Generate accurate APA citations for free
- Knowledge Base
- APA Style 7th edition
- How to create an APA Style appendix

How to Create an APA Style Appendix | Format & Examples
Published on October 16, 2020 by Jack Caulfield . Revised on August 9, 2022.
An appendix is a section at the end of an academic text where you include extra information that doesn’t fit into the main text. The plural of appendix is “appendices.”
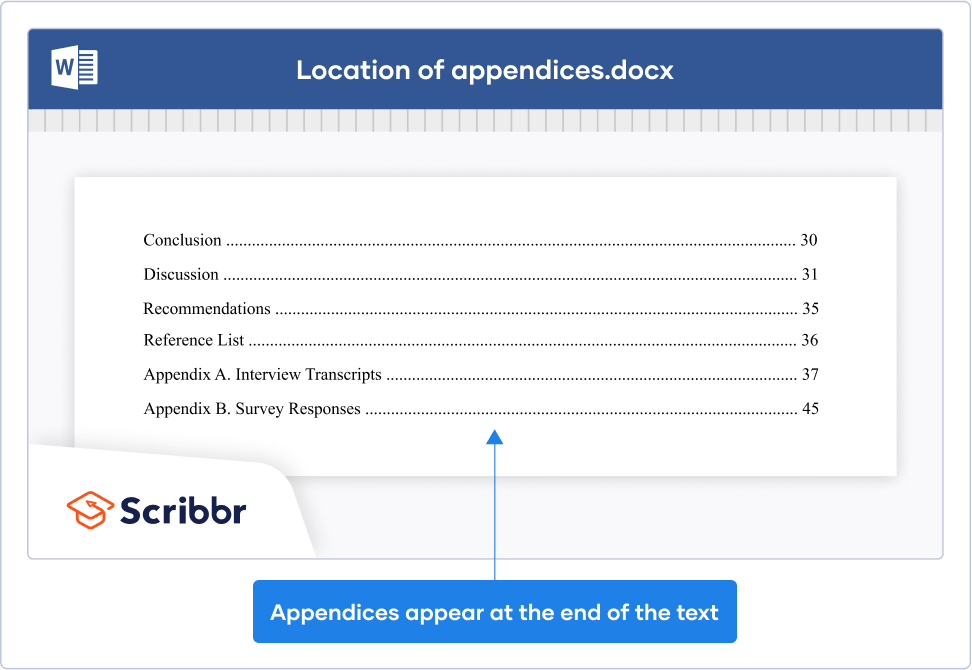
In an APA Style paper, appendices are placed at the very end, after the reference list .

Instantly correct all language mistakes in your text
Upload your document to correct all your mistakes in minutes
Table of contents
Do i need an appendix, appendix format example, organizing and labeling your appendices, frequently asked questions.
You don’t always need to include any appendices. An appendix should present information that supplements the reader’s understanding of your research but is not essential to the argument of your paper . Essential information is included in the main text.
For example, you might include some of the following in an appendix:
- Full transcripts of interviews you conducted (which you can quote from in the main text)
- Documents used in your research, such as questionnaires , instructions, tests, or scales
- Detailed statistical data (often presented in tables or figures )
- Detailed descriptions of equipment used
You should refer to each appendix at least once in the main text. If you don’t refer to any information from an appendix, it should not be included.
When you discuss information that can be found in an appendix, state this the first time you refer to it:
Note that, if you refer to the same interviews again, it’s not necessary to mention the appendix each time.
Scribbr Citation Checker New
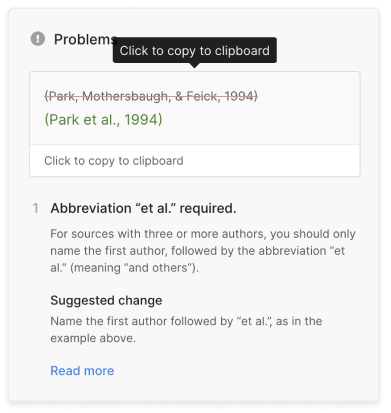
The AI-powered Citation Checker helps you avoid common mistakes such as:
- Missing commas and periods
- Incorrect usage of “et al.”
- Ampersands (&) in narrative citations
- Missing reference entries
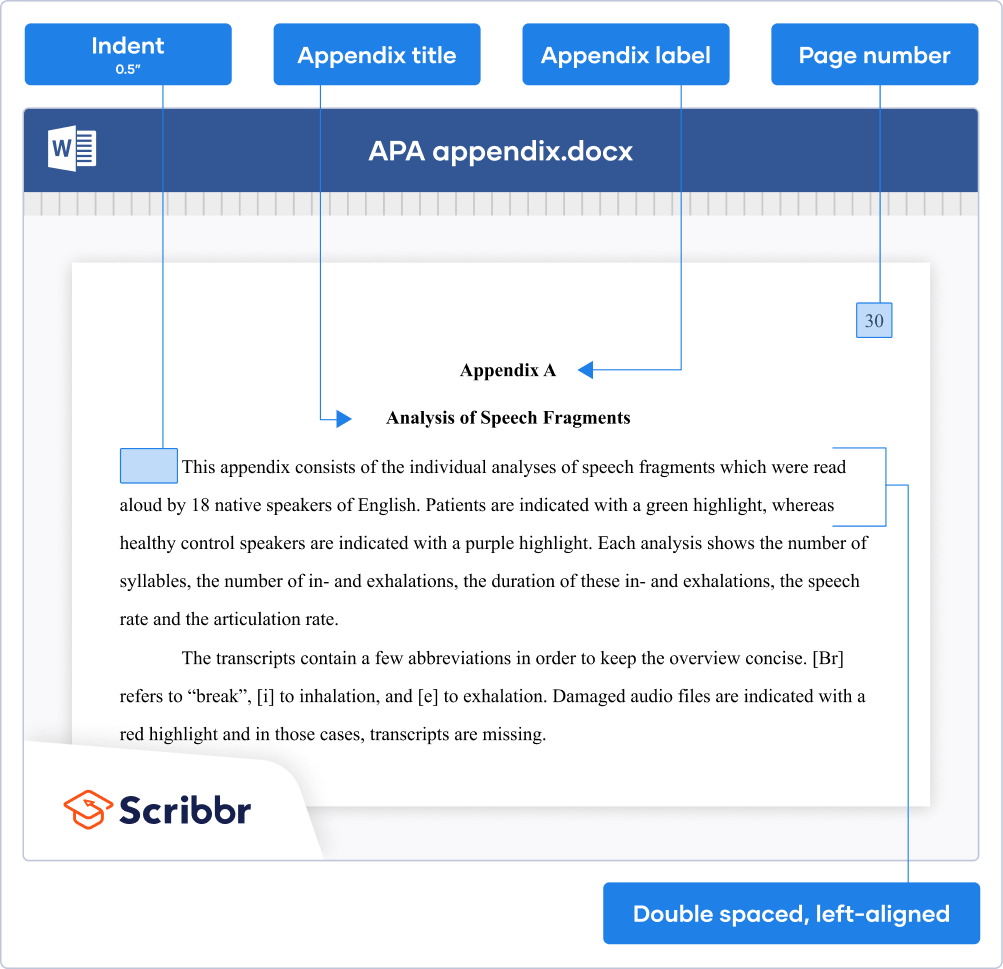
The appendix label appears at the top of the page, bold and centered. On the next line, include a descriptive title, also bold and centered.
The text is presented in general APA format : left-aligned, double-spaced, and with page numbers in the top right corner. Start a new page for each new appendix.
The example image below shows how to format an APA Style appendix.
If you include just one appendix, it is simply called “Appendix” and referred to as such in-text:
When more than one appendix is included, they are labeled “Appendix A,” “Appendix B,” and so on.
Present and label your appendices in the order they are referred to in the main text.
Labeling tables and figures in appendices
An appendix may include (or consist entirely of) tables and/or figures . Present these according to the same formatting rules as in the main text.
Tables and figures included in appendices are labeled differently, however. Use the appendix’s letter in addition to a number. Tables and figures are still numbered separately and according to the order they’re referred to in the appendix.
For example, in Appendix A, your tables are Table A1, Table A2, etc; your figures are Figure A1, Figure A2, etc.
The numbering restarts with each appendix: For example, the first table in Appendix B is Table B1; the first figure in Appendix C is Figure C1; and so on. If you only have one appendix, use A1, A2, etc.
If you want to refer specifically to a table or figure from an appendix in the main text, use the table or figure’s label (e.g. “see Table A3”).
If an appendix consists entirely of a single table or figure, simply use the appendix label to refer to the table or figure. For example, if Appendix C is just a table, refer to the table as “Appendix C,” and don’t add an additional label or title for the table itself.
An appendix contains information that supplements the reader’s understanding of your research but is not essential to it. For example:
- Interview transcripts
- Questionnaires
- Detailed descriptions of equipment
Something is only worth including as an appendix if you refer to information from it at some point in the text (e.g. quoting from an interview transcript). If you don’t, it should probably be removed.
Appendices in an APA Style paper appear right at the end, after the reference list and after your tables and figures if you’ve also included these at the end.
When you include more than one appendix in an APA Style paper , they should be labeled “Appendix A,” “Appendix B,” and so on.
When you only include a single appendix, it is simply called “Appendix” and referred to as such in the main text.
Yes, if relevant you can and should include APA in-text citations in your appendices . Use author-date citations as you do in the main text.
Any sources cited in your appendices should appear in your reference list . Do not create a separate reference list for your appendices.
Cite this Scribbr article
If you want to cite this source, you can copy and paste the citation or click the “Cite this Scribbr article” button to automatically add the citation to our free Citation Generator.
Caulfield, J. (2022, August 09). How to Create an APA Style Appendix | Format & Examples. Scribbr. Retrieved August 19, 2024, from https://www.scribbr.com/apa-style/appendices/
Is this article helpful?

Jack Caulfield
Other students also liked, creating an apa style table of contents, how to format tables and figures in apa style, apa format for academic papers and essays, "i thought ai proofreading was useless but..".
I've been using Scribbr for years now and I know it's a service that won't disappoint. It does a good job spotting mistakes”
Have a language expert improve your writing
Run a free plagiarism check in 10 minutes, automatically generate references for free.
- Knowledge Base
- Dissertation
- Research Paper Appendix | Example & Templates
Research Paper Appendix | Example & Templates
Published on 15 August 2022 by Kirsten Dingemanse and Tegan George. Revised on 25 October 2022.
An appendix is a supplementary document that facilitates your reader’s understanding of your research but is not essential to your core argument. Appendices are a useful tool for providing additional information or clarification in a research paper , dissertation , or thesis without making your final product too long.
Appendices help you provide more background information and nuance about your topic without disrupting your text with too many tables and figures or other distracting elements.
We’ve prepared some examples and templates for you, for inclusions such as research protocols, survey questions, and interview transcripts. All are worthy additions to an appendix. You can download these in the format of your choice below.
Download Word doc Download Google doc
Instantly correct all language mistakes in your text
Be assured that you'll submit flawless writing. Upload your document to correct all your mistakes.

Table of contents
What is an appendix in a research paper, what to include in an appendix, how to format an appendix, how to refer to an appendix, where to put your appendices, other components to consider, appendix checklist.
In the main body of your research paper, it’s important to provide clear and concise information that supports your argument and conclusions . However, after doing all that research, you’ll often find that you have a lot of other interesting information that you want to share with your reader.
While including it all in the body would make your paper too long and unwieldy, this is exactly what an appendix is for.
As a rule of thumb, any detailed information that is not immediately needed to make your point can go in an appendix. This helps to keep your main text focused but still allows you to include the information you want to include somewhere in your paper.
The only proofreading tool specialized in correcting academic writing
The academic proofreading tool has been trained on 1000s of academic texts and by native English editors. Making it the most accurate and reliable proofreading tool for students.
Correct my document today
An appendix can be used for different types of information, such as:
- Supplementary results : Research findings are often presented in different ways, but they don’t all need to go in your paper. The results most relevant to your research question should always appear in the main text, while less significant results (such as detailed descriptions of your sample or supplemental analyses that do not help answer your main question), can be put in an appendix.
- Statistical analyses : If you conducted statistical tests using software like Stata or R, you may also want to include the outputs of your analysis in an appendix.
- Further information on surveys or interviews : Written materials or transcripts related to things such as surveys and interviews can also be placed in an appendix.
You can opt to have one long appendix, but separating components (like interview transcripts, supplementary results, or surveys) into different appendices makes the information simpler to navigate.
Here are a few tips to keep in mind:
- Always start each appendix on a new page.
- Assign it both a number (or letter) and a clear title, such as ‘Appendix A. Interview transcripts’. This makes it easier for your reader to find the appendix, as well as for you to refer back to it in your main text.
- Number and title the individual elements within each appendix (e.g., ‘Transcripts’) to make it clear what you are referring to. Restart the numbering in each appendix at 1.
It is important that you refer to each of your appendices at least once in the main body of your paper. This can be done by mentioning the appendix and its number or letter, either in parentheses or within the main part of a sentence. It is also possible to refer to a particular component of an appendix.
Appendix B presents the correspondence exchanged with the fitness boutique. Example 2. Referring to an appendix component These results (see Appendix 2, Table 1) show that …
It is common to capitalise ‘Appendix’ when referring to a specific appendix, but it is not mandatory. The key is just to make sure that you are consistent throughout your entire paper, similarly to consistency in capitalising headings and titles in academic writing.
However, note that lowercase should always be used if you are referring to appendices in general. For instance, ‘The appendices to this paper include additional information about both the survey and the interviews.’
Prevent plagiarism, run a free check.
The simplest option is to add your appendices after the main body of your text, after you finish citing your sources in the citation style of your choice . If this is what you choose to do, simply continue with the next page number. Another option is to put the appendices in a separate document that is delivered with your dissertation.
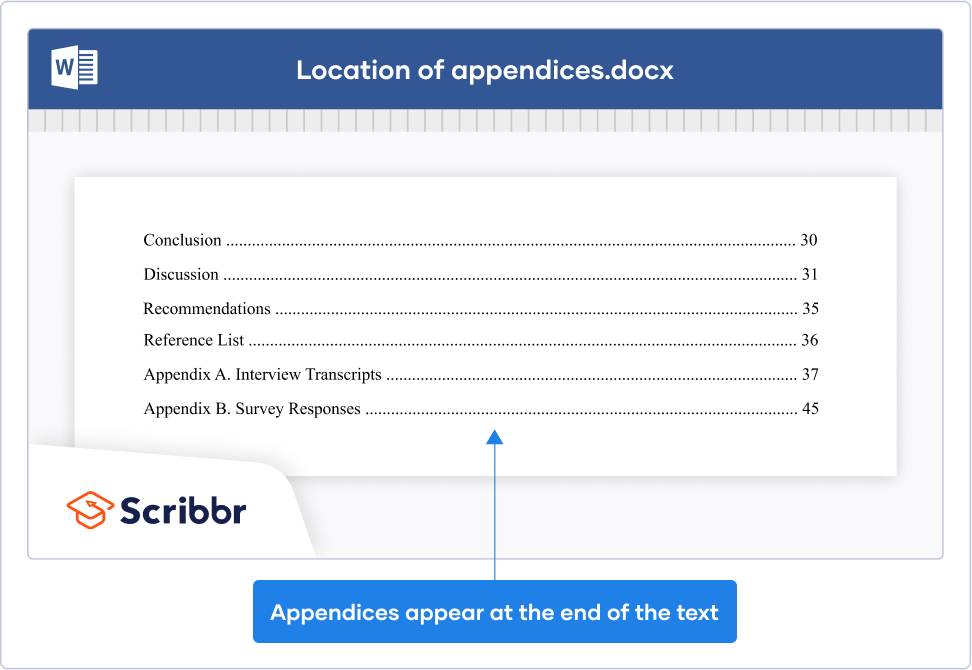

Remember that any appendices should be listed in your paper’s table of contents .
There are a few other supplementary components related to appendices that you may want to consider. These include:
- List of abbreviations : If you use a lot of abbreviations or field-specific symbols in your dissertation, it can be helpful to create a list of abbreviations .
- Glossary : If you utilise many specialised or technical terms, it can also be helpful to create a glossary .
- Tables, figures and other graphics : You may find you have too many tables, figures, and other graphics (such as charts and illustrations) to include in the main body of your dissertation. If this is the case, consider adding a figure and table list .
Checklist: Appendix
All appendices contain information that is relevant, but not essential, to the main text.
Each appendix starts on a new page.
I have given each appendix a number and clear title.
I have assigned any specific sub-components (e.g., tables and figures) their own numbers and titles.
My appendices are easy to follow and clearly formatted.
I have referred to each appendix at least once in the main text.
Your appendices look great! Use the other checklists to further improve your thesis.
Cite this Scribbr article
If you want to cite this source, you can copy and paste the citation or click the ‘Cite this Scribbr article’ button to automatically add the citation to our free Reference Generator.
Dingemanse, K. & George, T. (2022, October 25). Research Paper Appendix | Example & Templates. Scribbr. Retrieved 19 August 2024, from https://www.scribbr.co.uk/thesis-dissertation/appendix/
Is this article helpful?
Kirsten Dingemanse
Other students also liked, thesis & dissertation acknowledgements | tips & examples, dissertation title page, how to write a results section | tips & examples.
- USC Libraries
- Research Guides
Organizing Your Social Sciences Research Paper
- Purpose of Guide
- Design Flaws to Avoid
- Independent and Dependent Variables
- Glossary of Research Terms
- Reading Research Effectively
- Narrowing a Topic Idea
- Broadening a Topic Idea
- Extending the Timeliness of a Topic Idea
- Academic Writing Style
- Applying Critical Thinking
- Choosing a Title
- Making an Outline
- Paragraph Development
- Research Process Video Series
- Executive Summary
- The C.A.R.S. Model
- Background Information
- The Research Problem/Question
- Theoretical Framework
- Citation Tracking
- Content Alert Services
- Evaluating Sources
- Primary Sources
- Secondary Sources
- Tiertiary Sources
- Scholarly vs. Popular Publications
- Qualitative Methods
- Quantitative Methods
- Insiderness
- Using Non-Textual Elements
- Limitations of the Study
- Common Grammar Mistakes
- Writing Concisely
- Avoiding Plagiarism
- Footnotes or Endnotes?
- Further Readings
- Generative AI and Writing
- USC Libraries Tutorials and Other Guides
- Bibliography
An appendix contains supplementary material that is not an essential part of the text itself but which may be helpful in providing a more comprehensive understanding of the research problem. An appendix may also contain information that is too cumbersome to be included in the body of the paper. A separate appendix should be used for each distinct topic or set of data and always have a title descriptive of its contents [e.g., Appendix 1: Interview Protocol].
Tables, Appendices, Footnotes and Endnotes. The Writing Lab and The OWL. Purdue University.
Importance of...
Appendices are always supplementary to the research paper. As such, your study must be able to stand alone without the appendices, and the paper must contain all information including tables, diagrams, and results necessary to understand the research problem. The key point to remember when including an appendix or appendices is that the information is non-essential to understanding the research problem being investigated. In other words, if it were removed, the reader would still be able to comprehend the significance, validity , and implications of your research even if that additional data was missing.
It is appropriate to include appendices for the following reasons:
- Including this material in the body of the paper that would render it poorly structured or interrupt the narrative flow;
- Information is too lengthy and detailed to be easily summarized in the body of the paper;
- Inclusion of helpful, supporting, or useful material would otherwise distract the reader from the main content of the paper;
- Provides relevant information or data that is more easily understood or analyzed in a self-contained section of the paper;
- Can be used when there are constraints placed on the length of your paper; and,
- Provides a place to further demonstrate your understanding of the research problem by giving additional details about a new or innovative method, technical details, or design protocols.
Appendices. Academic Skills Office, University of New England; Chapter 12, "Use of Appendices." In Guide to Effective Grant Writing: How to Write a Successful NIH Grant . Otto O. Yang. (New York: Kluwer Academic, 2005), pp. 55-57; Tables, Appendices, Footnotes and Endnotes. The Writing Lab and The OWL. Purdue University.
Structure and Writing Style
I. General Points to Consider
When considering whether to include content in an appendix, keep in mind the following:
- It is usually good practice to include your raw data in an appendix, laying it out in a clear format so the reader can re-check your results. Another option if you have a large amount of raw data is to consider placing it online [e.g., on a Google drive] and note that this is the appendix to your research paper.
- Any tables and figures included in the appendix should be numbered as a separate sequence from the main paper . Remember that appendices contain non-essential information that, if removed, would not diminish a reader's ability to understand the research problem being investigated. This is why non-textual elements should not carry over the sequential numbering of non-textual elements in the body of your paper.
- If you have more than three appendices, consider listing them on a separate page in the table of contents . This will help the reader know what information is included in the appendices. Note that some works list appendices in the table of contents before the first chapter while other styles list the appendices after the conclusion but before your references. Consult with your professor to confirm if there is a preferred approach.
- The appendix can be a good place to put maps, photographs, diagrams, and other images , if you feel that it will help the reader to understand the content of your paper, while keeping in mind the study should be understood without them.
- An appendix should be streamlined and not loaded with a lot information . If you have a very long and complex appendix, it is a good idea to break it down into separate appendices, allowing the reader to find relevant information quickly as the information is covered in the body of the paper.
II. Content
Never include an appendix that isn’t referred to in the text . All appendices should be summarized in your paper where it is relevant to the content. Appendices should also be arranged sequentially by the order they were first referenced in the text [i.e., Appendix 1 should not refer to text on page eight of your paper and Appendix 2 relate to text on page six].
There are few rules regarding what type of material can be included in an appendix, but here are some common examples:
- Correspondence -- if your research included collaborations with others or outreach to others, then correspondence in the form of letters, memorandums, or copies of emails from those you interacted with could be included.
- Interview Transcripts -- in qualitative research, interviewing respondents is often used to gather information. The full transcript from an interview is important so the reader can read the entire dialog between researcher and respondent. The interview protocol [list of questions] should also be included.
- Non-textual elements -- as noted above, if there are a lot of non-textual items, such as, figures, tables, maps, charts, photographs, drawings, or graphs, think about highlighting examples in the text of the paper but include the remainder in an appendix.
- Questionnaires or surveys -- this is a common form of data gathering. Always include the survey instrument or questionnaires in an appendix so the reader understands not only the questions asked but the sequence in which they were asked. Include all variations of the instruments as well if different items were sent to different groups [e.g., those given to teachers and those given to administrators] .
- Raw statistical data – this can include any numerical data that is too lengthy to include in charts or tables in its entirety within the text. This is important because the entire source of data should be included even if you are referring to only certain parts of a chart or table in the text of your paper.
- Research instruments -- if you used a camera, or a recorder, or some other device to gather information and it is important for the reader to understand how, when, and/or where that device was used.
- Sample calculations – this can include quantitative research formulas or detailed descriptions of how calculations were used to determine relationships and significance.
NOTE: Appendices should not be a dumping ground for information. Do not include vague or irrelevant information in an appendix; this additional information will not help the reader’s overall understanding and interpretation of your research and may only distract the reader from understanding the significance of your overall study.
ANOTHER NOTE: Appendices are intended to provide supplementary information that you have gathered or created; it is not intended to replicate or provide a copy of the work of others. For example, if you need to contrast the techniques of analysis used by other authors with your own method of analysis, summarize that information, and cite to the original work. In this case, a citation to the original work is sufficient enough to lead the reader to where you got the information. You do not need to provide a copy of this in an appendix.
III. Format
Here are some general guideline on how to format appendices . If needed, consult the writing style guide [e.g., APA, MLS, Chicago] your professor wants you to use for more detail or choose the style you are most familiar with:
- Appendices may precede or follow your list of references.
- Each appendix begins on a new page.
- The order they are presented is dictated by the order they are mentioned in the text of your research paper.
- The heading should be "Appendix," followed by a letter or number [e.g., "Appendix A" or "Appendix 1"], centered and written in bold type.
- If there is a table of contents, the appendices must be listed.
- Depending on the type of information, the content can be presented in landscape format rather than regular portrait format.
- The page number(s) of the appendix/appendices will continue on with the numbering from the last page of the text.
Appendices. The Structure, Format, Content, and Style of a Journal-Style Scientific Paper. Department of Biology. Bates College; Appendices. Academic Skills Office, University of New England; Appendices. Writing Center, Walden University; Chapter 12, "Use of Appendices." In Guide to Effective Grant Writing: How to Write a Successful NIH Grant . Otto O. Yang. (New York: Kluwer Academic, 2005), pp. 55-57 ; Tables, Appendices, Footnotes and Endnotes. The Writing Lab and The OWL. Purdue University; Lunsford, Andrea A. and Robert Connors. The St. Martin's Handbook . New York: St. Martin's Press, 1989; What To Know About The Purpose And Format Of A Research Paper Appendix. LoyolaCollegeCulion.com.
Writing Tip
Consider Putting Your Appendices Online
Appendices are useful because they provide the reader with information that supports your study without breaking up the narrative or distracting from the main purpose of your paper. If you have a lot of raw data or information that is difficult to present in textual form, consider uploading it to an online site. This prevents your paper from having a large and unwieldy set of appendices and it supports a growing movement within academe to make data more freely available for re-analysis. If you do create an online portal to your data, note it prominently in your paper with the correct URL and access procedures if it is a secured site, or if needed, with clear directions on how to contact the author to obtain access.
Piwowar, Heather A., Roger S. Day, and Douglas B. Fridsma. “Sharing Detailed Research Data Is Associated with Increased Citation Rate.” PloS ONE (March 21, 2007); Wicherts, Jelte M., Marjan Bakker, and Dylan Molenaar. “Willingness to Share Research Data Is Related to the Strength of the Evidence and the Quality of Reporting of Statistical Results.” PLoS ONE (November 2, 2011).
- << Previous: 9. The Conclusion
- Next: 10. Proofreading Your Paper >>
- Last Updated: Aug 13, 2024 12:57 PM
- URL: https://libguides.usc.edu/writingguide
- Franklin University |
- Help & Support |
- Locations & Maps |

