value is based on t-statistic). c , Same as in a but with units divided into major area groups. No area group has significant correlation between intrinsic times scales and TF response width at a single cell level (p > 0.05, Pearson correlation, p-value is based on t-statistic). d , Same as Fig. 4g , but here areal intrinsic time scale is extracted from TF responsive units only. In agreement with Fig. 4g , there is no correlation (Pearson correlation, p-value is based on t-statistic) between areal intrinsic timescales and median TF response width. e , intrinsic timescales of TF responsive units are similar to the intrinsic timescales as areas as a whole (Pearson correlation, p-value is based on t-statistic).
Extended Data Fig. 9 Differences in timing of preparatory activity between TF responsive and TF non-responsive populations.
a , Fraction of active units (combined across TF responsive and TF non-responsive units) as a function of time from the hit lick onset, shown across brain regions. Shades of red indicate a higher level of activity. Time points with lower 95% confidence interval (bootstrap test, see Methods ) smaller than zero are shown as white. Brain regions are sorted according to the time of the first significant activation (blue line, see Methods ). Black line shows the time of first significant activation using the same criterion as for Fig. 5f,g . b , Difference in onsets of preparatory activity across TF responsive and TF non-responsive subpopulations. Positive values indicate that TF responsive subpopulation has an earlier preparatory activity. Significant differences from zero are indicated by number of stars and area shaded in grey indicates 95% confidence intervals (bootstrap test, see Methods ). * p < 0.05, ** p < 0.01, *** p < 0.001. c , Difference in levels of activity between TF responsive and TF non-responsive subpopulations within each brain region. Shades of red indicate a higher level of activity across TF responsive subpopulation. Time points with non-significant differences (p ≥ 0.05, bootstrap test) in activity are shown as white. Brain regions are sorted according to the latency of the first significant difference in activation between TF responsive and non-responsive subpopulations (black line). d , Pearson correlation (p-value is based on t-statistic) across brain regions between the latency of the first significant difference in activation between TF responsive and TF non-responsive subpopulations and the median half-peak width of response to a fast pulse.
Extended Data Fig. 10 Definition of movement and movement-null subspaces.
a , Cross-validated cumulative R-squared coefficient of activity aligned to the hit lick onset shown across first six principal components for each brain region. Brain regions are sorted by the maximum cumulative R-squared value. b , Projections onto first four principal components of orofacial nuclei activity aligned to the hit lick onset. Projections on the first two principal components define the temporal profiles of activity within the two-dimensional movement subspace. The amount of cross-validated variance (average across draws) captured by each principal component is indicated on each panel. c , Projections of MOs activity (orange) aligned to the hit lick onset onto two movement (top) and two movement-null (bottom) dimensions. Projections of orofacial nuclei activity onto movement dimensions are shown in brown. d , Average cross-validated R-squared coefficient of mapping onto the movement subspace, with brain regions ordered from the best to worst mapping accuracy. The minimal value of R-squared coefficient for a brain region to be considered to have a good mapping onto a movement subspace is shown as a dashed red line (0.8). In all panels shaded regions indicate non-parametric 95% confidence intervals (see Methods ).
Extended Data Fig. 11 Occupancy of movement and movement-null subspaces and contribution of TF-responsive subpopulation within them.
a , Peak-normalized occupancy of movement subspace as a function of time for each brain region, relative to the hit lick onset time. Here and on panels b,c the order of brain regions is the same as on Fig. 6f . b , Peak-normalized occupancy of movement-null subspace as a function of time for each brain region. c , Average time of the peak occupancy within the movement-null subspace (green line), shown for each brain region. Shading indicates 95% confidence intervals. d , Distribution of loadings values along the first movement-null dimension that correspond to TF responsive (blue) and TF non-responsive (black) units in MOs. e , Minus log of p-value (blue line) for a paired 2-sided t-test between absolute values of loadings along the first movement-null dimension that correspond to TF responsive and TF non-responsive units. Dashed grey line indicates p = 0.05 level. c , Related to Fig. 6h . Comparison (Wilcoxon signed-rank test) of half-peak width of response to fast TF pulse between brain regions that had a disproportionate contribution of TF responsive subpopulation to preparatory activity in movement-null subspace (left bar, n = 16 brain regions) and the rest of brain regions (right bar, n = 12 brain regions). Bars indicate the mean across brain regions, error bars – 95% confidence intervals of the mean (bootstrap test, 2000 times). f , Relative contribution of TF responsive subpopulation within the movement subspace as a function of time for each brain region. Brain regions are shown in the same order as on Fig. 6h .
Extended Data Fig. 12 Alignment and scaling of TF pulse response projections on movement and movement-null dimensions.
a , Cosine of the angle between population response to a fast TF pulse and movement or movement-null dimensions. Values that are significantly different from zero (p < 0.05, 2-sided bootstrap test) are indicated by the black outline. b , Peak value of projections onto the first movement-null dimension of responses to a slow, fast, two sequential slow, and two sequential fast TF pulses. Results are shown for each brain region as the mean and 95% confidence intervals over 2000 cross-validations (see Methods ). See Supplementary Table 1 for number of neurons in each brain region. Number of starts indicates a 2-sided bootstrap test p-value of difference from zero for population response to a single fast or slow TF pulse, or a significance of a difference between responses to one or two sequential TF pulses. * p < 0.05, ** p < 0.01, *** p < 0.001. Non-significant effects are not indicated.
Extended Data Fig. 13 Breakdown of projections on main principal components by contributions from TF responsive and TF-nonresponsive units.
Each row shows projections on four main principal components of population activity within a given brain region aligned to the hit lick onset (same as on Fig. 6 ). The time course of each projection (black) was decomposed into a sum of contributions from TF responsive (blue) and TF non-responsive (red) units. Grey line indicates projection expected from a random sample of the same size as there were TF responsive units, taken randomly (with replacement) from the full population. Data is shown as mean and 95% confidence intervals across 2000 cross-validations (see Methods ). The amount of cross-validated variance captured by each principal component is indicated on top of each panel.
Supplementary information
Supplementary information.
Supplementary Tables 1 and 2
Reporting Summary
Peer review file, source data, source data fig. 1, source data fig. 2, source data fig. 3, source data fig. 4, source data fig. 5, source data fig. 6, rights and permissions.
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/ .
Reprints and permissions
About this article
Cite this article.
Khilkevich, A., Lohse, M., Low, R. et al. Brain-wide dynamics linking sensation to action during decision-making. Nature (2024). https://doi.org/10.1038/s41586-024-07908-w
Download citation
Received : 13 July 2023
Accepted : 05 August 2024
Published : 11 September 2024
DOI : https://doi.org/10.1038/s41586-024-07908-w
Share this article
Anyone you share the following link with will be able to read this content:
Sorry, a shareable link is not currently available for this article.
Provided by the Springer Nature SharedIt content-sharing initiative
By submitting a comment you agree to abide by our Terms and Community Guidelines . If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.
Quick links
- Explore articles by subject
- Guide to authors
- Editorial policies
Sign up for the Nature Briefing newsletter — what matters in science, free to your inbox daily.

IMAGES
VIDEO
COMMENTS
Null hypothesis: The sample data provides no evidence to support some claim being made by an individual. Alternative hypothesis: The sample data does provide sufficient evidence to support the claim being made by an individual. For example, suppose it's assumed that the average height of a certain species of plant is 20 inches tall. However ...
To distinguish it from other hypotheses, the null hypothesis is written as H 0 (which is read as "H-nought," "H-null," or "H-zero"). A significance test is used to determine the likelihood that the results supporting the null hypothesis are not due to chance. A confidence level of 95% or 99% is common. Keep in mind, even if the confidence level is high, there is still a small chance the ...
The null hypothesis (H0) answers "No, there's no effect in the population.". The alternative hypothesis (Ha) answers "Yes, there is an effect in the population.". The null and alternative are always claims about the population. That's because the goal of hypothesis testing is to make inferences about a population based on a sample.
When your sample contains sufficient evidence, you can reject the null and conclude that the effect is statistically significant. Statisticians often denote the null hypothesis as H 0 or H A.. Null Hypothesis H 0: No effect exists in the population.; Alternative Hypothesis H A: The effect exists in the population.; In every study or experiment, researchers assess an effect or relationship.
An example of the null hypothesis is that light color has no effect on plant growth. The null hypothesis (H 0) is the hypothesis that states there is no statistical difference between two sample sets. In other words, it assumes the independent variable does not have an effect on the dependent variable in a scientific experiment.
The null and alternative hypotheses are two competing claims that researchers weigh evidence for and against using a statistical test: Null hypothesis (H0): There's no effect in the population. Alternative hypothesis (HA): There's an effect in the population. The effect is usually the effect of the independent variable on the dependent ...
Review. In a hypothesis test, sample data is evaluated in order to arrive at a decision about some type of claim.If certain conditions about the sample are satisfied, then the claim can be evaluated for a population. In a hypothesis test, we: Evaluate the null hypothesis, typically denoted with \(H_{0}\).The null is not rejected unless the hypothesis test shows otherwise.
Step 1: Figure out the hypothesis from the problem. The hypothesis is usually hidden in a word problem, and is sometimes a statement of what you expect to happen in the experiment. The hypothesis in the above question is "I expect the average recovery period to be greater than 8.2 weeks.". Step 2: Convert the hypothesis to math.