- | Research Guides
To access Safari eBooks,
- Select not listed in the Select Your Institution drop down menu.
- Enter your Franklin email address and click Go
- click "Already a user? Click here" link
- Enter your Franklin email and the password you used to create your Safari account.
Continue Close
APA Citation Style 7th Edition
- APA Style Overview
- Sample Documents & Guides
- Multiple Sources With the Same Author and Year
- Websites & Web Documents
- Course Materials (Slides, Lecture Notes, Specialty Software)
- Citing Business Databases
- Film, Videos, & Podcasts
- Art, Photos, Tables & Figures
- Legal Materials & Tax Codes
- Dissertations
- Pamphlet or Brochure
- Interviews, E-mail, Intranet, Religious Works, & Secondary Sources (7th edition)
- Footnotes This link opens in a new window
What goes into an Appendix?
Where is an appendix placed, labeling the appendix, formatting the appendix.
- Evaluating Sources This link opens in a new window
- Understanding Plagiarism
- RefWorks This link opens in a new window
"Material that supplements the content of the paper, but would be distracting or inappropriate to include in the body of the paper is to be placed in an appendix." This includes "materials that are relatively brief and that are easily presented in print format" ( Publication Manual of the APA: 6th edition , section 2.13; Publication Manual of the APA: 7th edition , section 2.14). Examples include "mathematical proofs, lists of words, a questionnaire used in the research, a detailed description of an apparatus used in the research, etc" ( Purdue OWL .)
An appendix (or appendices) follow the reference list. Use the following order for your paper:
- Abstract ( if required, start on a new page, numbered page 2)
- Text (start on a new page, numbered 3)
- References (start on a new page)
- Tables (start each on a new page)
- Figures (start each on a new page; include caption on page with figure)
- Appendices (start each on a new page)
- If only one appendix, label it Appendix
- If more than one appendix: label each one with a capital letter (Appendix A, Appendix B, etc.) in the order in which it is mentioned in the text
- Each appendix must have a title
- In the text, refer to appendices by their labels:
"produced the same results for both studies (see Appendices A and B for complete proofs)."
- Begin each appendix on a separate page
- At the top of the page, center the word Appendix and the identifying capital letters (A, B, etc.) in the order in which they are mentioned in the text.
- Center the title of the appendix using uppercase and lowercase letter on the next line
- Begin the text of the appendix flush left, followed by indented paragraphs.
A sample appendix is below:

- << Previous: Footnotes
- Next: Evaluating Sources >>
- Last Updated: May 29, 2024 3:56 PM
- URL: https://guides.franklin.edu/APA
- Privacy Policy

Home » Appendices – Writing Guide, Types and Examples
Appendices – Writing Guide, Types and Examples
Table of Contents

Definition:
Appendices refer to supplementary materials or documents that are attached to the end of a Book, Report , Research Paper , Thesis or other written work. These materials can include charts, graphs, tables, images, or other data that support the main content of the work.
Types of Appendices
Types of appendices that can be used depending on the content and purpose of the document. These types of Appendices are as follows:
Statistical Appendices
Statistical appendices are used to present raw data or statistical analysis that is relevant to the main text but would be too bulky to include in the main body of the document. These appendices may include tables, graphs, charts, or other types of visual aids that help to illustrate the data.
Technical Appendices
Technical appendices are used to provide detailed technical information that is relevant to the main text but would be too complex or lengthy to include in the main body of the document. These appendices may include equations, formulas, diagrams, or other technical details that are important for understanding the subject matter.
Bibliographical Appendices
Bibliographical appendices are used to provide additional references or sources that are relevant to the main text but were not cited in the main body of the document. These appendices may include lists of books, articles, or other resources that the author consulted in the course of their research.
Historical Appendices
Historical appendices are used to provide background information or historical context that is relevant to the main text but would be too lengthy or distracting to include in the main body of the document. These appendices may include timelines, maps, biographical sketches, or other historical details that help to contextualize the subject matter.
Supplemental Appendices
Supplemental appendices are used to provide additional material that is relevant to the main text but does not fit into any of the other categories. These appendices may include interviews, surveys, case studies, or other types of supplemental material that help to further illustrate the subject matter.
Applications of Appendices
Some applications of appendices are:
- Providing detailed data and statistics: Appendices are often used to include detailed data and statistics that support the findings presented in the main body of the document. For example, in a research paper, an appendix might include raw data tables or graphs that were used to support the study’s conclusions.
- Including technical details: Appendices can be used to include technical details that may be of interest to a specialized audience. For example, in a technical report, an appendix might include detailed calculations or equations that were used to develop the report’s recommendations.
- Presenting supplementary information: Appendices can be used to present supplementary information that is related to the main content but doesn’t fit well within the main body of the document. For example, in a business proposal, an appendix might include a list of references or a glossary of terms.
- Providing supporting documentation: Appendices can be used to provide supporting documentation that is required by the document’s audience. For example, in a legal document, an appendix might include copies of contracts or agreements that were referenced in the main body of the document.
- Including multimedia materials : Appendices can be used to include multimedia materials that supplement the main content. For example, in a book, an appendix might include photographs, maps, or illustrations that help to clarify the text.
Importance of Appendices
Appendices are important components of research papers, reports, Thesis, and other academic papers. They are supplementary materials that provide additional information and data that support the main text. Here are some reasons why appendices are important:
- Additional Information : Appendices provide additional information that is too detailed or too lengthy to include in the main text. This information includes raw data, graphs, tables, and charts that support the research findings.
- Clarity and Conciseness : Appendices help to maintain the clarity and conciseness of the main text. By placing detailed information and data in appendices, writers can avoid cluttering the main text with lengthy descriptions and technical details.
- Transparency : Appendices increase the transparency of research by providing readers with access to the data and information used in the research process. This transparency increases the credibility of the research and allows readers to verify the findings.
- Accessibility : Appendices make it easier for readers to access the data and information that supports the research. This is particularly important in cases where readers want to replicate the research or use the data for their own research.
- Compliance : Appendices can be used to comply with specific requirements of the research project or institution. For example, some institutions may require researchers to include certain types of data or information in the appendices.
Appendices Structure
Here is an outline of a typical structure for an appendix:
I. Introduction
- A. Explanation of the purpose of the appendix
- B. Brief overview of the contents
II. Main Body
- A. Section headings or subheadings for different types of content
- B. Detailed descriptions, tables, charts, graphs, or images that support the main content
- C. Labels and captions for each item to help readers navigate and understand the content
III. Conclusion
- A. Summary of the key points covered in the appendix
- B. Suggestions for further reading or resources
IV. Appendices
- A. List of all the appendices included in the document
- B. Table of contents for the appendices
V. References
- A. List of all the sources cited in the appendix
- B. Proper citation format for each source
Example of Appendices
here’s an example of what appendices might look like for a survey:
Appendix A:
Survey Questionnaire
This section contains a copy of the survey questionnaire used for the study.
- What is your age?
- What is your gender?
- What is your highest level of education?
- How often do you use social media?
- Which social media platforms do you use most frequently?
- How much time do you typically spend on social media each day?
- Do you feel that social media has had a positive or negative impact on your life?
- Have you ever experienced cyberbullying or harassment on social media?
- Have you ever been influenced by social media to make a purchase or try a new product?
- In your opinion, what are the biggest advantages and disadvantages of social media?
Appendix B:
Participant Demographics
This section includes a table with demographic information about the survey participants, such as age, gender, and education level.
Age Gender Education Level
- 20 Female Bachelor’s Degree
- 32 Male Master’s Degree
- 45 Female High School Diploma
- 28 Non-binary Associate’s Degree
Appendix C:
Statistical Analysis
This section provides details about the statistical analysis performed on the survey data, including tables or graphs that illustrate the results of the analysis.
Table 1: Frequency of Social Media Platforms
Use Platform Frequency
- Facebook 35%
- Instagram 28%
- Twitter 15%
- Snapchat 12%
Figure 1: Impact of Social Media on Life Satisfaction
Appendix D:
Survey Results
This section presents the raw data collected from the survey, such as participant responses to each question.
Question 1: What is your age?
Question 2: What is your gender?
And so on for each question in the survey.
How to Write Appendices
Here are the steps to follow to write appendices:
- Determine what information to include: Before you start writing your appendices, decide what information you want to include. This may include tables, figures, graphs, charts, photographs, or other types of data that support the main content of your paper.
- Organize the material: Once you have decided what to include, organize the material in a logical manner that follows the sequence of the main content. Use clear headings and subheadings to make it easy for readers to navigate through the appendices.
- Label the appendices: Label each appendix with a capital letter (e.g., “Appendix A,” “Appendix B,” etc.) and provide a brief descriptive title that summarizes the content.
- F ormat the appendices: Follow the same formatting style as the rest of your paper or report. Use the same font, margins, and spacing to maintain consistency.
- Provide detailed explanations: Make sure to provide detailed explanations of any data, charts, graphs, or other information included in the appendices so that readers can understand the significance of the material.
- Cross-reference the appendices: In the main text, cross-reference the appendices where appropriate by referring to the appendix letter and title (e.g., “see Appendix A for more information”).
- Review and revise: Review and revise the appendices just as you would any other part of your paper or report to ensure that the information is accurate, clear, and relevant.
When to Write Appendices
Appendices are typically included in a document when additional information needs to be provided that is not essential to the main text, but still useful for readers who want to delve deeper into a topic. Here are some common situations where you might want to include appendices:
- Supporting data: If you have a lot of data that you want to include in your document, but it would make the main text too lengthy or confusing, you can include it in an appendix. This is especially useful for academic papers or reports.
- Additional examples: I f you want to include additional examples or case studies to support your argument or research, but they are not essential to the main text, you can include them in an appendix.
- Technical details: I f your document contains technical information that may be difficult for some readers to understand, you can include detailed explanations or diagrams in an appendix.
- Background information : If you want to provide background information on a topic that is not directly related to the main text, but may be helpful for readers, you can include it in an appendix.
Purpose of Appendices
The purposes of appendices include:
- Providing additional details: Appendices can be used to provide additional information that is too detailed or bulky to include in the main body of the document. For example, technical specifications, data tables, or lengthy survey results.
- Supporting evidence: Appendices can be used to provide supporting evidence for the arguments or claims made in the main body of the document. This can include supplementary graphs, charts, or other visual aids that help to clarify or support the text.
- Including legal documents: Appendices can be used to include legal documents that are referred to in the main body of the document, such as contracts, leases, or patent applications.
- Providing additional context: Appendices can be used to provide additional context or background information that is relevant to the main body of the document. For example, historical or cultural information, or a glossary of technical terms.
- Facilitating replication: In research papers, appendices are used to provide detailed information about the research methodology, raw data, or analysis procedures to facilitate replication of the study.
Advantages of Appendices
Some Advantages of Appendices are as follows:
- Saving Space: Including lengthy or detailed information in the main text of a document can make it appear cluttered and overwhelming. By placing this information in an appendix, it can be included without taking up valuable space in the main text.
- Convenience: Appendices can be used to provide supplementary information that is not essential to the main argument or discussion but may be of interest to some readers. By including this information in an appendix, readers can choose to read it or skip it, depending on their needs and interests.
- Organization: Appendices can be used to organize and present complex information in a clear and logical manner. This can make it easier for readers to understand and follow the main argument or discussion of the document.
- Compliance : In some cases, appendices may be required to comply with specific document formatting or regulatory requirements. For example, research papers may require appendices to provide detailed information on research methodology, data analysis, or technical procedures.
About the author
Muhammad Hassan
Researcher, Academic Writer, Web developer
You may also like

APA Research Paper Format – Example, Sample and...

Research Questions – Types, Examples and Writing...

Research Paper Outline – Types, Example, Template

Tables in Research Paper – Types, Creating Guide...

Thesis Statement – Examples, Writing Guide

Limitations in Research – Types, Examples and...
How to Write a Research Paper Appendix
Link Copied
Share on Facebook
Share on Twitter
Share on LinkedIn

Presentation aced!
Writing a research paper isn’t just a work of mere writing. Writing the perfect research paper takes a lot of research, analysis, framing, formatting, and much more. Correctly writing one of the most essential and academically popular segments of a research paper, the appendix, is one such effort that goes into a dissertation. In this blog , we will discuss with you the functions of an appendix in-depth and give you some tried and tested tips to craft the perfect appendix section of a research paper! Let’s dive in!
What is an Appendix?
The appendix on a research paper is a supplementary segment at the end of a dissertation or the research paper. This section isn’t considered a part of the main body text of the dissertation, but it is an important part of doing research. Appendices often feature raw data in the form of tables, figures, maps, diagrams and statistics and thus contribute to the credibility of the research and make it a perfect research paper .
Using academic resources, books, and research tools can help frame an appendix better. Appendices are essential since they provide extra support to your research and make the dissertation seem more transparent regarding data.
However, the appendix section of a research paper should only be supplementary; thus, you cannot depend on it to help the reader understand the main text. Your dissertation text should be detailed enough to be understandable without appendices, and they should only be placed to support your arguments presented in the research report.
How to Write an Appendix for a Research Paper
Writing the perfect research paper appendix can be overwhelming if it’s your first time doing so. However, drafting the appendix section of a research paper can be quite fun if you know the basics and understand how exactly you should go about it. Here are our 5 tips on how to write the perfect appendix for your dissertation:
Step 1: Organize the Appendix
With all the raw data, stats, and information, an appendix on a research paper can be difficult to go through and understand if they’re drafted disorganizedly. So, while writing your research paper appendix, make sure you are not just ramming all information into it but organising it well so the reader can utilise it. Structure it well, for it can very well come across as a reflection of your daily choices.
Step 2: Consider Accessibility
A research paper appendix can include non-textual information like tables, diagrams, graphs, images, illustrations, etc. If you’re adding such visual data elements to your appendices, ensure the material is clear and readable so the reader can comprehend the data. You should also ensure you are labelling these elements well and adding brief descriptions to each figure.
Step 3: Review for Relevance
It is easy to lose track of the relevance of your data while preparing appendices since you have to work with many different types of data simultaneously. However, you have to remember that the goal is not to stuff your appendices with data. Rather, craft a precise, careful research paper appendix that can give your reader relevant and additional data that supports your research.
Step 4: Proofread and Revise
When it comes to dissertation writing, typos, grammatical errors, and spelling mistakes can cost you way more than just miscommunication. These seemingly harmless errors can make your work look casual and unprofessional, bringing in questions about the credibility of your work. It is a similar case when it comes to writing an appendix for a research paper.
Step 5: Seek Guidance
It is important to remember that seeking guidance when you feel stuck is pretty normal, and there is nothing to be embarrassed about it. You may feel lost while writing an appendix for a research paper, and it is the perfect time to seek guidance from your peers, advisor or even dissertation committee members.
Explore student accommodations for a focused academic experience!
Book with amber today!
How to Format an Appendix
Ensuring proper formatting is crucial for the seamless integration of the research paper appendix into the main body. Follow the guidelines below for a sharp-looking appendix:
Consistency with the Main Body
Formatting elements, fonts, font sizes and margins should have uniformity. Consistent and professional appearance gives your research paper a neat look.
Organisation and Structure
Use headings and subheadings to categorise your data logically. You can also use a well-structured numbering system to facilitate easy navigation.
Descriptive Elements
Introduce each content with short descriptions and paragraphs. Giving additional context makes the information more accessible and interpretable.
Consistent Formatting Style
Use a formatting style that goes well with the rest of your dissertation, along with font styles, sizes, and other formatting guidelines instructed by your academic institution.
Visual Accessibility
Any non-textual elements, such as tables, graphs, or images, should be clear and readable. Label these visual elements and add alternative texts for inclusivity in the digital appendix.
Where does the appendix go in your dissertation?
Although the appendix section of a research paper is an essential part of your dissertation, it is not to be included in the main body of the dissertation. As a compilation of supplementary material and raw data, your research paper appendix should go at the end of the dissertation, typically inserted after the reference lists. Some even present appendices as separate supplementary documents, mostly done in specially requested cases.
The format of the research paper appendix should be similar to the rest of your report for consistency. It should thus be drafted and formatted in the same style as the dissertation in terms of fonts, margins, and font sizes.
What to include in your appendix
While drafting your research paper appendix, remember that it needs to be as precise as possible. Thus, there cannot be unnecessary information in it. Typically, appendices include raw data that supports your research and is referenced in the dissertation you have prepared. Here are some of the elements that you should include in your appendix:
- Research results
- Transcribed interviews
- Survey/questionnaire details
- Table and figures
- Co-respondence
- List of abbreviations used
- Calculations and formulas
Referring appendix in-text
Only adding your appendix to the research paper at the end of the dissertation would not make sense if there are no references to them in the main text. To justify its existence and inclusion in the research report, you should reference the appendix at least once in the whole report. A neatly labelled and properly referred research paper appendix can make your dissertation look more professional and supported.
How to refer to an appendix
Referring to the research paper appendix within the main text is important in highlighting its relevance. Use these five methods for referencing:
In-text references
Specific references embedded in your sentences contextually shape your information. For example, "In Table 2 of Appendix B, the commonality between subjects A and B is illustrated.
Parenthetical references
You can use parentheses for concise references without disrupting the main text's flow. For instance, "The result [refer to Appendix C, Fig. 2] is not consistent with the previous findings."
Referring to the entire appendix
Refer to the entire research paper appendix in your text when appropriate. For example, "The data supporting this conclusion can be found in Appendix B."
Clarity and labelling
References should be clear and well-labelled. Proper labelling ensures easy identification of referenced material within the appendix, polishing your research paper professionally.
Cross-referencing
Cross-referencing helps you establish connections between the main text and the appendix. Phrases like "As discussed in Appendix A" guide readers to supporting material.
Crafting the perfect appendix section of a research paper involves meticulous attention to detail and adherence to formatting and referencing guidelines. As an integral part of your dissertation, the appendix contributes significantly to the transparency, credibility, and overall professionalism of your research. By following the comprehensive guidelines provided in this guide, you can ensure that your appendix not only complements your main text but also serves as a valuable resource for readers seeking additional insights.
Frequently Asked Questions
What do i write in a research paper appendix, why is an appendix important for a dissertation, where is the appendix placed in the research paper, is writing a research paper appendix difficult, what are the basic guidelines for writing an appendix.
Your ideal student home & a flight ticket awaits
Follow us on :

Related Posts

17 Reasons to Study Abroad

600+ EPQ Ideas to Secure an A* Grade

UCAS Application 2024-25: UCAS Fee, Deadlines, Process

amber © 2024. All rights reserved.
4.8/5 on Trustpilot
Rated as "Excellent" • 4800+ Reviews by students
Rated as "Excellent" • 4800+ Reviews by Students

APA 7th edition - Paper Format: Appendices
- Introduction
- Mechanics of Style
- Overall Paper
- Sample Papers
- Reference List
- Tables and Figures
- APA Citations This link opens in a new window
- Additional Resources
- Writing Skills This link opens in a new window
How to Format An Appendix - Tutorial
- APA Appendices - JIBC Tip Sheet All you need to know about appendices in APA Style.
Information in this section is as outlined in the APA Publication Manual (2020), sections 2.14, 2.17, 2.24, and 7.6.
Appendices are used to include information that supplement the paper’s content but are considered distracting or inappropriate for the overall topic. It is recommended to only include an appendix if it helps the reader comprehend the study or theoretical argument being made. It is best if the material included is brief and easily presented. The material can be text, tables, figures, or a combination of these three.
Placement :
Appendices should be placed on a separate page at the end of your paper after the references, footnotes, tables, and figure. The label and title should be centre aligned. The contents of the appendix and the note should be left-aligned.
- If you are choosing to include tables and figures in your appendix, then you can list each one on a separate page or you may include multiple tables/figures in one appendix, if there is no text and each table and/or figure has its own clear number and title within the appendix.
- Tables and figures in an appendix receive a number preceded by the letter of the appendix in which it appears, e.g. Table A1 is the first table in Appendix A or of a sole appendix that is not labeled with a letter.
The follow elements are required for appendices in APA Style:
Appendix Labels:
Each appendix that you place in your paper is labelled “Appendix.” If a paper has more than one appendix, then label each with a capital letter in the order the appendices are referred to in your paper (“Appendix A” is referred to first, “Appendix B” is referred to second, etc).
- The label of the appendix should be in bold font, centre-aligned, follow Title Casing, and is located at the top of the page.
- If your appendix only contains one table or figure (and no text), then the appendix label takes the place of the table/figure number, e.g. the table may be referred to as “Appendix B” rather than “Table B1.”
Appendix Titles:
Each appendix should have a title, that describes its contents. Titles should be brief, clear, and explanatory.
- The title of the appendix should be in bold font, centre-aligned, follow Title Casing, and is one double-spaced line down from the appendix label.
- If your appendix only contains one table or figure (and no text), then the appendix title takes the place of the table/figure title.
Appendix Contents:
- Left aligned and indented; written the same as paragraphs within the body of the paper
- Double-spaced and with the same font as the rest of the paper
- If the appendix contains a table and/or figure, then the table/figure number must contain a letter to correlate the table and/or figure to the appendix and not the body of the paper, e.g. “Table A1” rather than “Table 1” to clarify that the table appears in the appendix and not in the body of the paper.
- All tables and figures in an appendix must be mentioned in the appendix and numbered in order of mention.
- All tables and figures must be aligned to the left margin, (not center aligned), and positioned after a paragraph break, preferably the paragraph in which they are referred to, with a double-spaced blank line between the table and the text.
- Each table and figure should include a note afterwards to further explain the supplement or clarify information in the table or figure to your paper/appendix and can be general, specific, and probability. See “Table Notes” in the section “Table and Figures” above for more details.
Referring to Appendices in the Text:
In your paper, refer to every appendix that you have inserted. Do not include an appendix in your work that you do not clearly explain in relation to the ideas in your paper.
- In general, only refer to the appendix by the label (“Appendix” or “Appendix A” etc.) and not the appendix title.
Reprinting or Adapting:
If you did not create the content in the appendix yourself, for instance if you found a figure on the internet, you must include a copyright attribution in a note below the figure.
- A copyright attribution is used instead of an in-text citation.
- Each work should also be listed in the reference list.
Please see pages 390-391 in the Manual for example copyright attributions.
- << Previous: Tables and Figures
- Next: Workshops >>
- Last Updated: May 1, 2024 11:05 AM
- URL: https://libguides.jibc.ca/apa/formatyourpaper
- Research Paper Guides
- Basics of Research Paper Writing
- How to Write an Appendix: Step-by-Step Guide & Examples
- Speech Topics
- Basics of Essay Writing
- Essay Topics
- Other Essays
- Main Academic Essays
- Research Paper Topics
- Miscellaneous
- Chicago/ Turabian
- Data & Statistics
- Methodology
- Admission Writing Tips
- Admission Advice
- Other Guides
- Student Life
- Studying Tips
- Understanding Plagiarism
- Academic Writing Tips
- Basics of Dissertation & Thesis Writing
- Essay Guides
- Formatting Guides
- Basics of Research Process
- Admission Guides
- Dissertation & Thesis Guides
How to Write an Appendix: Step-by-Step Guide & Examples

Table of contents
Use our free Readability checker
While composing your work, you may stumble upon a question on how to write an appendix.
An appendix is a supplemental section of a research paper that provides additional information, data, or materials to support the main content. The appendix is usually placed at the end of the document and is numbered with letters or numbers, such as "Appendix A," "Appendix B," etc. The purpose of an appendix is to provide readers with supplementary details that are not included in the main text but are relevant to the topic.
Once you decide on writing appendices, you should collect additional information and format your text as required. Here, we will talk about how you can work with appendices. We will also show some nuances of their preparation process using a real example. Is the deadline around the corner? Consider using professional research paper help from expert scholars.
What Is an Appendix: Definition
Experienced researchers know what an appendix in a paper is. But aspiring authors often have problems with this section of the work. First of all, you should understand that appendices are an additional section of a dissertation or any other scientific paper that includes additional information. Main points are not placed in an appendix meanwhile at the end of your work it can expand on some context or clarify author’s position on a particular issue. Also, an appendix is often placed after the citation page of a work. It is indicated with the help of references in a main text.
What Is the Purpose of an Appendix
Quite often, authors don’t understand the purpose of an appendix. This usually looks like a table and is not included in a main text. Remember that content of your dissertation should be concise and clear. It is also undesirable if you deviate from your theme so as not to confuse readers. Therefore, you can provide a reference, which will lead a reader to an appendix of a thesis. Typically, the purpose of an appendix is to extra information that is usually not included in the text's body. It expresses author's point of view, and provides additional information. It may not address the immediate topic of your dissertation or expand on current research. As a reminder, your work should be clear even without studying an appendix. So make sure you don't put important details there.
What Can You Include in an Appendix
An appendix in a paper is a supplement to a main text, not a replacement. You can put different elements there. It is better if you separate appendices, highlighting one element in each of them. Don’t forget about separate references in your text. Otherwise it will be difficult for a reader to understand your information better. Thus, the following information can be added:
- diagrams with illustrative figures;
- abbreviations ;
- interviews;
- statistics, and much more.
There are no restrictions on content added to your dissertation's appendices. Theoretically, you can attach absolutely any information that is relevant to your topic. Thus, possibilities for evidence base are almost unlimited. All you need to do is add tables or any other information.
How to Write an Appendix: Full Guide
If you already have experience working on dissertations and other scientific texts, you will not wonder how to make an appendix. However, it is still important that you get some advice on how to properly structure an appendices section. This will help add information that may be redundant in the main part of your paper. We offer 4 simple steps to create an informative and readable appendix block.
Step 1. Make an Appendix: Include Your Data
When creating an appendix, include extra data in their raw form. That is, you might not have used some details in your main paper. But you want a reader to know more information. For example, it can be calculations, some results of which are mentioned in your main text. Or maybe, you can add some statistics that clearly demonstrate your research paper conclusion . You can also include facts from other scientific sources that support your position. One thing is important — information should complement your text but not contradict it.
Step 2. Include Visual Supporting Documents in an Appendix
When you are writing an appendix, you can’t avoid visual additions that clearly demonstrate an information and save an author from lengthy descriptions in the text. Should you need to support your conclusions drawn in the scientific text, these can be used:
Don’t forget: you should quote and indicate the authorship of graphics used in your work. If you took it from any third-party sources, of course. Thus, a reader will be able to find additional data that explains the content of your text. It is good if you personally put results of your research in a graphic form. To do this, you can use Office programs, graphic editors and other programs available to PC users.
Step 3. Describe the Instruments of Your Research in Your Appendices
It is good if your appendix in the research paper has a section for indicating tools that were used during the preparation of your dissertation writing . This way, your reader will understand how you collected information and do it themselves. For example, it could be a dictaphone or tape recorder on which an interview with your expert was recorded. Or you might have used a video camera for recording facts and interviews. In such case, it is advisable to indicate these instruments in your appendix. Specialized equipment for measuring, calculating and making graphics should also be added at the beginning of the appendix. This way, you will demonstrate your skills and knowledge. Research units don’t require extra tools, so make sure they are listed. You can do it even in a short format.
Step 4. Include an Interview and Transcripts in an Appendix
When conducting interviews and surveys for collecting information, make an appendix with photocopies of handwritten materials or electronic copies of digital surveys. Their order is not important. The main thing is that your research text contains references. This will allow you to quickly study the sources. You should not only show that the source contains important data but also explain it. So, even additional content, including questions and answers, needs to be listed. But if you originally had a readable format, you don’t need to do this. In addition to interviews, also add screenshots or photos of correspondences used for surveys. For example, you can refer to a significant researcher with whom you exchanged letters. Or maybe you studied subject, together with this researcher, and they gave some comments on a particular issue. Do not know how to write a discussion section of a research paper ? Do not worry, we have the whole article dedicated to this topic.
Formatting an Appendix: Main Rules
Formatting of appendices is required in any case. First of all, provide correct citations. APA, MLA, and Chicago are the most commonly used standards. Although, you should clarify what formatting requirements your institution has. Correct formatting includes:
- Appendix title. Write it at the top of the content page, indicate its title, using letters or numbers for ordering.
- Sorted by mention. Don’t add appendices randomly, it is better to do it in chronological order. That is, as information from it is given in main text.
- Location after bibliography. This is a general requirement that cannot always be met. For example, if your professor wants the appendices to be put before the bibliography, this will have to be done.
- Page numbers. All dissertation pages should be numbered, even if they are blank. This will make the appendix block the part of main text.
Also, review your appendix before approval. Make sure that its content is clear, error-free, and correctly quoted.
Appendix Example
To do the job successfully, it is recommended to have an example of an appendix at hand. Without it, there are usually problems with a choice of font and mentions that appear in main text. We will show you what the appendix itself looks like at the end of the dissertation using a short interview as an example.

We have one more blog in case you wonder what is an abstract in a paper or need some examples and writing tips.
How to Make an Appendix: Final Thoughts
Thus, we talked about how to write an appendix. It allows you to include additional details, while avoiding writing them in the body of your text. To do this, one can use graphics, transcriptions of conversations, tables and statistics — anything that complements your research. Be sure to clarify formatting requirements of your university. Arrange appendices in an order in which they appear in your text. Try to use your own materials and not take other people's work. In case of unique findings, they can be used in your work.
Please contact us if you have any difficulties preparing an academic work! Our professional paper writers guarantee high quality and loyal prices. Just choose a writer to your liking, send your requirements and you're good to go!
Frequently Asked Questions About Appendix Writing
1. how do you add an appendix to an essay.
The inclusion of appendix to an essay is the same as to any other paper. You need to provide references in your text of an essay itself, as well as submit attachments after a bibliography. Don't forget to specify name of an appendix for easy navigation.
2. Do I add references to the appendix?
Yes, this is not only recommended but must be done. In this case the appendix will allow your reader to check the reliability of sources you used. Moreover, if you took any information from third-party sources, this protect you from plagiarism charges.
4. How do you create an appendix in Word?
It is not difficult to prepare an appendix in Word, because this Office program contains all the necessary tools. To get started, choose the same font, font size and indentation that were used in the main text, so as not to visually break away from it. We also recommend that you apply title formatting with built-in Word tools. Place the appendix titles at the top in the center of a page. In this case it will be much easier to navigate the paper.
3. What is an appendix in a report example?
You can include a wide range of information into an appendix in a report. It is better to opt for descriptive formats, though. For example, it can be graphical or mathematical research results, statistics of a certain phenomenon, and questionnaires filled in by other people.

Joe Eckel is an expert on Dissertations writing. He makes sure that each student gets precious insights on composing A-grade academic writing.
You may also like

- PRO Courses Guides New Tech Help Pro Expert Videos About wikiHow Pro Upgrade Sign In
- EDIT Edit this Article
- EXPLORE Tech Help Pro About Us Random Article Quizzes Request a New Article Community Dashboard This Or That Game Happiness Hub Popular Categories Arts and Entertainment Artwork Books Movies Computers and Electronics Computers Phone Skills Technology Hacks Health Men's Health Mental Health Women's Health Relationships Dating Love Relationship Issues Hobbies and Crafts Crafts Drawing Games Education & Communication Communication Skills Personal Development Studying Personal Care and Style Fashion Hair Care Personal Hygiene Youth Personal Care School Stuff Dating All Categories Arts and Entertainment Finance and Business Home and Garden Relationship Quizzes Cars & Other Vehicles Food and Entertaining Personal Care and Style Sports and Fitness Computers and Electronics Health Pets and Animals Travel Education & Communication Hobbies and Crafts Philosophy and Religion Work World Family Life Holidays and Traditions Relationships Youth
- Browse Articles
- Learn Something New
- Quizzes Hot
- Happiness Hub
- This Or That Game
- Train Your Brain
- Explore More
- Support wikiHow
- About wikiHow
- Log in / Sign up
- Education and Communications
- Technical Writing
How to Write an Appendix
Last Updated: October 4, 2023
This article was co-authored by Stephanie Wong Ken, MFA . Stephanie Wong Ken is a writer based in Canada. Stephanie's writing has appeared in Joyland, Catapult, Pithead Chapel, Cosmonaut's Avenue, and other publications. She holds an MFA in Fiction and Creative Writing from Portland State University. This article has been viewed 1,740,135 times.
Like the appendix in a human body, an appendix contains information that is supplementary and not strictly necessary to the main body of the writing. An appendix may include a reference section for the reader, a summary of the raw data or extra details on the method behind the work. You may be required to write an appendix for school or you may decide to write an appendix for a personal project you are working on. You should start by collecting content for the appendix and by formatting the appendix properly. You should then polish the appendix so it is accessible, useful, and engaging for your reader.
Collecting Content for the Appendix

- Raw data may include sample calculations that you refer to in the body of the paper as well as specialized data that expands on data or information you discuss in the paper. Raw statistical data can also be included in the appendix.
- You may also include contributory facts from other sources that will help to support your findings in the paper. Make sure you properly cite any information you are pulling from other sources.

- You may include graphs or charts you have created yourself or graphs or charts from another source. Make sure you properly cite any visuals that are not your own in the appendix.

- For example, you may note in the appendix: “All interviews and surveys were conducted in person in a private setting and were recorded with a tape recorder.”

- You should also include any correspondences you had with subjects in your research, such as copies of emails, letters, or notes written to or from your research subjects.
Formatting the Appendix

- If you have more than one appendix, order them by letter or number and be consistent about the ordering. For example, if you are using letters, make sure the appendices are titled “Appendix A,” “Appendix B,” etc. If you are using numbers, make sure the appendices are titled “Appendix 1,” “Appendix 2,” etc.
- If you have more than one appendix, make sure each appendix begins on a new page. This will ensure the reader is not confused as to where one appendix ends and another begins.

- For example, if raw data is mentioned in the first line of your paper, place that raw data first in your appendix. Or if you mention interview questions at the very end of your paper, make sure the interview questions appear as the last point in your appendix.

- You should also make sure you list the appendix in your table of contents for the paper, if you have one. You can list it based on title, for example, “Appendix”, or “Appendix A” if you have more than one appendix.

- For example, if the text ends on page 17, continue numbering from page 17 when you put in the page numbers for the appendix.
Polishing the Appendix

- You may find it helpful to have someone else read through the appendix, such as a peer or a mentor. Ask them if they feel all the included information is relevant to the paper and remove any information they deem unnecessary.

- Read through the appendix backwards so you can make sure there are no spelling errors. You want the appendix to appear as professional as possible.

- For example, you may note an appendix in the text with: “My research produced the same results in both cases (see Appendix for raw data)” or “I feel my research was conclusive (see Appendix A for interview notes).”
Sample Appendices

Community Q&A

You Might Also Like

- ↑ https://libguides.usc.edu/writingguide/appendices
- ↑ http://libguides.usc.edu/writingguide/appendices
- ↑ https://askus.library.wwu.edu/faq/116707
About This Article

Medical Disclaimer
The content of this article is not intended to be a substitute for professional medical advice, examination, diagnosis, or treatment. You should always contact your doctor or other qualified healthcare professional before starting, changing, or stopping any kind of health treatment.
Read More...
To write an appendix, start by writing “Appendix” at the top of the document, using the same font you used for your chapter headings. Then, order the contents, such as graphs, surveys, or interview transcripts, based on the order in which they appear in your paper. Next, number the pages so they follow sequentially, coming after your paper and your reference list or list of sources. Finally, make sure to check for spelling and grammar errors, so everything will look polished and professional. For more tips from our English co-author, including how to refer to the appendix in your paper, keep reading! Did this summary help you? Yes No
- Send fan mail to authors
Reader Success Stories
Apr 20, 2018

Did this article help you?

Apr 10, 2017
Stella Cheboi
Dec 4, 2017
Daniella Saarn
May 20, 2018
Apr 5, 2017

Featured Articles

Trending Articles

Watch Articles

- Terms of Use
- Privacy Policy
- Do Not Sell or Share My Info
- Not Selling Info
Don’t miss out! Sign up for
wikiHow’s newsletter
- Essay Editor
What is an Appendix in a Research Paper? | Aithor

What is an appendix in a research paper?
An appendix is a supplementary section at the end of a research paper after the list of references. It contains more information that helps explain the main ideas in the paper. It's not needed in the main part because it would make the paper too long or go “off-topic." The appendix gives readers more details to help them understand the research better without making the main argument hard to follow.
The length of an appendix can differ depending on what kind of research it is and how much extra information there is. But it shouldn't be more than 10-15% of the total number of pages. An appendix document can have many types of information, like tables, figures, charts, graphs, images, interview notes, survey questions, or anything else that helps support the research findings.
What type of information does a research paper appendix include?
A research paper appendix can include a wide range of supporting materials, such as:
- Raw data sets or statistical tables that are too extensive for the main text
- Detailed descriptions of research methodologies, instruments, or protocols
- Interview transcripts or survey questionnaires
- Correspondence with research participants or collaborators
- Visual aids, such as graphs, charts, images, or diagrams
- Glossaries or abbreviation lists
- Copies of relevant documents, such as consent forms or legal agreements
The specific content of an appendix depends on the nature of the research and the requirements of the academic discipline or publication venue.
How to structure an appendix
When considering how to write an appendix, follow these guidelines:
- Place the appendix after the references list, starting on a new page
- Use a clear and descriptive title, such as "Appendix A: Survey Questionnaire"
- Organize the content logically and label each item systematically (e.g., Table A1, Figure B2)
- Refer to each appendix item in the main text using parenthetical citations (e.g., "(see Appendix A)")
- Use consistent formatting throughout the appendix, following the style guide requirements (e.g., APA, MLA, and Chicago)
Remember, the goal is to make it easy for readers to find and understand the extra information without removing the main points you're making.
General rules for completing appendices
In addition to the basic structure, there are some general rules to follow when making appendices. The exact rules might be a little different depending on the citation style you're using, but here are some common ones:
Appendix APA format
The APA format is the most popular at colleges and universities. When using APA format for your appendices, there are a few specific rules to keep in mind:
- Use the heading "Appendix" followed by a letter (A, B, C) for each distinct appendix
- Center the appendix title beneath the heading
- Arrange appendices in the order they are mentioned in the main text
- Start each appendix on a new page, regardless of its length
- Use double spacing and indent the first line of each paragraph
- Include page numbers and place the appendix after the references list
Appendix MLA format
The rules for MLA format are similar to APA, but the difference is that the MLA appendix should be placed before the list of references. Here are some requirements for MLA format:
- Place the appendix before the references list
- Use the heading "Appendix," followed by a letter for each distinct appendix
- Start each appendix on a new page
- Use double spacing and a hanging indent for each entry
- Italicize titles of standalone sources (e.g., books, websites)
Appendix Chicago style
Used when assigned academic papers on History. Here are some requirements for Chicago style:
- Use the plural heading "Appendices" for multiple appendices
- Use 12-point Times New Roman font
- Number pages in the top right corner, starting with "Page 1"
- Omit the page number on the title page
- Place appendices before the bibliography
No matter which citation style you use, the most important thing is to be consistent and clear when labeling and referencing your appendices.
How do I refer to an appendix?
To refer to an appendix in the main text, follow these guidelines:
- Mention each appendix at least once in the text, using a parenthetical citation (e.g., "(see Appendix A)")
- Capitalize "Appendix" when referring to a specific appendix (e.g., "As shown in Appendix B")
- Use lowercase when referring to appendices in general (e.g., "The appendices contain additional data")
- Be consistent in your references throughout the paper
For example:
- "The survey questionnaire (see Appendix A ) was distributed to 100 participants."
- " Figure B3 in Appendix B illustrates the relationship between variables X and Y."
- "More information on the parallel between both eras can be found in Appendix C ."
Remember to consult your chosen style guide for specific formatting requirements and guidelines on how to write an appendix and appendix format.
When you're writing a research paper, an appendix can be a helpful way to provide more information supporting your paper's main ideas. By following some simple rules for organizing and mentioning appendices, you can share extra data and back up your points without making the main part of your paper too long or hard to understand.
Want to make writing your research paper as prestigious as possible? Aithor can help make your life easier! This helpful essay-writing tool uses AI to assist you in creating academic and creative works that meet your specific needs in just a short time.
Give Aithor a try today and see how it can help make writing your academic papers a better experience!
Related articles
How to write a dialogue in an essay: useful tips.
A correct usage of dialogues in essays may seem quite difficult at first sight. Still there are special issues, for instance, narrative or descriptive papers, where this literary technique will be a good helper in depicting anyone's character. How to add dialogues to the work? How to format them correctly? Let's discuss all relevant matters to master putting conversation episodes into academic essays. Essay Dialogue: Definition & Purpose A dialogue is a literary technique for presenting a con ...
What Is Self-Plagiarism & How To Avoid It
Have you ever thought about whether using your own work again could be seen as copying? It might seem strange, but self-plagiarism is a real issue in school and work writing. Let's look at what this means and learn how to avoid self-plagiarism so your work stays original and ethical. What is self-plagiarism? Self-plagiarism, also called auto-plagiarism or duplicate plagiarism, happens when a writer uses parts of their old work without saying where it came from. This isn't just about copying w ...
Paraphrasing vs Plagiarism: Do They Really Differ?
Academic assignments require much knowledge and skill. One of the most important points is rendering and interpreting material one has ever studied. A person should avoid presenting word-for-word plagiarism but express his or her thoughts and ideas as much as possible. However, every fine research is certain to be based on the previous issues, data given, or concepts suggested. And here it's high time to differentiate plagiarism and paraphrasing, to realize its peculiarities and cases of usage. ...
How To Write Essays Faster Using AI?
Creating various topical texts is an obligatory assignment during studies. For a majority of students, it seems like a real headache. It is quite difficult to write a smooth and complex work, meeting all the professors' requirements. However, thanks to modern technologies there appeared a good way of getting a decent project – using AI to write essays. We'd like to acquaint you with Aithor, an effective tool of this kind, able to perform fine and elaborated texts, and, of course, inspiration, i ...
Can Plagiarism Be Detected on PDF?
Plagiarism has been a challenge for a long time in writing. It's easy to find information online, which might make some people use it without saying where it came from. But plagiarism isn't just taking someone else's words. Sometimes, we might do it by accident or even use our own old work without mentioning it. When people plagiarize, they can get into serious trouble. They might lose others' trust or even face legal problems. Luckily, we now have tools to detect plagiarism. But what about PDF ...
Top 10 Use Cases for AI Writers
Writing is changing a lot because of AI. But don't worry — AI won't take human writers' jobs. It's a tool that can make our work easier and help us write better. When we use AI along with our own skills, we can create good content faster and better. AI can help with many parts of writing, from coming up with ideas to fixing the final version. Let's look at the top 10 ways how to use AI for content creation and how it can make your writing better. What Is AI Content Writing? AI content writin ...
Plagiarism: 7 Types in Detail
Your professor says that it is necessary to avoid plagiarism when writing a research paper, essay, or any project based on the works of other people, so to say, any reference source. But what does plagiarism mean? What types of it exist? And how to formulate the material to get rid of potential bad consequences while rendering original texts? Today we try to answer these very questions. Plagiarism: Aspect in Brief Plagiarism is considered to be a serious breach, able to spoil your successful ...
What is Citation and Why Should You Cite the Sources When Writing Content
When we write something for school, work, or just for fun, we often use ideas and facts from other places. This makes us ask: what is a citation in writing? Let's find out what this means and why it's really important when we write. What is Citation? Citation in research refers to the practice of telling your readers where you got your information, ideas, or exact words from. It's like showing them the path to the original information you used in your writing. When you cite something, you us ...
Verify originality of an essay
Get ideas for your paper
Cite sources with ease
Guide to What is an Appendix in a Research Paper: Structure and Format
Updated 09 Jul 2024