The actual test begins by considering two hypotheses.They are called the null hypothesis and the alternative hypothesis.These hypotheses contain opposing viewpoints. H 0, the —null hypothesis: a statement of no difference between sample means or proportions or no difference between a sample mean or proportion and a population mean or proportion. In other words, the difference equals 0.
Null Hypothesis Examples. "Hyperactivity is unrelated to eating sugar " is an example of a null hypothesis. If the hypothesis is tested and found to be false, using statistics, then a connection between hyperactivity and sugar ingestion may be indicated. A significance test is the most common statistical test used to establish confidence in a ...
5. Phrase your hypothesis in three ways. To identify the variables, you can write a simple prediction in if…then form. The first part of the sentence states the independent variable and the second part states the dependent variable. If a first-year student starts attending more lectures, then their exam scores will improve.
When your p-value is less than or equal to your significance level, you reject the null hypothesis. In other words, smaller p-values are taken as stronger evidence against the null hypothesis. Conversely, when the p-value is greater than your significance level, you fail to reject the null hypothesis. In this case, the sample data provides ...
Write a research null hypothesis as a statement that the studied variables have no relationship to each other, or that there's no difference between 2 groups. Write a statistical null hypothesis as a mathematical equation, such as. μ 1 = μ 2 {\displaystyle \mu _ {1}=\mu _ {2}} if you're comparing group means.
Present the findings in your results and discussion section. Though the specific details might vary, the procedure you will use when testing a hypothesis will always follow some version of these steps. Table of contents. Step 1: State your null and alternate hypothesis. Step 2: Collect data. Step 3: Perform a statistical test.
A possible null hypothesis is that the mean male score is the same as the mean female score: H 0: μ 1 = μ 2. where H 0 = the null hypothesis, μ 1 = the mean of population 1, and μ 2 = the mean of population 2. A stronger null hypothesis is that the two samples have equal variances and shapes of their respective distributions.
10.1 - Setting the Hypotheses: Examples. A significance test examines whether the null hypothesis provides a plausible explanation of the data. The null hypothesis itself does not involve the data. It is a statement about a parameter (a numerical characteristic of the population). These population values might be proportions or means or ...
Concept Review. In a hypothesis test, sample data is evaluated in order to arrive at a decision about some type of claim.If certain conditions about the sample are satisfied, then the claim can be evaluated for a population. In a hypothesis test, we: Evaluate the null hypothesis, typically denoted with H 0.The null is not rejected unless the hypothesis test shows otherwise.
A null hypothesis is a general assertion or default position that there is no relationship or effect between two measured phenomena. It's a critical part of statistics, data analysis, and the scientific method. This concept forms the basis of testing statistical significance and allows researchers to be objective in their conclusions.
The alternative hypothesis (Ha H a) is a claim about the population that is contradictory to H0 H 0 and what we conclude when we reject H0 H 0. Since the null and alternative hypotheses are contradictory, you must examine evidence to decide if you have enough evidence to reject the null hypothesis or not. The evidence is in the form of sample data.
Most technical papers rely on just the first formulation, even though you may see some of the others in a statistics textbook. Null hypothesis: " x is equal to y.". Alternative hypothesis " x is not equal to y.". Null hypothesis: " x is at least y.". Alternative hypothesis " x is less than y.". Null hypothesis: " x is at most ...
If the p-value that corresponds to the test statistic t with (n 1 +n 2-1) degrees of freedom is less than your chosen significance level (common choices are 0.10, 0.05, and 0.01) then you can reject the null hypothesis. Two Sample t-test: Assumptions. For the results of a two sample t-test to be valid, the following assumptions should be met:
What is Null Hypothesis? Null Hypothesis in statistical analysis suggests the absence of statistical significance within a specific set of observed data. Hypothesis testing, using sample data, evaluates the validity of this hypothesis. Commonly denoted as H 0 or simply "null," it plays an important role in quantitative analysis, examining theories related to markets, investment strategies ...
Confusingly, however, you cannot interpret the p-value as telling you the probability (how likely) that the null hypothesis is true. If, however, the test statistic is less than the critical value, then the conclusion is that the null hypothesis is to be provisionally accepted. The test statistic can be assigned a probability or p-value.
The actual test begins by considering two hypotheses.They are called the null hypothesis and the alternative hypothesis.These hypotheses contain opposing viewpoints. H 0: The null hypothesis: It is a statement of no difference between the variables—they are not related. This can often be considered the status quo and as a result if you cannot accept the null it requires some action.
For every quantity of interest, the value computed from the real data was compared to values computed from each synthetic dataset, comprising 10,000 samples from the null distribution.