Completing academic assignments requires understanding the basic concepts. One of the questions students often ask is, “What is an appendix in a paper?” This part of the text improves a reader's understanding of a research paper without adding excessive length to the final result. It also provides detailed information about your topic without interrupting the flow of your thoughts with too many tables and figures.
To learn more about writing an appendix in a research paper according to specific formatting styles, such as APA, Chicago, or MLA, read this article from Edubirdie.
What is an appendix in a research paper?
The definition of this term is simple.
An appendix is an academic work section that contains additional information (statistics, references, tables, figures, etc.) that cannot be included in the main text.
This component is usually placed after the reference list at the end of a research paper or dissertation. The purpose of this text component is to provide additional information that may need to be explained fully in the main body. It’s a useful tool for giving context and clarity to the reader about the subject matter analyzed in the paper.
The length of an appendix can vary depending on the type of research, the amount of data collected, and the academic institution’s requirements. Generally, it should be as long as necessary to provide important and relevant information to support arguments and answer the research question . This element should not be too long or contain irrelevant data, as it can distract the audience. Usually, the appendix doesn’t exceed 10-15% of the total research pages.
What type of information does a research paper appendix include?
An appendix is a means of providing additional data that can further illustrate the research topic. As a result, the information included in an appendix section of a research paper can take various forms. Let’s see them in detail.
Surveys are frequently used in research methodology and are often included in appendices. To provide readers with a clear understanding of the findings, you should add surveys precisely as given to the respondents, along with their exact answers.
Whether transcribed or recorded, interviews are typically included in appendices. To ensure transparency, you should include the full list of questions and the corresponding answers.
- Correspondence
Researchers often add correspondence with collaborators related to the research subject in their papers as an appendix. These can include text messages, emails, transcripts of audio messages, letters, and other forms of communication.
- Research Tools
Cameras, audio recorders, specialized software, and other research tools should be acknowledged in an appendix to provide readers with insight into the research process.
- Non-Textual Items
Non-textual items such as tables, graphs, illustrations, figures, charts, or excessively numerous photos should be included as appendices.
- Statistical Data
Statistical data that is too extensive should be added to the research as an appendix, even if only a portion of the data was used. It’s also necessary to provide complete data.
How to structure an appendix
When you understand a research paper definition learning its writing style and structure is crucial. Maintaining the academic writing style and presenting information concisely and scientifically enhance the work's credibility. It applies to raw information and public discussion results, interview transcripts, summarized evaluation results, and copies of private letters. To ensure a smooth and effective organization of the appendix, it is recommended to learn expert tips on writing a research paper outline and create a separate appendix for each part of the paper.
Appendices can include tables, text, footnotes, and other supporting items that may be useful for readers, and each item should be titled. Each appendix should be mentioned at least once in the document. Selecting and using reliable sources, figures, and authors is important to create a credible research paper. The same principles apply to the appendices, which may be placed before or after the list of references depending on the requirements and formatting style.
General rules for completing appendices
When you create an appendix, it's essential to follow particular guidelines. It's also recommended to consult the citation format requirements for details before starting your research work.
Let’s see some general rules for creating an appendix:
- A common heading for an appendix is “Appendix 1 or A,” in bold and centered, with a title that describes its content.
- It's recommended to divide it by a set of data or a topic, which should be indicated in the table of contents.
- The appendix should be located before or after the list of references and should start on a new page with a page number.
- Lastly, the appendices should be arranged in sequential order based on the references made in the paper.
If you find it complicated and are thinking, "It's better to pay someone to write my research paper ," seeking professional assistance can be particularly helpful, especially when it comes to creating an appendix.
There are three formats for organizing research papers appendices, which your professor may require you to follow. Although similar, they have distinct features and rules that must be followed. Let’s see them in detail.
Appendix APA format
This format is the most popular at colleges and universities, and it’s usually required for academic papers on Social Sciences, such as Psychology, Education, Sociology, Criminology, and others. Many professors often ask students to produce their assignments in this style. Following the guidelines is important to ensure the structure and information are correct. These are the key points that professors look for when a paper is required to be written in APA format:
- Create the heading "Appendix", which should be followed by A, B, C, etc.;
- Write the appendix title centered under the heading;
- Follow the order of information stated in the paper;
- Indicate page numbers;
- Start each appendix from a new page, even if it’s smaller than the page size;
- Use double spacing;
- Write the first paragraph without indentation, while the rest should be indented.
- Add footnotes;
- Place the appendix just after the reference list;
- Include “see Appendix A” after the text to reference an appendix in the body of your thesis.
You should learn the general guidelines on this format or note them.
Appendix MLA format
The MLA style is recommended when researching the Humanities, like Philosophy, Languages, and Arts. This format is very similar to the APA format, but a few differences exist. The essential peculiarity is that the MLA appendix should be placed before the list of references. Here are some requirements for MLA format:
- Insert the appendix after the main document body and before the reference list;
- Use A, B, and C when writing headings for several appendices;
- Center the title;
- Create an appendix following the order of information stated in the research work;
- Add page numbers for every appendix;
- Place each appendix on its page, no matter its size;
- Double-space the list;
- Use a “hanging indent” format, where the first line is in the left margin, and every subsequent line is indented;
- Use italics for the titles of Internet sites, complete writings, books, and recordings when you use them in your appendices;
- Do not use italics for reference titles that only refer to a part of a source, such as short papers, poetry, tabloids, scholarly entries, sections of a PDF document, etc.
Appendix Chicago style
Consider this format if you’re assigned academic papers on History. It’s also required for academic journals and books. Creating research papers in Chicago style is not more difficult than in APA format. These two formats are almost identical. Still, there is a slight difference. Look at the guidelines for writing an appendix in a research paper in Chicago style:
- Create the title “Appendices” to describe more than one appendix;
- Use Times New Roman font with a 12-point text size;
- Place page numbers on the top right of every page labeled as "Page 1, 2, 3," etc.;
- Do not indicate the number of the page on the front cover;
- The appendices should be placed before the bibliography, which should come with footnotes and be on a separate page finalizing your research work.
Feel free to look at samples formatted according to the Chicago style requirements. While completing appendices, you’ll find many useful things to implement in your papers.
How to refer to an appendix?
If you’re looking for an answer to the question, “What is an appendix in a paper?” you’ll definitely find a lot of useful recommendations about how to refer to appendices. It’s necessary to mention every appendix at least once in a document. It can be done by stating the appendix's letter or number within the sentence or in parentheses. You can also refer to a specific component within an appendix. Let’s see an example of an appendix in a research paper.
Example 1. Referring to the whole appendix:
As shown in Appendix A, the participants' demographic information indicates that…
In the table (see Appendix B), you’ll see…
Example 2. Referring to an appendix component:
These data (see Appendix 3, Table 2) indicate that…
Photo 3 in Appendix 1 presents…
The word "Appendix" should be capitalized when referring to a particular appendix. It’s essential to ensure consistency throughout the paper. You should always use lowercase when you refer to appendices in general.
Different citation styles have specific formatting requirements and rules for appendices, particularly for APA Style and labeling figures and tables within the appendices. For detailed information, it's important to refer to the guidelines. Understanding what an appendix in a research paper is and how to format it correctly can be challenging, so some students choose to buy thesis paper to ensure all components are professionally compiled.
If you are unsure about the originality of your research paper, it is always a good idea to utilize a " check my paper for plagiarism " tool.
Where to place appendices in a research paper?
One way to place your appendix for a paper is to insert it after the main text with the citation references. In this case, you can proceed with the next page number. Another approach is to complete a separate document containing the appendices, which can be submitted along with your dissertation. It's important to remember that all appendices should be indicated in the table of contents of your thesis.
Does the appendix mean the same as references?
The appendix and references are different. An appendix is an extra material that can include tables, diagrams, or graphs that support the main text, while references are a list of sources cited or consulted in the main text. Both can be used to provide further information but serve different purposes and contain different types of information.
Is it possible to cite sources in an appendix?
Yes, using APA in-text citations in your appendix is acceptable if it's relevant. You should use an author-date citation format, the same way you do it in the main text. Remember to include all the cited sources in your reference list in your appendices. There is no need to complete an individual reference list for your appendices.
Is it appendices or appendixes?
Both "appendices" and "appendixes" are accepted plural forms of the word “appendix”. Still, "appendices" is the more commonly used plural form in academic writing. It’s also the preferred form in APA style. Using the same spelling throughout the whole document is essential.
Should I number my appendices in APA style?
In an APA guide about how to write an appendix for a research paper, there is a recommendation to label multiple appendices in sequential order using uppercase letters, such as “Appendix A,” “Appendix B,” etc. Still, if you only have one appendix, it should be titled simply as "Appendix" and mentioned as such in the document (for example, “see Appendix”).
What title to give to an appendix?
The typical title for an appendix is "Appendix A" or "Appendix 1," which should be centered and written in bold. Following the appendix number, a descriptive heading outlining the appendix content should be provided.
Was this helpful?
Thanks for your feedback.

Written by Meredith Anderson
Meredith, a dedicated editor at EduBirdie, specializes in academic writing. Her keen eye for grammar and structure ensures flawless papers, while her insightful feedback helps students improve their writing skills and achieve higher grades.
Related Blog Posts
How to write a research question: common types and winning examples.
The starting point for any investigation is a research question. Still, formulating valid and relevant questions can be challenging for many writer...
What is qualitative research? Approaches, methods, and examples
Students in social sciences frequently seek to understand how people feel, think, and behave in specific situations or relationships that evolve ov...
Delimitations in research: meaning, types, and examples
Working on academic papers can make it easy to feel overwhelmed by the huge amount of available data and information. One of the most crucial consi...
Join our 150K of happy users
- Get original papers written according to your instructions
- Save time for what matters most

Organizing Academic Research Papers: Appendices
- Purpose of Guide
- Design Flaws to Avoid
- Glossary of Research Terms
- Narrowing a Topic Idea
- Broadening a Topic Idea
- Extending the Timeliness of a Topic Idea
- Academic Writing Style
- Choosing a Title
- Making an Outline
- Paragraph Development
- Executive Summary
- Background Information
- The Research Problem/Question
- Theoretical Framework
- Citation Tracking
- Content Alert Services
- Evaluating Sources
- Primary Sources
- Secondary Sources
- Tertiary Sources
- What Is Scholarly vs. Popular?
- Qualitative Methods
- Quantitative Methods
- Using Non-Textual Elements
- Limitations of the Study
- Common Grammar Mistakes
- Avoiding Plagiarism
- Footnotes or Endnotes?
- Further Readings
- Annotated Bibliography
- Dealing with Nervousness
- Using Visual Aids
- Grading Someone Else's Paper
- How to Manage Group Projects
- Multiple Book Review Essay
- Reviewing Collected Essays
- About Informed Consent
- Writing Field Notes
- Writing a Policy Memo
- Writing a Research Proposal
- Acknowledgements
An appendix contains supplementary material that is not an essential part of the text itself but which may be helpful in providing a more comprehensive understanding of the research problem and/or is information which is too cumbersome to be included in the body of the paper. A separate appendix should be used for each distinct topic or set of data and always have a title descriptive of its contents .
Importance of...
Your research paper must be complete without the appendices, and it must contain all information including tables, diagrams, and results necessary to address the research problem. The key point to remember when you are writing an appendix is that the information is non-essential; if it were removed, the paper would still be understandable.
It is appropriate to include appendices...
- When the incorporation of material in the body of the work would make it poorly structured or it would be too long and detailed and
- To ensure inclusion of helpful, supporting, or essential material that would otherwise clutter or break up the narrative flow of the paper, or it would be distracting to the reader.
Structure and Writing Style
I. General Points to Consider
When considering whether to include content in an appendix, keep in mind the following points:
- It is usually good practice to include your raw data in an appendix, laying it out in a clear format so the reader can re-check your results. Another option if you have a large amount of raw data is to consider placing it online and note this as the appendix to your research paper.
- Any tables and figures included in the appendix should be numbered as a separate sequence from the main paper . Remember that appendices contain non-essential information that, if removed, would not diminish a reader's understanding of the overall research problem being investigated. This is why non-textual elements should not carry over the sequential numbering of elements in the paper.
- If you have more than three appendices, consider listing them on a separate page at the beginning of your paper . This will help the reader know before reading the paper what information is included in the appendices [always list the appendix or appendices in a table of contents].
- The appendix can be a good place to put maps, photographs, diagrams, and other non-textual elements , if you feel that it will help the reader to understand the content of your paper, but remembering that the paper should be understandable without them.
- An appendix should be streamlined and not loaded with a lot information . If you have a very long and complex appendix, it is a good idea to break it down into separate appendices, allowing the reader to find relevant information quickly.
II. Contents
Appendices may include some of the following, all of which should be referred to or summarized in the text of your paper:
- Supporting evidence [e.g. raw data]
- Contributory facts or specialized data [raw data appear in the appendix, but with summarized data appearing in the body of the text].
- Sample calculations
- Technical figures, graphs, tables, statistics
- Detailed description of research instruments
- Maps, charts, photographs, drawings
- Letters, emails, and other copies of correspondance
- Questionnaire/survey instruments, with the results appearing in the text
- Complete transcripts of interviews
- Complete field notes from observations
- Specification or data sheets
NOTE: Do not include vague or irrelevant information in an appendix; this additional information will not help the reader’s overall understanding and interpretation of your research and may only succeed in distracting the reader from understanding your research study.
III. Format
Here are some general guideline on how to format appendices, but consult the writing style guide [e.g., APA] your professor wants you to use for the class, if needed:
- Appendices may precede or follow your list of references.
- Each appendix begins on a new page.
- The order they are presented is dictated by the order they are mentioned in the text of your research paper.
- The heading should be "Appendix," followed by a letter or number [e.g., "Appendix A" or "Appendix 1"], centered and written in bold.
- Appendices must be listed in the table of contents [if used].
- The page number(s) of the appendix/appendices will continue on with the numbering from the last page of the text.
Appendices . The Structure, Format, Content, and Style of a Journal-Style Scientific Paper. Department of Biology. Bates College; Tables, Appendices, Footnotes and Endnotes . The Writing Lab and The OWL. Purdue University; Lunsford, Andrea A. and Robert Connors. The St. Martin's Handbook. New York: St. Martin's Press, 1989.
- << Previous: 9. The Conclusion
- Next: 10. Proofreading Your Paper >>
- Last Updated: Jul 18, 2023 11:58 AM
- URL: https://library.sacredheart.edu/c.php?g=29803
- QuickSearch
- Library Catalog
- Databases A-Z
- Publication Finder
- Course Reserves
- Citation Linker
- Digital Commons
- Our Website
Research Support
- Ask a Librarian
- Appointments
- Interlibrary Loan (ILL)
- Research Guides
- Databases by Subject
- Citation Help
Using the Library
- Reserve a Group Study Room
- Renew Books
- Honors Study Rooms
- Off-Campus Access
- Library Policies
- Library Technology
User Information
- Grad Students
- Online Students
- COVID-19 Updates
- Staff Directory
- News & Announcements
- Library Newsletter
My Accounts
- Interlibrary Loan
- Staff Site Login
FIND US ON
Reference management. Clean and simple.
What is an appendix in a paper

What is an appendix?
What type of information includes an appendix, the format of an appendix, frequently asked questions about appendices in papers, related articles.
An appendix is a section of a paper that features supporting information not included in the main text.
The appendix of a paper consists of supporting information for the research that is not necessary to include in the text. This section provides further insight into the topic of research but happens to be too complex or too broad to add to the body of the paper. A paper can have more than one appendix, as it is recommended to divide them according to topic.
➡️ Read more about what is a research paper?
An appendix can take many types of forms. Here are some examples:
- Surveys. Since many researchers base their methodology on surveys, these are commonly found attached as appendices. Surveys must be included exactly as they were presented to the respondents, and exactly how they were answered so the reader can get a real picture of the findings.
- Interviews . Whether it’s a transcript or a recording, interviews are usually included as an appendix. The list of questions and the real answers must be presented for complete transparency.
- Correspondence . All types of communication with collaborators regarding the research should be included as an appendix. These can be emails, text messages, letters, transcripts of audio messages, etc.
- Research tools . Any instrument used to perform the research should be acknowledged in an appendix to give the reader insight into the process. For instance, audio recorders, cameras, special software, etc.
- Non-textual items . If the research includes too many graphs, tables, figures, illustrations, photos or charts, these should be added as an appendix.
- Statistical data . When raw data is too long, it should be attached to the research as an appendix. Even if only one part of the data was used, the complete data must be given.
➡️ Learn more about surveys, interviews, and other research methodologies .
The format of an appendix will vary based on the type of citation style you’re using, as well as the guidelines of the journal or class for which the paper is being written. Here are some general appendix formatting rules:
- Appendices should be divided by topic or by set of data.
- Appendices are included in the table of contents.
The most common heading for an appendix is Appendix A or 1, centered, in bold, followed by a title describing its content.
- An appendix should be located before or after the list of references.
- Each appendix should start on a new page.
- Each page includes a page number.
- Appendices follow a sequential order, meaning they appear in the order in which they are referred to throughout the paper.
An appendix is usually added before or after the list of references.
There is no specific space limit to an appendix, but make sure to consult the guidelines of the citation format you are using.
Yes, all appendices must be included in the table of contents.
Appendices feature different types of material, for instance interviews, research tools, surveys, raw statistical data, etc.

Easy Guide on How to Write an Appendix

Understanding What Is an Appendix
Many students ask, 'What is an appendix in writing?'. Essentially, an appendix is a compilation of the references cited in an academic paper, prevalent in academic journals, which can be found in any academic publication, including books. Professors frequently require their students to include an appendix in their work.
Incorporating an appendix in your written piece can aid readers in comprehending the information presented. It is important to note that different professors may have varying guidelines on how to write an appendix. To learn more about how to write an appendix for a research paper according to APA, Chicago, and MLA styles, check out the following paragraphs prepared by our PRO nursing essay writing service !
Meanwhile, note that an appendix comprises all the information utilized in a paper, including references and statistics from several authors and sources (the number varies according to the type of academic paper). The purpose of the appendix is to prevent vague or irrelevant information and improve the reader's understanding of the paper.
The Purpose of an Appendix
To understand what an appendix tries to accomplish and how to write an appendix example, after all, we must first answer the key question, 'What is the purpose of an appendix?'. In short, an appendix is crucial for further explaining complex information that may be difficult to fully convey within the main text of an essay. It is intended to offer readers additional information about the topic addressed in the paper.
The material presented in an appendix has the potential to bolster the argument and sway the reader's opinion. Nonetheless, you should try to incorporate supporting material and examples toward the end of the paper to avoid disrupting the flow of the main text. Furthermore, the likelihood of including an appendix increases as a paper becomes more advanced. The use of an appendix is especially prevalent in the academic writing of a research document and journal-style scientific paper, in which extra information is usually needed to support a main point of view.
How to Structure an Appendix
While there are variations between formats, each one follows a basic structure. Thus, understanding the general structure is an essential first step in learning about this topic. No matter if you're tasked with 'how to write an appendix MLA or APA style?' - remember that both adhere to this structure, despite their differences:

Order an Essay Now & Get These Features For Free :
Every Appendix Should Contain:
- A clear title: The title of the appendix should be concise and descriptive, clearly indicating what information is contained within it. For example, 'Appendix A: Data Tables for Study Results or 'Appendix B: Images of Experimental Setup.'
- A list of contents: Including a table of contents in the appendix can be helpful for readers to navigate the information provided. For example:
Table of Contents:
A. Data Tables for Study Results
B. Images of Experimental Setup
C. Survey Questions and Responses
D. Sample Interview Transcripts
- Page numbers: The appendix should be a separate page, independently numbered from the main body of the paper, and specified uniformly (e.g., 'Appendix A,' 'Appendix B,' etc.). For example:
Page 1 of 5
- Relevant information: The appendix should contain all the relevant information supporting the main arguments of the document, including tables of data, raw statistical data, charts, or other documents. For example:
Figure 1: Experimental Results
[insert graph or chart here]
- Proper formatting: The appendix should be formatted in accordance with the specific requirements of the chosen citation style (e.g., APA, MLA, Chicago). For example:
Appendix B: Survey Questions and Responses
[insert survey questions and responses here, formatted following APA style guidelines]
- Clear labeling: Each element should have a clear appendix label so readers can easily understand its relevance to the paper. For example:
Table 1: Demographic Characteristics of Survey Respondents
- Concise explanation: It is important to provide short detailed descriptions of each element in the Appendix so that readers can understand its importance. For example:
Appendix C: Sample Interview Transcripts
Transcripts of the three interviews with the study participants shall be included for reference. These interviews provide further insights into the experiences of participants and their views on the subject addressed in this document.
Need college essay help ? You can always ask us to do a custom term paper from our professional writers.
General Appendix Format
To ensure proper formatting, it is important to understand the basics of how to structure an appendix. Although it may seem overwhelming, the basic format is relatively easy to comprehend and serves as a foundation for understanding the APA and MLA formats. Additionally, mastering the basic format can be helpful when writing an appendix for a book or dissertation.

- Heading “Appendix #” . Contains a number or letter, that could be 1 or A.
- Reference List.
- Index Table followed a list of appendices.
- Page Number.
Do You Need More Help With Your APPENDIX?
Get help from professional writers asap!
How to Write an Appendix in Different Styles
There are two distinct styles for creating an appendix, and it's important to familiarize yourself with both since a professor may request one or the other. Our expert writers have compiled guidelines and rules for both formats - the Appendix APA format and the Appendix MLA format. Although they share some similarities, they also have unique features and regulations that must be strictly followed.
Appendix APA
Many professors require students to write an appendix in a paper of this format. To master how to write an appendix APA format and get the structure correct, it's a good idea to follow these guidelines and rules:
The guidelines for Appendix APA:
- The appendix begins with the heading 'Appendix' followed by ABC.
- It should also be written on top of the appendix title.
- Every appendix follows the order of the stated information in the paper.
- Include the appendix after the reference list.
- Include page numbers for each appendix.
- Appendices are to have their own page, regardless of the size.
- Include Footnotes.
The general rules for Appendix APA are to be followed when writing. This is what professors look for when a paper is required when apprentices are to be written in this format. Learn the general rules to master how to write an appendix APA style and get you onto the right path to success. You may find it useful to memorize this information or keep a note of it.
Rules for APA:
- All appendices should include their own point.
- Include a title for each appendix.
- For multiple appendices, use ABC for tilting them.
- For reference within the body, include (see appendix a) after the text.
- The title should be centered.
- All appendices are to have their own page, regardless of the size.
- Paragraph One should be written without indents.
- The rest of the paragraphs should have the intended formatting.
- Include double spacing.
Whether you're tackling how to write an interview paper in APA appendix or any other type of academic work, the following example can serve as a valuable blueprint to guide you through the process.
Appendix Chicago Style
Writing an appendix Chicago style is rather similar to APA. Though, there are some minor differences. Take a look at these guidelines for this form of an appendix.
Guidelines for an Appendix Chicago Style
- More than one appendix is described as appendices.
- The font required for the appendix Chicago style is Times New Roman.
- The text size should be 12 points.
- The page numbers should be displayed on the top right of each page.
- The page numbers should also be labeled as 'Page 1,2,3'.
- Avoid including a page number on the front cover.
- The bibliography should be the final new page. It should not share a page with any other content.
- It is possible to include footnotes in the bibliography.
To better comprehend how to write an appendix in Chicago style, glance through the example below:
Appendix MLA Format
The guidelines and regulations for creating an appendix in MLA format are largely similar to those in APA format. However, there are some differences between the two, the most notable being that the MLA appendix is placed before the reference list.
The guidelines for MLA Format:
- The appendix is included before the list of references.
It may be useful to follow the example of an appendix to better understand how to write an appendix in MLA style. Doing so can increase the chances of getting a grasp of the MLA rules to fulfill the requirements of your professor on your academic paper.
Rules for MLA
- The title is to be centered.
- The list should be double-spaced.
- The first line should include each reference in the left margin. Every subsequent line is to be formatted so it's invented. This can be referred to as 'hanging indent' to make things easier.
- The reference list must be in alphabetical order. This can be done with the first letter of the title of the reference. Though, this is usually done if the writer is unknown. If the writer is known, you can also use the first letter of the surname.
- If you include the name of the known writer, use this order. SURNAME, FIRST NAME, YEAR.
- Italic fonts are required for the titles of complete writings, internet sites, books, and recordings.
- It is important not to use an italic font on reference titles that only refer to the part of a source. This includes poetry, short papers, tabloids, sections of a PDF, and scholarly entries.
Before we conclude, let's dive deeper into the world of appendix writing by exploring an example of how to write an appendix MLA style.
Let's wrap this up! It's safe to say that following the APA, Chicago, and MLA formats is crucial when crafting an appendix. As we've seen, starting with an APA appendix example can help ease you in mastering how to write an appendix of paper. Once you have a handle on the precise formats and guidelines, creating an appendix becomes a piece of cake. Also, memorizing the format can help you whip up accurate appendices for any type of paper, whether an essay or a dissertation. Trust us, mastering this topic is a must if you want to excel in knowing how to write an appendix in a report or any other academic work.
Moreover, if you ever find yourself in need of additional academic assistance, be sure to check out our resources on how to write an article review . Or, better yet, why not let us handle your most challenging tasks with ease by simply sending us a ' write my paper request? We are here to support you every step of the way.
Get Immediate Writing Help
Our appendix pages are written with the customer’s needs in mind, and plagiarism-free. Why not give it a try?
What Is An Appendix In Writing?
What is the purpose of an appendix, how to format an appendix.

Daniel Parker
is a seasoned educational writer focusing on scholarship guidance, research papers, and various forms of academic essays including reflective and narrative essays. His expertise also extends to detailed case studies. A scholar with a background in English Literature and Education, Daniel’s work on EssayPro blog aims to support students in achieving academic excellence and securing scholarships. His hobbies include reading classic literature and participating in academic forums.

is an expert in nursing and healthcare, with a strong background in history, law, and literature. Holding advanced degrees in nursing and public health, his analytical approach and comprehensive knowledge help students navigate complex topics. On EssayPro blog, Adam provides insightful articles on everything from historical analysis to the intricacies of healthcare policies. In his downtime, he enjoys historical documentaries and volunteering at local clinics.
.webp)
We apologize for any inconvenience as we update our site to a new look.

- Walden University
- Faculty Portal
General Research Paper Guidelines: Appendices
If you have some information you would like to include in your research but it could potentially be distracting to readers or inappropriate within the body of your research paper, you can always include supplemental information as an appendix to your work. An appendix or appendices should always be inserted after your Reference List; however, the appropriateness of appendix content really depends on the nature and scope of your research paper.
For a more in-depth review of what supplemental materials might be included in a social science appendix, be sure to review Section 2.14 “Appendices” (pp. 41-42) of your 7 th edition APA manual.
Appendices Formatting
APA 7 addresses appendices and supplemental materials in Section 2.14 and on page 41:
- The appendices follow the reference list.
- They are lettered "Appendix A," "Appendix B," "Appendix C," and so forth. If you have only one appendix, however, simply label it Appendix.
- Put figures and tables in separate appendices. The appendix title serves as the title for a table if it is the only table in the appendix.
- If you decide that certain figures and tables should appear in the same appendix, number them A1, A2, A3, and so forth, according to the appendix in which they appear.
- The materials in the appendix must not extend beyond the margins of the rest of the paper: Reduce the appendix materials as needed.
As a general guide, appendices are appropriate for any material that, if presented in the main body of the document, would unnecessarily interrupt the flow of the writing. Note that it is unlikely that you will use appendices in Walden course papers. For doctoral capstone studies, you might include some appendices with supplementary information.
- Previous Page: References
- Office of Student Disability Services
Walden Resources
Departments.
- Academic Residencies
- Academic Skills
- Career Planning and Development
- Customer Care Team
- Field Experience
- Military Services
- Student Success Advising
- Writing Skills
Centers and Offices
- Center for Social Change
- Office of Academic Support and Instructional Services
- Office of Degree Acceleration
- Office of Research and Doctoral Services
- Office of Student Affairs
Student Resources
- Doctoral Writing Assessment
- Form & Style Review
- Quick Answers
- ScholarWorks
- SKIL Courses and Workshops
- Walden Bookstore
- Walden Catalog & Student Handbook
- Student Safety/Title IX
- Legal & Consumer Information
- Website Terms and Conditions
- Cookie Policy
- Accessibility
- Accreditation
- State Authorization
- Net Price Calculator
- Contact Walden
Walden University is a member of Adtalem Global Education, Inc. www.adtalem.com Walden University is certified to operate by SCHEV © 2024 Walden University LLC. All rights reserved.
We use cookies to give you the best experience possible. By continuing we’ll assume you’re on board with our cookie policy

- A Research Guide
- Research Paper Guide
How to Make an Appendix for a Research Paper
What is an appendix, what can you include in an appendix.
- Texts or paragraph
- Graphs or Charts
- Examples with images, photographs, and illustrations
- Drawings, diagrams, and maps
- Links to websites
- List of suggested reading
The content of an appendix
Visual documents, instruments used, transcripts of interviews and surveys, the format of an appendix, title of the appendix, content order, placement and page numbers, make your appendix perfect.
Read also: Who will do assignment for me and make it plagiarism-free?
Review and revise
Check for quality, check if the appendix is cited in the text properly.
By clicking "Log In", you agree to our terms of service and privacy policy . We'll occasionally send you account related and promo emails.
Sign Up for your FREE account
Organizing Your Social Sciences Research Paper: Appendices
- Purpose of Guide
- Writing a Research Proposal
- Design Flaws to Avoid
- Independent and Dependent Variables
- Narrowing a Topic Idea
- Broadening a Topic Idea
- The Research Problem/Question
- Academic Writing Style
- Choosing a Title
- Making an Outline
- Paragraph Development
- The C.A.R.S. Model
- Background Information
- Theoretical Framework
- Citation Tracking
- Evaluating Sources
- Reading Research Effectively
- Primary Sources
- Secondary Sources
- What Is Scholarly vs. Popular?
- Is it Peer-Reviewed?
- Qualitative Methods
- Quantitative Methods
- Common Grammar Mistakes
- Writing Concisely
- Avoiding Plagiarism [linked guide]
- Annotated Bibliography
- Grading Someone Else's Paper
An appendix contains supplementary material that is not an essential part of the text itself but which may be helpful in providing a more comprehensive understanding of the research problem or it is information that is too cumbersome to be included in the body of the paper. A separate appendix should be used for each distinct topic or set of data and always have a title descriptive of its contents.
Tables, Appendices, Footnotes and Endnotes . The Writing Lab and The OWL. Purdue University.
Importance of...
Appendices are always supplementary to the research paper. As such, your study must be able to stand alone without the appendices, and the paper must contain all information including tables, diagrams, and results necessary to understand the research problem. The key point to remember when including an appendix is that the information is non-essential; if it were removed, the reader would still be able to comprehend the significance, validity , and implications of your research.
It is appropriate to include appendices for the following reasons:
- Including this material in the body of the paper that would render it poorly structured or interrupt the narrative flow;
- Information is too lengthy and detailed to be easily summarized in the body of the paper;
- Inclusion of helpful, supporting, or useful material would otherwise distract the reader from the main content of the paper;
- Provides relevant information or data that is more easily understood or analyzed in a self-contained section of the paper;
- Can be used when there are constraints placed on the length of your paper; and,
- Provides a place to further demonstrate your understanding of the research problem by giving additional details about a new or innovative method, technical details, or design protocols.
Appendices . Academic Skills Office, University of New England; Chapter 12, "Use of Appendices." In Guide to Effective Grant Writing: How to Write a Successful NIH Grant . Otto O. Yang. (New York: Kluwer Academic, 2005), pp. 55-57; Tables, Appendices, Footnotes and Endnotes . The Writing Lab and The OWL. Purdue University.
Structure and Writing Style
I. General Points to Consider
When considering whether to include content in an appendix, keep in mind the following:
- It is usually good practice to include your raw data in an appendix, laying it out in a clear format so the reader can re-check your results. Another option if you have a large amount of raw data is to consider placing it online and note that this is the appendix to your research paper.
- Any tables and figures included in the appendix should be numbered as a separate sequence from the main paper . Remember that appendices contain non-essential information that, if removed, would not diminish a reader's ability to understand the research problem being investigated. This is why non-textual elements should not carry over the sequential numbering of non-textual elements in the body of your paper.
- If you have more than three appendices, consider listing them on a separate page at the beginning of your paper . This will help the reader know what information is included in the appendices [always list the appendix or appendices in a table of contents].
- The appendix can be a good place to put maps, photographs, diagrams, and other images , if you feel that it will help the reader to understand the content of your paper, while keeping in mind the study should be understood without them.
- An appendix should be streamlined and not loaded with a lot information . If you have a very long and complex appendix, it is a good idea to break it down into separate appendices, allowing the reader to find relevant information quickly as the information is covered in the body of the paper.
II. Content
Never include an appendix that isn’t referred to in the text . All appendices should be summarized in your paper where it is relevant to the content. Appendices should also be arranged sequentially by the order they were first referenced in the text [i.e., Appendix 1 should not refer to text on page eight of your paper and Appendix 2 relate to text on page six].
There are very few rules regarding what type of material can be included in an appendix, but here are some common examples:
- Correspondence -- if your research included collaborations with others or outreach to others, then correspondence in the form of letters, memorandums, or copies of emails from those you interacted with could be included.
- Interview Transcripts -- in qualitative research, interviewing respondents is often used to gather information. The full transcript from an interview is important so the reader can read the entire dialog between researcher and respondent. The interview protocol [list of questions] should also be included.
- Non-textual elements -- as noted above, if there are a lot of non-textual items, such as, figures, tables, maps, charts, photographs, drawings, or graphs, think about highlighting examples in the text of the paper but include the remainder in an appendix.
- Questionnaires or surveys -- this is a common form of data gathering. Always include the survey instrument or questionnaires in an appendix so the reader understands not only the questions asked but the sequence in which they were asked. Include all variations of the instruments as well if different items were sent to different groups [e.g., those given to teachers and those given to administrators] .
- Raw statistical data – this can include any numerical data that is too lengthy to include in charts or tables in its entirety within the text. This is important because the entire source of data should be included even if you are referring to only certain parts of a chart or table in the text of your paper.
- Research instruments -- if you used a camera, or a recorder, or some other device to gather information and it is important for the reader to understand how, when, and/or where that device was used.
- Sample calculations – this can include quantitative research formulas or detailed descriptions of how calculations were used to determine relationships and significance.
NOTE: Appendices should not be a dumping ground for information. Do not include vague or irrelevant information in an appendix; this additional information will not help the reader’s overall understanding and interpretation of your research and may only distract the reader from understanding the significance of your overall study.
ANOTHER NOTE : Appendices are intended to provide supplementary information that you have gathered or created; it is not intended to replicate or provide a copy of the work of others. For example, if you need to contrast the techniques of analysis used by other authors with your own method of analysis, summarize that information, and cite to the original work. In this case, a citation to the original work is sufficient enough to lead the reader to where you got the information. You do not need to provide a copy of this in an appendix.
III. Format
Here are some general guideline on how to format appendices . If needed, consult the writing style guide [e.g., APA, MLS, Chicago] your professor wants you to use for more detail:
- Appendices may precede or follow your list of references.
- Each appendix begins on a new page.
- The order they are presented is dictated by the order they are mentioned in the text of your research paper.
- The heading should be "Appendix," followed by a letter or number [e.g., "Appendix A" or "Appendix 1"], centered and written in bold type.
- If there is a table of contents, the appendices must be listed.
- The page number(s) of the appendix/appendices will continue on with the numbering from the last page of the text.
Appendices . The Structure, Format, Content, and Style of a Journal-Style Scientific Paper. Department of Biology. Bates College; Appendices . Academic Skills Office, University of New England; Appendices . Writing Center, Walden University; Chapter 12, "Use of Appendices." In Guide to Effective Grant Writing: How to Write a Successful NIH Grant . Otto O. Yang. (New York: Kluwer Academic, 2005), pp. 55-57 ; Tables, Appendices, Footnotes and Endnotes . The Writing Lab and The OWL. Purdue University; Lunsford, Andrea A. and Robert Connors. The St. Martin's Handbook . New York: St. Martin's Press, 1989; What To Know About The Purpose And Format Of A Research Paper Appendix . LoyolaCollegeCulion.com.
Writing Tip
Consider Putting Your Appendices Online
Appendices are useful because they provide the reader with information that supports your study without breaking up the narrative or distracting from the main purpose of your paper. If you have a lot of raw data or information that is difficult to present in textual form, consider uploading it to an online site. This prevents your paper from having a large and unwieldy set of appendices and it supports a growing movement within academe to make data more freely available for re-analysis. If you do create an online portal to your data, note it prominently in your paper with the correct URL and access procedures if it is a secured site.
Piwowar, Heather A., Roger S. Day, and Douglas B. Fridsma. “Sharing Detailed Research Data Is Associated with Increased Citation Rate.” PloS ONE (March 21, 2007); Wicherts, Jelte M., Marjan Bakker, and Dylan Molenaar. “Willingness to Share Research Data Is Related to the Strength of the Evidence and the Quality of Reporting of Statistical Results.” PLoS ONE (November 2, 2011).
- << Previous: Quantitative Methods
- Next: Proofreading Your Paper >>
- Last Updated: Sep 8, 2023 12:19 PM
- URL: https://guides.library.txstate.edu/socialscienceresearch

Home » Education » How to Write an Appendix for a Research Paper
How to Write an Appendix for a Research Paper
If you are in the process of writing a research paper and wondering how to write an appendix for a research paper, then this article is for you. Writing could come in different forms serving different purposes. Over the years, the term ‘writing’ has expanded into many horizons that sub categories of writing have emerged: academic writing, literary writing, business writing, technical writing, legal writing and so on. In each of these sub-categories of writing, there are many diverse forms of writing that fall under them, for example, in academic writing, there could be many subdivisions or types of academic writing as writing a research paper, writing reports, writing essays, etc., and all of these diverse pieces of writing have their own structure and style of writing. This article seeks to explore the arena of writing an appendix for a research paper.
What is an Appendix?
Appendix is defined as a supplement to a document, form a part of a main document, but not essential for its completeness. It contains supporting information, and usually, appears at the end of the document. Research papers are lengthy and precise (containing only what is strictly relevant) at the same time. Yet, apart from what is provided in the research, if a writer feels that some additional document could be of use to complement his facts or information, he/she can attach it at the end of the paper as an appendix. Appendices could, usually, contain maps, graphs, questionnaires used for the study, raw data, etc. Also, the reference section for the reader could also be a part of the appendix.
How to write an appendix for a Research Paper
Since an appendix is not a type of writing that has to follow a particular structure or form but additional documents, there is no acknowledged structure to write one. There are things that writer should consider when they are writing an appendix and each of those is described in separate paragraphs below.
First, you can review previous work, study what other writers have done when attaching an appendix to their research paper. This would always provide you with the benefit of having some prior background knowledge as to how to the task.
Next, it is best to review your own work. Go through it from cover to cover carefully thinking what should be included inside your research paper (usually, the most important things go inside) and what documents would be best to be attached at the end if further referral is required by the reader. Take notes. Also, just to be on the safe side, you can always get somebody else’s opinion to see if the documents and information you are going to place in your appendix would make sense.
Then, gather all the information you need to place in your appendix and assess their relevance to the research paper. Remember, your purpose should not be to attach each and every little detail you found in your research.
Next, organize your appendix; the order it should appear should be properly prearranged. Use sections, headings, subheading, numbering, etc. to arrange your appendix. If there are too many documents that you can even categorize them under multiple appendices. Give different subjects to make it easier for the reader to understand.
Finally, proofread your appendix before publishing your research paper.

As such, when you are writing a research paper, you can always insert any documents that you think could be important if further referral is sought by the reader in such a way that you are not burdening the reader with heaps of facts and information written inside your research paper. An appendix is always additional, hence readers have a choice whether to read it or not.
About the Author: admin
you may also like these, leave a reply cancel reply.
- Testimonials
- How it works
- Paper Writers Team
- Essay Writing Guide
- Free plagiarism checker
- Essay title generator
- Conclusion Generator
- Citation Generator
- Can ChatGPT Write Essays?
How to Include Appendices in Your Research Paper

November 01, 2023
Did your professor tell you to have a research paper appendix, but you can’t imagine what goes into it? Don’t fret; we’re about to show you! The appendix is a section at the end of the paper with additional information that doesn’t belong in the main text. It shouldn’t have any arguments, quotes, or conclusions. You must use it to support the claims you already made in your essay — appendices help illustrate everything better. Take a look at ten types of content that will fit right in.
Ten Pieces of Information that Belong in Appendices
You finished your research paper and composed a list with references. Now what? That’s right, now it is time for appendices, especially if your essay has a lot of complex information! Look through our list and include relevant options in your work.
- Supplementary Data. Often, students working on a complex paper include a ton of information. There is no place for any additional illustrations, and this is where an appendix could help you. Were you explaining a tricky market situation to your readers? Add a graph to the appendix that helps demonstrate your point better. Were you listing all the numbers to show growth or decline? Create a table that will let your readers see all the data in one concise section. Raw data always has its place there: it could clutter your paper if you included it in the body in its unprocessed form, but it won’t hurt anyone in the appendix. Charts could also come in handy. Imagine comparing three different companies and analyzing their capabilities and limitations. You already explained their similarities and differences with text and numbers. Is adding a chart for a better illustration of these findings necessary? Not at all. But if it could make the picture clearer visually, why not? That’s what supplementary material is for!
- Research Instruments. Most dissertations and other layered research works involve an experiment of a sort. Students present a hypothesis, and then they need to perform a study to either prove or refute it. They rely on various instruments, such as surveys, quizzes, questionnaires, and other mediums. They are vital but don’t have a place in a paper in their full form. Let’s say you have five interview transcripts from questioning five people about their job satisfaction. This interview had ten questions and required ten expanded answers. You cannot possibly copy-paste them all into a body. No, you’ll reference and summarize them, but you won’t show them entirely. You can do it in the appendix, though! Put your raw data in there, and if your audience feels interested, they will look to see just what you asked your interviewees and what they responded with. It creates transparency because there is no place for double meanings left, and it also gives other researchers a chance to use your questions in their work.
- Detailed Methodology. Methodology is a crucial component of every research. You need to decide if it will be quantitative, qualitative, mixed, or experimental. Every student must mention relevant information about their chosen methodology in a paper: they have to say what they plan to do, in what way, and through which methods. But some of them might want to go beyond this. For instance, your experiment might be complex, or you might be so passionate that you want to share everything you did step by step with your readers. It’s not a problem, do it! Create a fitting appendix title and go ahead. That’s the whole point of appendices: they host extra info. Sometimes, students use unique equipment to do their experiments. If this is your case, describe this equipment in detail and put it at the end of the paper in another appendix. Any interested party can go there and take a closer look.
- Code and Algorithms. This point exists mostly for students who study IT or other similar spheres. If your paper deals with endless codes and tricky algorithms, putting them in the body of the paper is not an option. It’ll stretch for eternity, ruin the readability of your text, and your readers will feel bored very quickly. Creating an appendix and shoveling this info in there is a great solution! You could include the entire source code you made for a program, showing how it exists. Multiple readers appreciate the inclusion of raw data like this. If you’re a bit of a nerd, add a detailed algorithm description in the appendix. Some people will ignore it, but others will enjoy studying it. If you’re having trouble with this paper and you keep thinking, “I’d like to pay someone to write my paper because it’s annoying,” ask for professional help. Our experts are one click away from you: we can perfect your academic writing or show you how to deal with appendices. You only need to mention, “I need someone good to write my research paper today.” One word from you, and we’ll get started.
- Participant Consent Forms. This is a point that many students overlook. Sometimes, professors ask to include complete consent forms personally, but in most cases, it’s up to you. You need to realize why this is essential. You had to find people to test your hypothesis and to write your research paper: most likely, they answered some questions for you or gave you a full interview. You need to prove that you got their consent and didn’t mislead them into participating in your research without telling them everything. This is why consent forms are there and why you must use them. Compose them carefully, covering every detail you shared with your living sample, and then paste this info onto a new page. Yeah, it’ll be the appendix. While your general readers might not be interested in it, we guarantee that your professor will take a look. Ethical considerations are everything where human subjects are concerned, and you should prove that you took them into account when working on your research paper.
- Maps, Images, or Photographs. Another reason appendices are common is their ability to host visual illustrations of your research. If adding photos or images is essential, you could include them in the body, but putting them under your appendix title is a much better idea if there are too many of them or their presence is not required. Did you measure the distance between locations to highlight some historical facts? Add a map to the appendix! It’ll be interesting, especially if you customize it, even if it’s not critical. Maybe you were exploring diseases some people have and how they are displayed. Your research paper mentions symptoms, but for particularly curious readers, you could include images that illustrate everything you described visually. Appendices are a perfect place for this. The same principle applies to photographs. You could take them of your subjects, equipment, and other elements involved in your research. Pasting them within the body isn’t important; it’d distract your audience from your main points, but including them in a separate appendix would do the trick.
- 7) Extended Explanations. Let us warn you, this form of material isn’t the best because most of your readers will find it boring or redundant. But it still exists and is common enough to be mentioned in this list. While it is obvious that you must do everything in your power to explain every point of your research in the body of the paper, sometimes it’s not enough. You might need to provide even more explanations, and this is something you can easily do with appendices. Imagine that you came up with a good formula to solve some task. You quickly explain how you did it, providing the basics and avoiding the boring details. But you want to show every aspect of your work — good news! — appendices allow you to do this. Include your complete formula in there. True, not everyone likes raw data like this, but some will appreciate it, and chances are, your professor will be among them. This is called being thorough. You could do the same with any background information. If you’ve been investigating a psychological profile of someone and this person experienced abuse & shared all the details with you, you can briefly disclose them in the text and then provide a complete overview in the appendix.
- Statistical Analysis. Many students shudder at the thought of doing any calculations, but some love it. Whether you fall into the first or the second category, you might need to write research papers like this, and in this case, you won’t be able to avoid appendices. They might become your best friends. If you had to show how you got from point A to point B, writing down all the formulas and analyses isn’t smart. It’ll turn your essay into a number-filled mess, and by the time people reach the end of the calculation, they’ll forget what they were reading in the first place and what this calculation is for. Place raw statistical data at the end of your research in an appendix. It’ll be there for extra thorough readers who are interested in seeing how you arrived at your conclusions, and, in turn, it won’t be cluttering your main text. As it always is with appendices, include crucial info in a body and mark the rest as supplementary material.
- Supporting Documentation. When students do their research, they accumulate some documentation. It might be full of vague or irrelevant information, but it’s still there, and if it can underline some of your points, you should include it. If you had to organize an extensive interview with your participant, you could have ten or even twenty pages. The most important stuff goes into the body of your essay. All the pages can be linked in an appendix: this will give readers a fuller picture of who the participant is and what they think. If you have some other files, like a case study with all observations or a detailed methodology, you could create multiple appendices and situate all the info there. Make as many of them as you need — unless your professor explicitly restricts it, there are no limits.
- Abbreviations and Acronyms. The last important type of content that has its place in appendices is abbreviations and similar elements. You might be working on a journal-style scientific paper with numerous complex concepts and phenomena. You can spell them out in their complete form once, but doing it repeatedly will be distracting and unnecessary. You’ll make your word count huge for no reason, which might result in a penalty. The solution lies in abbreviating and explaining these concepts on a new page in the appendix. Do the same with acronyms. If, at some point, your readers will forget what this or that abbreviation means, they’ll go to an appendix and look it up.
Format Your Appendices Correctly to Impress Your Readers Further
A clear appendix is a sure way to simplify your info for your readers. They’ll enjoy seeing understandable illustrations of the content they’ve just read. Remember that you could use several appendices simultaneously; there is no rule against it. Some of the materials we listed can go in Appendix A, while others will go to Appendix B. Be thorough, ask for help if you’re in trouble, and keep producing awesome research papers.

While being committed to a number of charitable causes, like volunteering at special events or giving free art lessons to children, Marie doesn’t forget her vocation – writing. She can write about almost anything but has focused on time management, motivation, academic and business writing.
Related posts

November 01 2023

Don`t have an account?
Password recovery instructions have been sent to your email
Back to Log in
Nixed: The Upside of Getting Dumped
- RA Publications
- Journal & Working Papers
- Explore Smart Beta
- ESG Investing
- Asset Allocation Interactive
- Mutual Funds & ETFs
- Institutional, SMAs, and Commingled
- In the News
- 20th Anniversary

Historically, index deletions have beaten the Russell 2000 Value Index in spectacular fashion and could add an abnormal upside to a portfolio when the current growth-dominated bubble starts to deflate.
Deletions lag the market by more than half in the year leading up to their removal from an index, but they historically outperform the market for at least five years after the breakup.
A deletions strategy relies on two growth drivers to fuel performance: long horizon mean reversion and a liquidity effect.
No one enjoys getting dumped. This holds true in finance and investing as much as it does in romantic relationships. When companies are dumped from the major indexes, their managers and shareholders may feel jilted and their stock may flounder post-breakup. But according to our research, deletion-related downward spirals are hardly inevitable. As it turns out, getting dumped by an index can have an impressive upside, just as a romantic breakup can sow seeds for personal growth. Dumped companies and their shareholders fare surprisingly well on average, better even than the stocks that replaced them.
Create your free account or log in to keep reading.
Register or Log in
Here's how it works. All the established index providers have roughly the same modus operandi. They add stocks that have become valuable and popular enough to catch the investment community’s eye, whether through the subjective judgment of an investment committee like the S&P or MSCI indexes or through a formula based on market capitalization or float like Russell’s. On the other hand, unless the breakup results from a merger or acquisition, dumped stocks are almost always unloved, out of favor, and no longer valuable enough for the index provider to pay them any attention. They have, in dating parlance, let themselves go. The S&P 500, the Nasdaq-100, the Russell 1000 all drop the fading older companies to make way for the Teslas and Nvidias, the exciting and frothy new additions.
How Do Nixed Companies Fare?
We have explored index additions and deletions and associated issues in an array of research papers over the years, and our findings have inspired a new approach to cap-weighted indexing. 1 In each paper, we keep a narrow focus on one aspect of this work: the reconstitution of a cap-weighted index. The reconstitution puzzle is a rich and nuanced topic, and our analysis has revealed several striking characteristics:
- Over the past 33 years, additions have generally outperformed the market in the year before the index reconstitution, while discretionary deletions have underperformed. How powerful is this performance difference? Additions have typically more than doubled compared to the stocks they replaced.
- When an index is reconstituted, added and deleted stocks tend to exhibit a wide valuation gap relative to their underlying fundamentals. Exhibit 1 shows that additions are priced at roughly twice the market multiple and discretionary deletions are priced at roughly half the market multiple. This enormous four-fold spread implies an equally enormous gap in the companies’ expected future prospects.
- Index reconstitutions generate stupendous momentum that should reward additions and punish deletions. But with a half-life of about three months, momentum tends to be a short-lived phenomenon. It also has a largely unrecognized reversal of its own: When held on a long-term basis rather than with monthly rebalancing, past winners reverse course relative to losers after 6 to 12 months and give back all their early gains and become net losses within 12 to 24 months. 2 What’s worse, over the last quarter-century, “standard momentum” hasn’t even worked! It has a negative average alpha since 1999.
- The substantial outperformance of additions relative to deletions, between the date a change is announced and the date when the change takes effect, front-loads that entire momentum effect. Empirically, there’s nothing left beyond a day or two following index reconstitution.
- Additions modestly underperform their index over the subsequent year on average, with S&P 500 additions lagging the market by 1% to 2% from 1990 through 2022. Deletions, by contrast, outperform by more than 5% per year for the next five years.
This paper focuses on deletions, the stocks dumped by major index providers, and their potential for robust post-breakup outperformance.
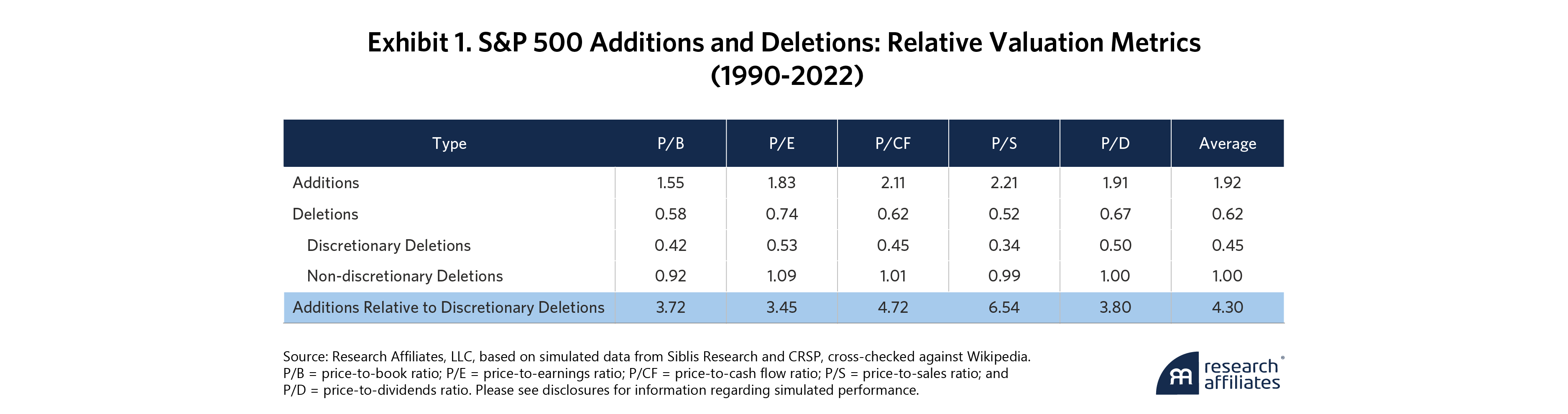
A deletions strategy benefits from long horizon mean reversion and from a liquidity effect: Deletion decisions initiate rapid selloffs. Total assets indexed to the S&P 500 are over 20% of the index’s total market capitalization. So, 20% of the total market capitalization of new additions and deletions changes hands in the days immediately surrounding the index reconstitution. Most of this trading occurs in a single “market on close” block trade that coincides with the reconstitution of the index. 3 In this way, index funds create the illusion of zero trading costs, in order to achieve near-zero tracking error. Deletions experience massive selling pressure since index fund investors must unload their shares; as a result, the market clearing prices of dumped stocks are often much lower than what they would have attracted before the deletion decision. This sets the stage for an impressive rebound.
One nuance is beyond the scope of this paper: flip-flops! Sometimes after a deeply depressed stock is deleted, its circumstances improve and it is re-added to the index at a newly inflated multiple. Sometimes this cycle repeats several times.
Portfolios of Deletions
Exhibit 2 shows how nixed S&P 500, Russell 1000, and Nasdaq-100 stocks perform in the year prior to, and in the five years after, their deletion. 4 The gray band illustrates the range of excess individual stock returns during those sample periods, from the 10th to 90th percentile. As the far left of the graph indicates, rare outliers (less than 10% of deletions) were at least four times as expensive one year before their deletion. They are sold after a 75% drop! At the other end of the spectrum, about 10% of deletions were cheaper than the S&P 500 one year before they were dumped. Five years later, the outlying deciles were down by more than 80% and up by 150% or more.
As a portfolio, deletions have fallen by more than half relative to the market in the year before their removal. That’s an even stronger result than we found in our 2023 examination of index rebalancing. 5 Deletions historically beat the market for at least five years post-breakup. Their annual outperformance is remarkably persistent and statistically significant, averaging over 5% per annum, compounded over the first five years.
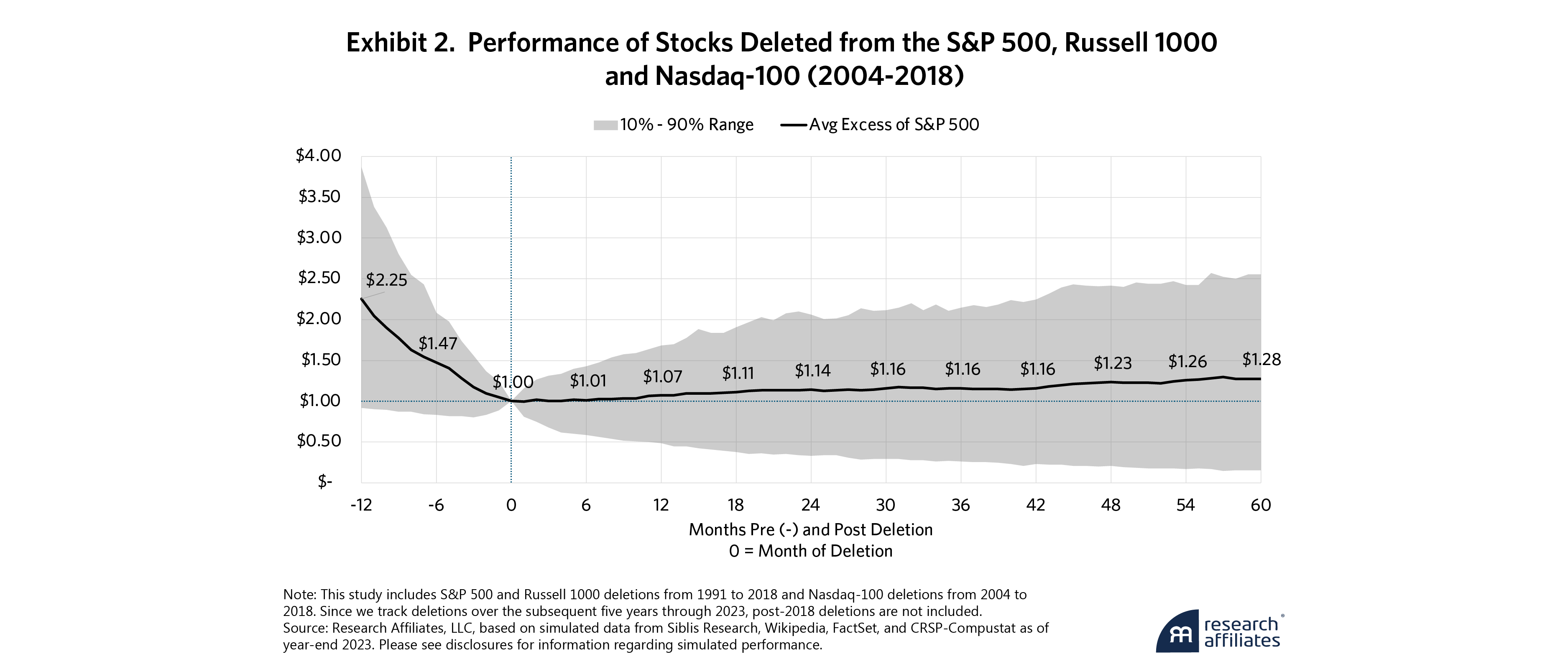
Exhibit 3 shows the growth of one dollar invested in a deletions strategy vs. the Russell 2000 Value and the indexes that dropped them. Why the Russell 2000 Value? Because any S&P 500 deletion is, by definition, not a member of the S&P 500. By the time they are dumped by the S&P 500, many S&P 500 deletions have also fallen out of the Russell 1000. Russell 1000 deletions are likewise, by definition, not members of the Russell 1000. That means most deletions are trading on the Russell 2000 at steep discounts. They are small-cap value stocks. That is why a small cap value index is the appropriate benchmark for a portfolio of dumped S&P 500 or Russell 1000 stocks. 6
Growth’s dominance will likely come to an end, and when it does, almost anything should beat the S&P 500 and Nasdaq-100. If the Russell 2000 Value is winning big and deletions can beat the Russell 2000 Value, a deletions portfolio could deliver truly remarkable outperformance.
Not everybody recognizes this. We recently shared commentary, entitled “Imitation is the Sincerest Form of Flattery,” with our Research Affiliates Insights subscribers in response to analysis of the performance of index additions and deletions by Dimensional Fund Advisors 7 and S&P . 8 Their results support ours. But they use the S&P 500 as their benchmark. That is a mistake. How could deletions not lag the S&P 500 during the small-cap value rout of the last decade and the stunning ascent of the “Magnificent Seven” in 2023 and the first half of 2024? Since deletions are generally small- and mid-cap, and are almost always deep value stocks, the Russell 2000 Value is much more relevant, and deletions fared well compared to it during much of this difficult span. We can only imagine how they will perform should small-cap and value stocks stage a significant recovery, as we think is likely in the coming decade.
Exhibit 3 also shows that the S&P 500’s performance is nearly identical to the Russell 1000’s, with the Nasdaq-100 following a more tumultuous trajectory. Because this is on a log scale, if one line is rising faster than another, that index is winning. Absent fees, taxes, spending, and other charges, an investor in a broad deletions portfolio for the five post-deletion years would be 74-times wealthier at year-end 2023 than in January 1991. Only a Nasdaq-100 investor would fare as well, albeit with higher volatility and the harrowing drawdown of the dot-com crash. After 30-plus years, S&P 500, Russell 1000, and Russell 2000 Value investors would be 55% to 65% poorer than our brave deletions investor.
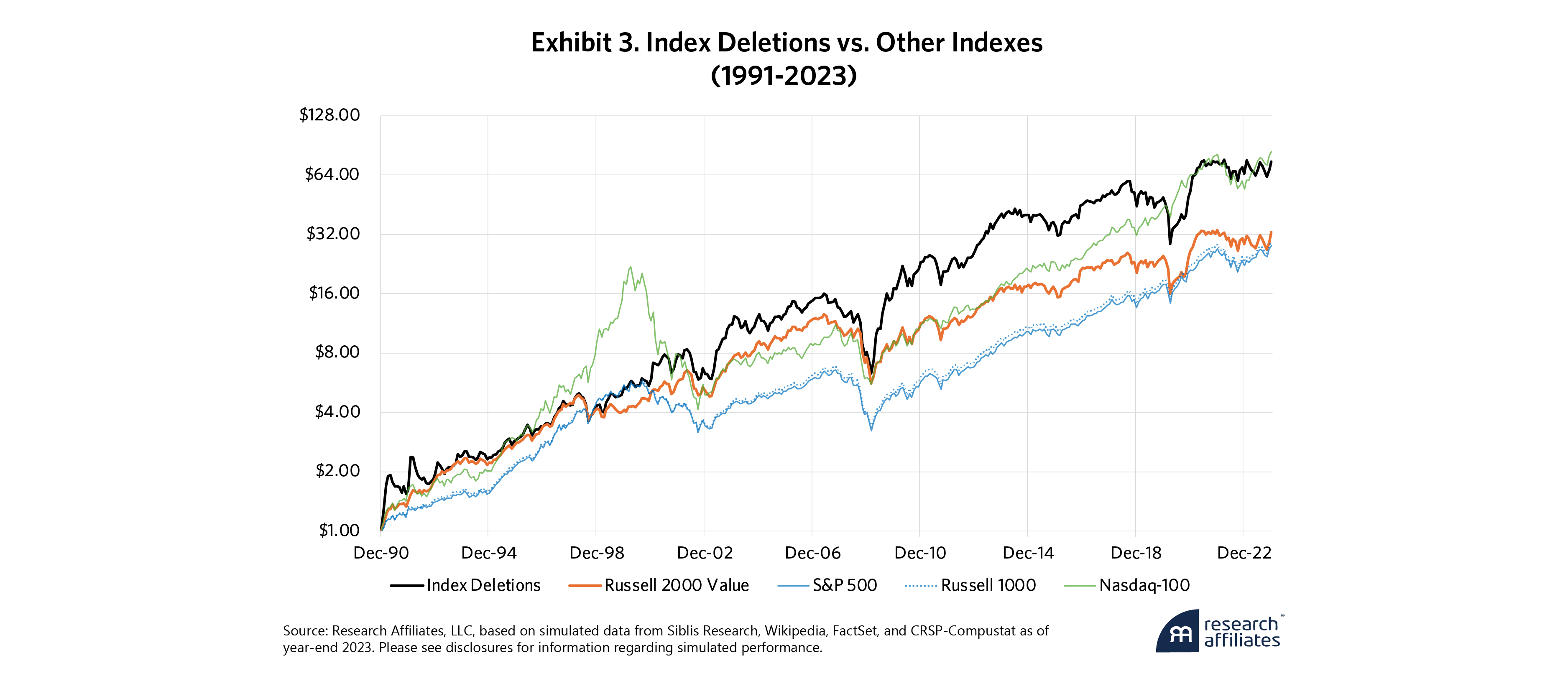
Deletions haven’t beaten the Nasdaq-100, S&P 500, or Russell 1000 over the last decade. This raises the question: Why bother? It all comes down to value and value’s recent travails. The current growth-dominated bull market has left value and small-cap stocks in the dust. In this climate, nothing beats the S&P 500 or Nasdaq-100. But growth’s dominance will likely come to an end, and when it does, almost anything should beat the S&P 500 and Nasdaq-100. If the Russell 2000 Value is winning big and deletions can beat the Russell 2000 Value, a deletions portfolio could deliver truly remarkable outperformance.
Absent fees, taxes, spending, and other charges, an investor in a broad deletions portfolio for the five post-deletion years would be 74-times wealthier at year-end 2023 than in January 1991.
Historically, deletions have beaten the Russell 2000 Value Index in impressive fashion. Exhibit 4 shows that when deletions outpace the Russell 2000 Value, they win by more than 18%, on average. In its best year, 2009, the deletions portfolio exceeded its benchmark by 90%, and in seven additional years, deletions outperformed the Russell 2000 Value by more than 10%. When deletions underperform, they lag by 5.3% on average, with a shortfall of over 10% only twice. Deletions may well add abnormal upside to a portfolio when the current growth-dominated bubble (and, yes, it is a bubble) starts to deflate.
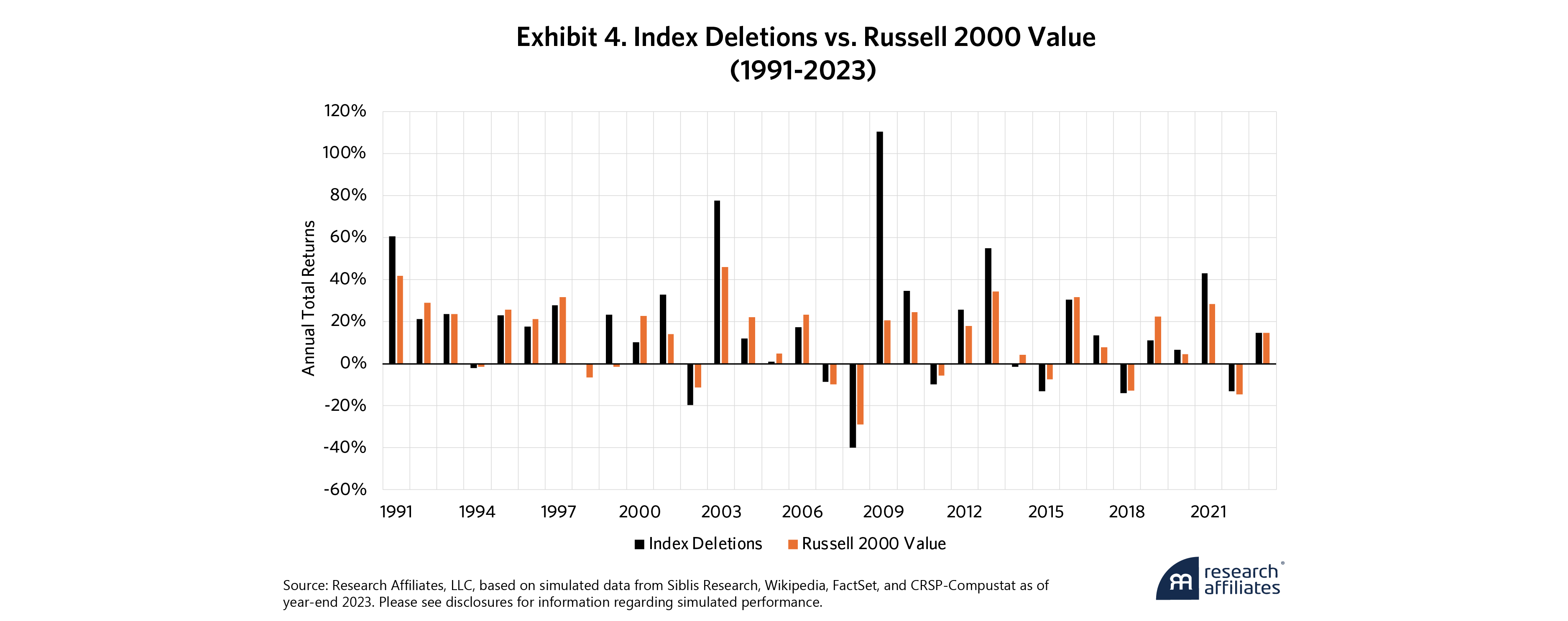
Exhibit 5 shows how well the deletions strategy holds up against various alternative indexes. To be sure, as of year-end 2023, it lagged the S&P 500 and Russell 1000, but it matched the Russell 2000 Value, with which it had the highest correlation and the lowest tracking error. The value-added, 14% per annum, is nearly as high as the rather large 19% tracking error to the S&P 500. But again, the deletions strategy has a penchant for extreme upside returns, so the total tracking can be misleading. By excluding the upside volatility and instead focusing on the downside tracking error, the portfolio delivers a much more palatable 8.0% tracking error to the S&P 500.
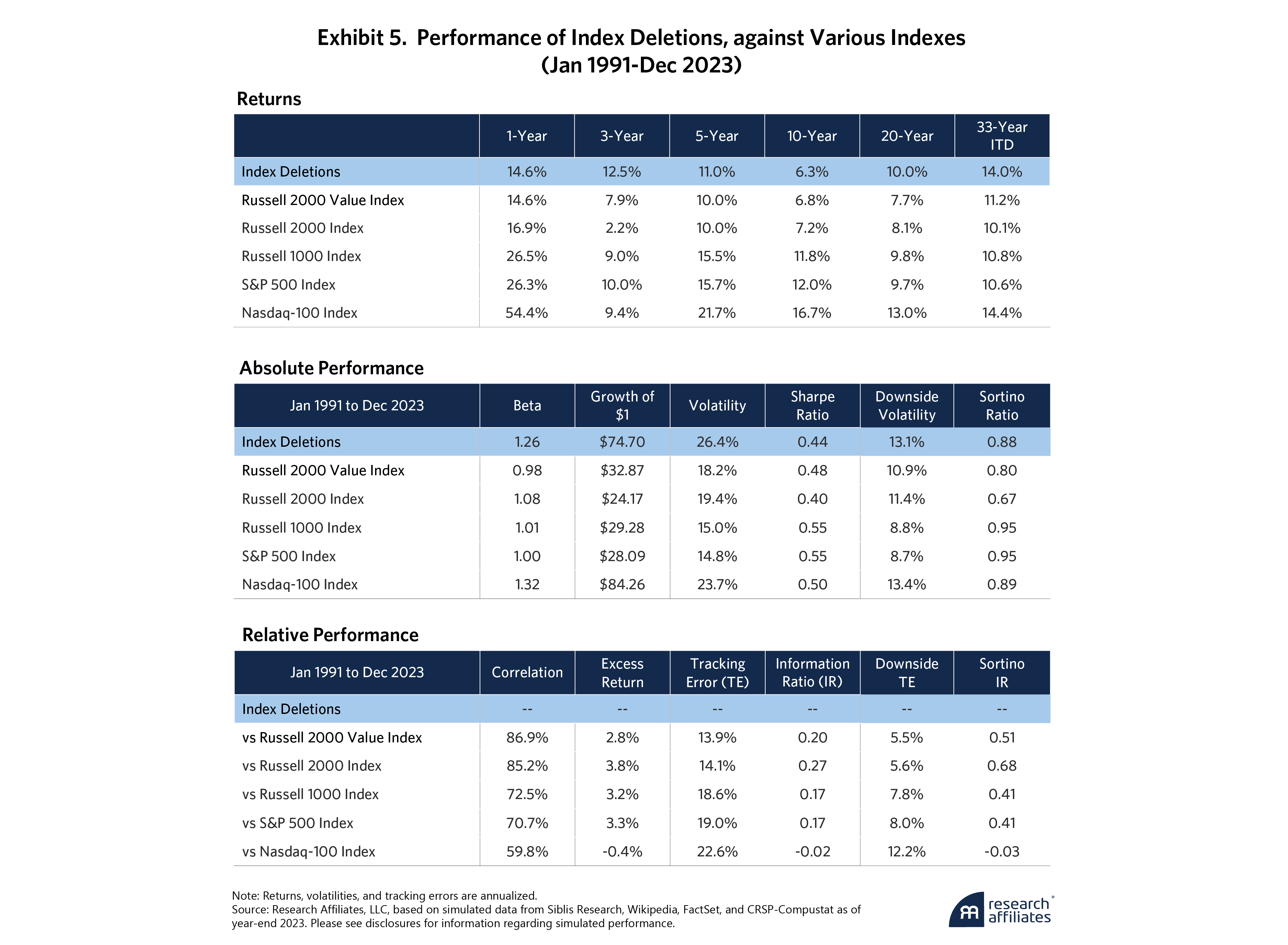
We carried out robustness checks to see whether our results were unique to published indexes like the S&P 500 and Russell 1000. We created hypothetical indexes of the 500 and 1,000 largest market-cap stocks to approximate the decisions of the S&P Index Committee and FTSE-Russell, respectively. 9 We call these “True Cap” indexes because the decision process is utterly trivial, with no subjectivity whatsoever.
Do dumped True Cap stocks also win in the years following their deletion? The results are surprisingly similar. In other words, this work is not in any way a criticism of the decisions that the major index providers are making. Quite the contrary. The alpha stems from the market inefficiencies created by the process of adding stocks that have recently soared onto the top 500 or top 1,000 list and deleting those that have tumbled off the list. There is no need to use the actual S&P, Russell, or Nasdaq indexes. A roster of “deletions” from any cap-selected and cap-weighted index will do!
Closing Observations
Of course, searching through data for relationships and fitting our preferred narrative to those data is not scientific method, it is backtesting. Even if the results are not data-mined, our inquiries thus far suffer from that flaw. To our knowledge, no one has managed a deletions strategy over the last 30 years. Too many in our industry use backtests to improve backtest results. That is an excellent way to develop a great backtest and a terrible way to win in the future. Scientific method requires we use data to test a prior hypothesis, not to refine the model.
Index producers aren’t especially subtle in their addition and deletion decisions. They inevitably add popular and expensive stocks and dump cheap and unloved ones. This simple truth provided much of the motivation for our launch of the RAFI concept, which selects and weights index stocks by the fundamental size, the macroeconomic footprint, of the underlying business. That the data demonstrates deletions historically outperform additions in the years after index reconstitution is no surprise to us.
Deletions historically outperform the market for at least five years post-breakup. Their annual outperformance is remarkably persistent and statistically significant, averaging over 5% per annum, compounded over the first five years.
The deletions strategy is a wonderful Occam’s razor to help identify the most depressed and unloved stocks. These may enjoy abnormally powerful performance rebounds, especially since the deletion process will only depress them more.
Such intriguing results have inspired us to take these investigations further and test these hypotheses in the real world. Today’s launch of the Research Affiliates Deletions Index (NIXT) is the first step in that direction. The NIXT index buys deletions from top 500 and top 1,000 market-cap weighted indexes, holds them for five years, and rebalances them annually to equal weight. This empirically has substantial overlap with the Russell 1000 and S&P 500 deletions and with very similar results. For the past 30 years, stocks have rebounded well after being dumped by an index. We’re looking forward to seeing if they maintain that resilience in the decades ahead.
Please read our disclosures concurrent with this publication below.
Introducing: Research Affiliates Deletions Index

Appendix 1. Separate Results for S&P 500, Russell 1000, and Nasdaq-100 Deletions
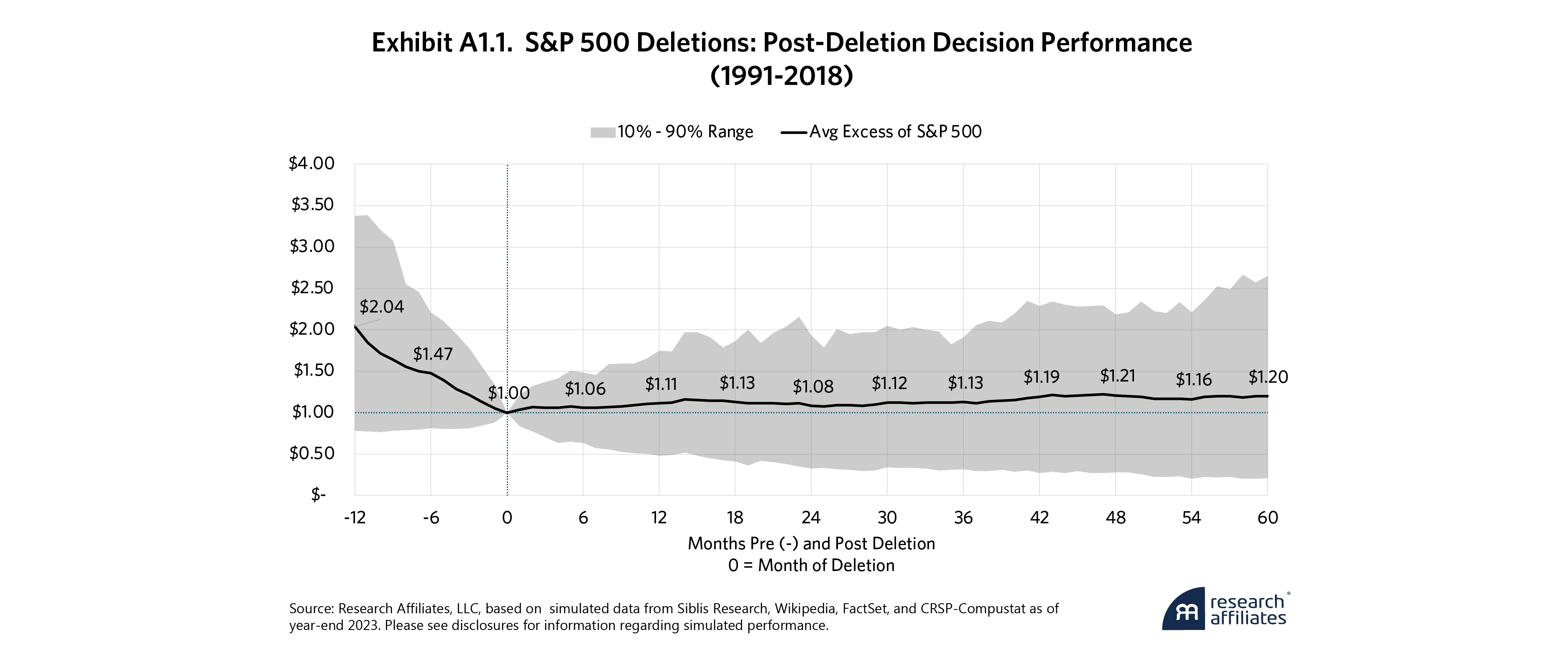
Appendix 2. Selecting the “Correct” Benchmark and Factor Attribution
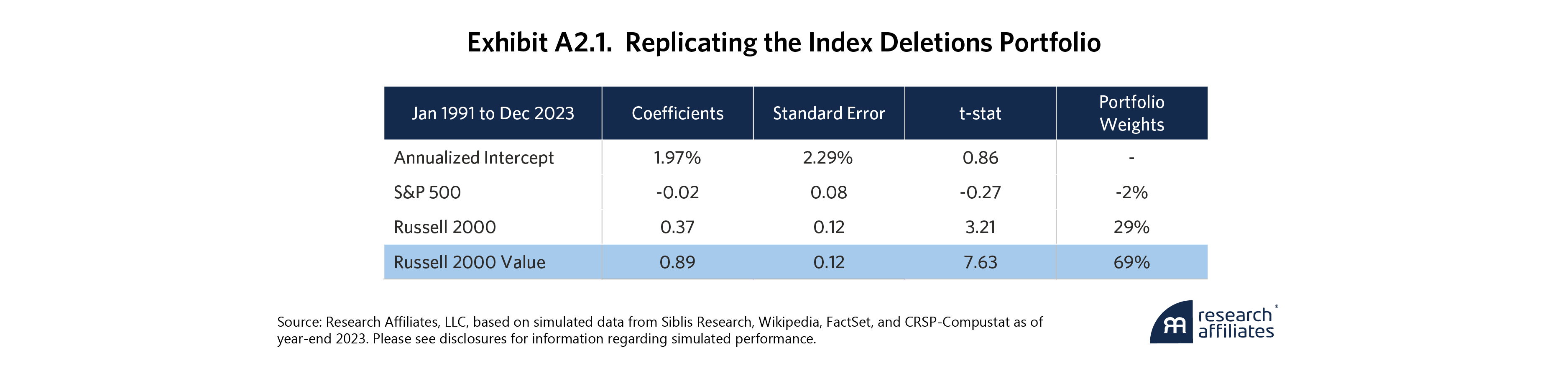
1. Arnott, Rob, Vitali Kalesnik, and Lillian Wu. “ Buy High and Sell Low with Index Funds! ” June 2018. Research Affiliates; Arnott, Rob, et al. “ Reimagining Index Funds .” 2023. Journal of Investment Management ; Arnott, Rob, et al. “ Earning Alpha by Avoiding the Index Rebalancing Crowd. ” 2023. Financial Analysts Journal .
2. See Arnott, Rob, and Engin Kose. “ Can Momentum Investing Be Saved? ” October 2017. Research Affiliates.
3. Chinco, Alex, and Marco Sammon. “ The Passive-Ownership Share Is Double What You Think It Is .” July 2024. Journal of Financial Economics .
4. Separate results for the S&P 500, Russell 1000, and Nasdaq 100 deletions are presented in the appendix. For simplicity, we start the clock at the end of the deletion month. Russell deletions underperform by more than S&P 500 deletions during their deletion year, with the deletions more than halving relative to the Russell 1000.
5. Arnott, Rob, et al. “ Earning Alpha by Avoiding the Index Rebalancing Crowd. ” 2023. Financial Analysts Journal .
6. Regression analysis indicates this is the “correct” benchmark. Per Table A2.1 , when we regress this deletions portfolio against the S&P 500, Russell 2000, and Russell 2000 Value, the majority weight (nearly 70%) is on the Russell 2000 Value. Nasdaq 100 deletions are a bit different. Since they are usually no longer S&P 500 members by the time they are dumped from the Nasdaq100, they are mid-cap value stocks. The Russell 2000 Value is still the better fit, however, because top 500 stocks dominate the Russell 1000 Value.
7. Hendrix, Kaitlin, Jerry Liu, and Trey Roberts. “ Measuring the Cost of Index Reconstitution: A 10-Year Perspective. ” 2024. Dimensional Fund Advisors
8. van der Beck, Philippe, Jean-Philippe Bouchaud, and Dario Villamaina. “ Ponzi Funds. ” 2024. S&P Global Market Intelligence via SSRN.
9. We also added 10% banding, so that we won’t introduce undue turnover. So stocks aren’t added until they are in the top 450 or 900, and deletions typically have to fall to rank 550 or 1100. The result closely resembles the S&P 500 and the Russell 1000 without intruding on their IP. Unsurprisingly, there is substantial overlap with S&P and Russell deletions.
Disclosures
The material contained in this document is for informational purposes only. This material is not intended as an offer or solicitation for the purchase or sale of any security or financial instrument, nor is it advice or a recommendation to enter into any transaction.
Certain performance information presented represents simulated performance or performance based on combined simulated index data (pre-index launch) and live index data (post-index launch). Indexes are unmanaged and cannot be invested in directly. Past simulated performance is no guarantee of future performance and does not represent actual performance of an investment product based on an index. No allowance has been made for trading costs, management fees, or other costs associated with asset management, as the information provided relates only to the index itself. Actual investment results will differ. The simulated data may have under- or over-compensated for the impact, if any, of certain market factors. Simulated returns may not reflect the impact that material economic and market factors might have had on the advisor's decision making if the advisor were actually managing clients' money. Simulated data is subject to the fact that it is designed with the benefit of hindsight. Simulated returns carry the risk that actual performance is not as depicted due to inaccurate predictive modeling. Simulated returns cannot predict how an investment strategy will perform in the future. Simulated returns should not be considered indicative of the skill of the advisor. Investors may experience loss of all or some of their investment.
© 2024 Research Affiliates, LLC. All rights reserved. Duplication or dissemination prohibited without prior written permission.
- Publications
- Journal Papers
- RA Multimedia
- How to Invest
Cookies Disclaimer
The content provided on this website is informational, subject to change and is not investment advice or any offer or solicitation for the purchase or sale of investments. Any use of this content is subject to and evidence of the user’s acceptance of all important legal disclosures, disclaimers, terms of use and provisions found at https://www.researchaffiliates.com/en_us/legal/privacy-policy, including the user’s complete release of liability for any use of the content, which may contain inaccuracies.
This site uses cookies on our website to distinguish you from other users of our website. Our objective is to optimize your experience when you browse our website and to continually improve our site. Consenting to the use of these conditions is not a condition of using the website, however, if you do not consent, you will be redirected to a static website with limited information. Please select “Accept” if you consent to our use of cookies, pursuant to our Cookies Policy, during your visit to our website. Otherwise, please select “Decline” if you do not consent to our use of cookies during your visit to our website. A copy of our Cookies Policy can be found at https://www.researchaffiliates.com/en_us/legal/privacy-policy .
You are now leaving the Research Affiliates, LLC website. The following link may contain information concerning investments, products or other information. Research Affiliates, LLC is not responsible for the accuracy or completeness of information on non-affiliated websites and does not make any representation regarding the advisability of investing in any investment fund or other investment product or vehicle. Importantly, Research Affiliates, LLC is not compensated for linking you to any non-affiliated website and instead is only compensated with an asset-based fee as either a licensor of intellectual property or a sub-adviser to an investment adviser. The material available on non-affiliated websites has been produced by entities that are not affiliated with Research Affiliates, LLC. Descriptions of, references to, or links to products or publications within any non-affiliated linked website does not imply endorsement or recommendation of that product or publication by Research Affiliates, LLC. Any opinions or recommendations from non-affiliated websites are solely those of the independent providers and are not the opinions or recommendations of Research Affiliates, LLC, which is not responsible for any inaccuracies or errors. THIS INFORMATION IS NOT AN OFFER TO BUY OR A SOLICITATION TO SELL ANY SECURITY OR INVESTMENT PRODUCT. SUCH AN OFFER OR SOLICITATION IS MADE ONLY BY THE SECURITIES’ OR INVESTMENT PRODUCTS’ ISSUER OR SPONSOR THROUGH A PROSPECTUS OR OTHER OFFERING DOCUMENTATION.

How to cite ChatGPT

Use discount code STYLEBLOG15 for 15% off APA Style print products with free shipping in the United States.
We, the APA Style team, are not robots. We can all pass a CAPTCHA test , and we know our roles in a Turing test . And, like so many nonrobot human beings this year, we’ve spent a fair amount of time reading, learning, and thinking about issues related to large language models, artificial intelligence (AI), AI-generated text, and specifically ChatGPT . We’ve also been gathering opinions and feedback about the use and citation of ChatGPT. Thank you to everyone who has contributed and shared ideas, opinions, research, and feedback.
In this post, I discuss situations where students and researchers use ChatGPT to create text and to facilitate their research, not to write the full text of their paper or manuscript. We know instructors have differing opinions about how or even whether students should use ChatGPT, and we’ll be continuing to collect feedback about instructor and student questions. As always, defer to instructor guidelines when writing student papers. For more about guidelines and policies about student and author use of ChatGPT, see the last section of this post.
Quoting or reproducing the text created by ChatGPT in your paper
If you’ve used ChatGPT or other AI tools in your research, describe how you used the tool in your Method section or in a comparable section of your paper. For literature reviews or other types of essays or response or reaction papers, you might describe how you used the tool in your introduction. In your text, provide the prompt you used and then any portion of the relevant text that was generated in response.
Unfortunately, the results of a ChatGPT “chat” are not retrievable by other readers, and although nonretrievable data or quotations in APA Style papers are usually cited as personal communications , with ChatGPT-generated text there is no person communicating. Quoting ChatGPT’s text from a chat session is therefore more like sharing an algorithm’s output; thus, credit the author of the algorithm with a reference list entry and the corresponding in-text citation.
When prompted with “Is the left brain right brain divide real or a metaphor?” the ChatGPT-generated text indicated that although the two brain hemispheres are somewhat specialized, “the notation that people can be characterized as ‘left-brained’ or ‘right-brained’ is considered to be an oversimplification and a popular myth” (OpenAI, 2023).
OpenAI. (2023). ChatGPT (Mar 14 version) [Large language model]. https://chat.openai.com/chat
You may also put the full text of long responses from ChatGPT in an appendix of your paper or in online supplemental materials, so readers have access to the exact text that was generated. It is particularly important to document the exact text created because ChatGPT will generate a unique response in each chat session, even if given the same prompt. If you create appendices or supplemental materials, remember that each should be called out at least once in the body of your APA Style paper.
When given a follow-up prompt of “What is a more accurate representation?” the ChatGPT-generated text indicated that “different brain regions work together to support various cognitive processes” and “the functional specialization of different regions can change in response to experience and environmental factors” (OpenAI, 2023; see Appendix A for the full transcript).
Creating a reference to ChatGPT or other AI models and software
The in-text citations and references above are adapted from the reference template for software in Section 10.10 of the Publication Manual (American Psychological Association, 2020, Chapter 10). Although here we focus on ChatGPT, because these guidelines are based on the software template, they can be adapted to note the use of other large language models (e.g., Bard), algorithms, and similar software.
The reference and in-text citations for ChatGPT are formatted as follows:
- Parenthetical citation: (OpenAI, 2023)
- Narrative citation: OpenAI (2023)
Let’s break that reference down and look at the four elements (author, date, title, and source):
Author: The author of the model is OpenAI.
Date: The date is the year of the version you used. Following the template in Section 10.10, you need to include only the year, not the exact date. The version number provides the specific date information a reader might need.
Title: The name of the model is “ChatGPT,” so that serves as the title and is italicized in your reference, as shown in the template. Although OpenAI labels unique iterations (i.e., ChatGPT-3, ChatGPT-4), they are using “ChatGPT” as the general name of the model, with updates identified with version numbers.
The version number is included after the title in parentheses. The format for the version number in ChatGPT references includes the date because that is how OpenAI is labeling the versions. Different large language models or software might use different version numbering; use the version number in the format the author or publisher provides, which may be a numbering system (e.g., Version 2.0) or other methods.
Bracketed text is used in references for additional descriptions when they are needed to help a reader understand what’s being cited. References for a number of common sources, such as journal articles and books, do not include bracketed descriptions, but things outside of the typical peer-reviewed system often do. In the case of a reference for ChatGPT, provide the descriptor “Large language model” in square brackets. OpenAI describes ChatGPT-4 as a “large multimodal model,” so that description may be provided instead if you are using ChatGPT-4. Later versions and software or models from other companies may need different descriptions, based on how the publishers describe the model. The goal of the bracketed text is to briefly describe the kind of model to your reader.
Source: When the publisher name and the author name are the same, do not repeat the publisher name in the source element of the reference, and move directly to the URL. This is the case for ChatGPT. The URL for ChatGPT is https://chat.openai.com/chat . For other models or products for which you may create a reference, use the URL that links as directly as possible to the source (i.e., the page where you can access the model, not the publisher’s homepage).
Other questions about citing ChatGPT
You may have noticed the confidence with which ChatGPT described the ideas of brain lateralization and how the brain operates, without citing any sources. I asked for a list of sources to support those claims and ChatGPT provided five references—four of which I was able to find online. The fifth does not seem to be a real article; the digital object identifier given for that reference belongs to a different article, and I was not able to find any article with the authors, date, title, and source details that ChatGPT provided. Authors using ChatGPT or similar AI tools for research should consider making this scrutiny of the primary sources a standard process. If the sources are real, accurate, and relevant, it may be better to read those original sources to learn from that research and paraphrase or quote from those articles, as applicable, than to use the model’s interpretation of them.
We’ve also received a number of other questions about ChatGPT. Should students be allowed to use it? What guidelines should instructors create for students using AI? Does using AI-generated text constitute plagiarism? Should authors who use ChatGPT credit ChatGPT or OpenAI in their byline? What are the copyright implications ?
On these questions, researchers, editors, instructors, and others are actively debating and creating parameters and guidelines. Many of you have sent us feedback, and we encourage you to continue to do so in the comments below. We will also study the policies and procedures being established by instructors, publishers, and academic institutions, with a goal of creating guidelines that reflect the many real-world applications of AI-generated text.
For questions about manuscript byline credit, plagiarism, and related ChatGPT and AI topics, the APA Style team is seeking the recommendations of APA Journals editors. APA Style guidelines based on those recommendations will be posted on this blog and on the APA Style site later this year.
Update: APA Journals has published policies on the use of generative AI in scholarly materials .
We, the APA Style team humans, appreciate your patience as we navigate these unique challenges and new ways of thinking about how authors, researchers, and students learn, write, and work with new technologies.
American Psychological Association. (2020). Publication manual of the American Psychological Association (7th ed.). https://doi.org/10.1037/0000165-000
Related and recent
Comments are disabled due to your privacy settings. To re-enable, please adjust your cookie preferences.
APA Style Monthly
Subscribe to the APA Style Monthly newsletter to get tips, updates, and resources delivered directly to your inbox.
Welcome! Thank you for subscribing.
APA Style Guidelines
Browse APA Style writing guidelines by category
- Abbreviations
- Bias-Free Language
- Capitalization
- In-Text Citations
- Italics and Quotation Marks
- Paper Format
- Punctuation
- Research and Publication
- Spelling and Hyphenation
- Tables and Figures
Full index of topics
Information
- Author Services
Initiatives
You are accessing a machine-readable page. In order to be human-readable, please install an RSS reader.
All articles published by MDPI are made immediately available worldwide under an open access license. No special permission is required to reuse all or part of the article published by MDPI, including figures and tables. For articles published under an open access Creative Common CC BY license, any part of the article may be reused without permission provided that the original article is clearly cited. For more information, please refer to https://www.mdpi.com/openaccess .
Feature papers represent the most advanced research with significant potential for high impact in the field. A Feature Paper should be a substantial original Article that involves several techniques or approaches, provides an outlook for future research directions and describes possible research applications.
Feature papers are submitted upon individual invitation or recommendation by the scientific editors and must receive positive feedback from the reviewers.
Editor’s Choice articles are based on recommendations by the scientific editors of MDPI journals from around the world. Editors select a small number of articles recently published in the journal that they believe will be particularly interesting to readers, or important in the respective research area. The aim is to provide a snapshot of some of the most exciting work published in the various research areas of the journal.
Original Submission Date Received: .
- Active Journals
- Find a Journal
- Proceedings Series
- For Authors
- For Reviewers
- For Editors
- For Librarians
- For Publishers
- For Societies
- For Conference Organizers
- Open Access Policy
- Institutional Open Access Program
- Special Issues Guidelines
- Editorial Process
- Research and Publication Ethics
- Article Processing Charges
- Testimonials
- Preprints.org
- SciProfiles
- Encyclopedia

Article Menu

- Subscribe SciFeed
- Recommended Articles
- Google Scholar
- on Google Scholar
- Table of Contents
Find support for a specific problem in the support section of our website.
Please let us know what you think of our products and services.
Visit our dedicated information section to learn more about MDPI.
JSmol Viewer
Agrifood sustainability transitions in firms and industry: a bibliographic analysis of research themes.

1. Introduction
1.1. the global agrifood system, 1.2. agrifood sustainability transitions, 1.3. sustainability transitions research, 1.4. agrifood sustainability transitions—firms and industries.
- Evaluate and describe the current state of the literature on the (STRN) theme of firms and industries.
- Identify gaps and propose areas for future research to address these areas.
- Is the theme of “industries and firms” still a marginal topic within the agrifood sustainability transition literature?
- How has the agrifood sustainability transition literature on “industries and firms” evolved?
- What are the main themes and topics within this theme and what gaps exist within this literature?
- What are the potential areas for future research on firms and industries in sustainability transitions?
2. Materials and Methods
2.1. data collection, 2.2. data analysis, 3.1. descriptive analysis of publications, 3.2. co-citation analysis, 3.3. co-occurrence of keyword analysis, 3.3.1. agriculture, 3.3.2. innovation, 3.3.3. governance, 3.3.4. food system, 3.3.5. agroecology, 4. discussion and conclusions, 4.1. key findings, 4.2. research gaps and future research, 4.3. theoretical contribution, 4.4. implications for managers and policymakers, author contributions, data availability statement, conflicts of interest, appendix a. list of papers.
| | | |||
| Food safety risks, disruptive events and alternative beef production: A case study of agricultural transition in Alberta [ ] | Alternative food producers, organic production | Case study | Alternative beef producers in Alberta, Canada | Sustainable transitions theory. Multi-level perspective |
| The power of corporate lock-ins and how they shape digital agriculture in Germany [ ] | Critique of agro-industrial farming models | Case study | Stakeholders in the German agriculture industry | Political economy |
| Translating Environmental Potential to Economic Reality: Assessment of Commercial Aquaponics through Sustainability Transitions [ ] | Circular economy | Semi-structured interviews, literature and policy review | American aquaponic producers | Technology innovation system framework, multi-level perspective (MLP) |
| The politics of expertise in assessing alternatives to glyphosate in France [ ] | Alternatives to pesticide use (glyphosate) | Case study | Actors in the agricultural sector and experts in pesticide policy | Boundary work concept |
| Integrating sustainability transitions and food systems research to examine consultation failures in Canadian food policymaking [ ] | Food sovereignty | Case study | Canadian food policymaking | Multi-level perspective (MLP) |
| Adaptive transition management for transformations to agricultural sustainability in the Karnali mountains of Nepal [ ] | Resilience, biodiversity | Single case study | Karnali mountains Nepal | Multi-level perspective (MLP). Adaptive transition management |
| Connecting business with the agricultural landscape: business strategies for sustainable rural development [ ] | Agricultural landscape sustainability | Multiple case studies | Four cases of business-landscape engagement | Landscape perspective |
| Conceptualizing sustainable food entrepreneurship [ ] | Relocalisation | Literature review and case study | Dutch city region of Almere-Flevoland | Sustainable food entrepreneurship framework |
| Unlearning in sustainability transitions: Insight from two Dutch community-supported agriculture farms [ ] | Community-supported agriculture | Case study | Phase-out and ban of battery cages for laying hens in the Netherlands | Political economy |
| Agriculture 4.0 and climate change in Brazil [ ] | Low carbon/climate change | Single case study | Brazilian agribusiness | Multi-level perspective (MLP) |
| Synthesising the diversity of European agri-food networks: A meta-study of actors and power-laden interactions [ ] | None | Meta-analysis | European case studies | |
| Spheres of transformation: Exploring personal, political and practical drivers of farmer agency and behaviour change in the Netherlands [ ] | Regenerative farming practices | Case study | Dutch farmers adopting regenerative farming practices | Spheres of transformation |
| Giagnocavo et al., 2022 [ ] | Agroecology | |||
| McInnes [ ] | Food system | |||
| Farhangi et al., 2020 [ ] | Food system | |||
| Manuel-Navarrete and Gallopín, 2012 [ ] | Governance | |||
| van der Gaast et al., 2022 [ ] | Innovation | |||
| van Oers et al., 2021 [ ] | Food system | |||
| Kinniburgh, 2023 [ ] | Governance | |||
| Selecting technologies to engage in sustainability transitions—A multi-stakeholder perspective [ ] | Sustainability transitions from a fossil-based towards a bio-based economy | Mixed-method research design | Stakeholders from multiple stages of the value chain | Transition theory. Selection criteria for sustainability-orientated technologies SOT’s |
| Enacting transitions—the combined effect of multiple niches in whole system reconfiguration [ ] | Organic farming | Ethnographic study and archival work | Drôme valley in southeast France | Multi-level perspective (MLP) |
| Relation between innovation and sustainability in the agro-food system [ ] | UN sustainable development goals | Literature review | Sustainability literature | Multi-level perspective (MLP) |
| Transformational adaptation on the farm: Processes of change and persistence in transitions to ‘climate-smart’ regenerative agriculture [ ] | Regenerative agriculture. | Semi-structured interviews, participant observation, and document analysis | Farmers grazing sheep or cattle New South Wales, Australia | sustainability transitions framework |
| Agency in regime destabilization through the selection environment: The Finnish food system’s sustainability transition [ ] | Nutrient recycling, vegetarian diet and organic foods | Discourse analysis | Finnish food system | Tripple embeddedness framework. Agency in the selection environment |
| A framework of disruptive sustainable innovation: An example of the Finnish food system [ ] | Reduced meat consumption, local food, direct farm sales, organic production | Multiple case studies | Finnish food system | Practice-based view on disruptive innovation, multi-level perspective (MLP) |
| The diffusion of climate-smart agricultural innovations: Systems-level factors that inhibit sustainable entrepreneurial action [ ] | Earth system biophysical thresholds, climate, smart agriculture | Multi-level perspective (MLP) | Semi-structured interviews. Climate-smart providers and policymakers A–Z | Multi-level perspective (MLP). Entrepreneurial ecosystem perspective |
| Lost in mainstreaming? Agrifood and urban mobility grassroots innovations with multiple pathways and outcomes [ ] | Grassroots innovation: Fairtrade, organic, veganism, car-sharing, cycling and shared spaces | Multiple case studies | Grassroots innovations | Sustainability transitions framework |
| Designing coupled innovations for the sustainability transition of agrifood systems. [ ] | Reduced energy use, increased biodiversity, improved soil and water quality, decreased pesticide use, preventing nutritional deficits and obesity | Multiple case study | Examples of coupled innovations | Multi-level perspective (MLP), innovative design theory |
| The role of supply chains for the sustainability transformation of global food systems: A large-scale, systematic review of food cold chains [ ] | Sustainable development goals | Literature review | Food cold chain | None |
| Conceptualizing sustainable food entrepreneurship [ ] | Relocalisation | Literature review and case study | Dutch city region of Almere-Flevoland | Sustainable food entrepreneurship framework |
| The governance features of social enterprise and social network activities of collective food buying groups [ ] | Collective food-buying groups | Semi-structured questionnaire | 104 collective buying groups, Belgium | Sustainability transitions theory |
| A methodological framework to initiate and design transition governance processes [ ] | Alternate food systems, urban farming, community gardens, local production | Single case study | Sustainable food systems in Ontario, Canada | Multi-level learning processes |
| Translating Environmental Potential to Economic Reality: Assessment of Commercial Aquaponics through Sustainability Transitions Theory [ ] | Aquaponic food production, circular economy | Semi-structured interviews | 25 North American producers | Technological innovation system (TIS) assessment multi-level perspective (MLP) |
| The politics of expertise in assessing alternatives to glyphosate in France [ ] | Reduced pesticide use. | Single case study | French pesticide regulation on glyphosate alternatives | Concepts of co-production and boundary work |
| Lost in mainstreaming? Agrifood and urban mobility grassroots innovations with multiple pathways and outcomes [ ] | Grassroots innovation: Fairtrade, organic, veganism, carsharing, cycling and shared space | Multiple case studies | Grassroots innovations | Sustainability transitions framework |
| Feeding the world sustainably: Knowledge governance and sustainable agriculture in the Argentine Pampas [ ] | No-till practices. Reduced soil degradation and ecosystem disruption | Single case study | Argentine farmers | Actor-centred approach |
| Crafting actionable knowledge on ecological intensification: Lessons from co-innovation approaches in Uruguay and Europe [ ] | Ecological intensification | Six case studies from three co-innovation projects | Co-innovation research projects in Uruguay and Europe | Complex adaptive systems, social learning, dynamic monitoring and evaluation |
| A Framework for Sustainability Transition: The Case of Plant-Based Diets [ ] | Social, economic, environmental, cultural, and ethical dimensions of sustainability | Case study | Plant-based diets | Sustainability transitions theory |
| Outside-in and bottom-up: Using sustainability transitions to understand the development phases of mainstreaming plant-based in the food sector in a meat and dairy focused economy [ ] | Planetary boundaries, reducing GHG, land use change and biodiversity loss | Semi-structured interviews with businesses and experts | Denmark plant-based products | Multi-level perspective (MLP) |
| Structuring tensions and key relations of Montreal seasonal food markets in the sustainability transition of the agri-food sector [ ] | Local food, seasonal production | Action research, case study | Three Montreal seasonal food markets | Multi-level perspective (MLP) |
| Systemic ethics and inclusive governance: two key prerequisites for sustainability transitions of agri-food systems [ ] | Small-scale production, local food | Case study | Belgian supermarkets | Sustainability transitions perspective |
| Systemic ethics and inclusive governance: Two key prerequisites for sustainability transitions of agri-food systems [ ] | Local, low-input, small-scale farmers’ products | Case study | Local sourcing in Belgian supermarkets | Sustainability transitions perspective |
| Incumbent entry modes and entry timing in sustainable niches: The plant-based protein transition in the United States, the Netherlands and the United Kingdom [ ] | Plant-based meat substitutes | Multiple case studies | Firms adopting plant-based meat substitutes in the US, the Netherlands and the UK | Entry mode theory |
| Productivity growth as a barrier to a sustainability transition [ ] | Small-scale artisan bakers | Case study | Australian baking industry | Multifactor productivity |
| Digital fooding, cashless marketplaces and reconnection in intermediated third places: Conceptualizing metropolitan food provision in the age of prosumption [ ] | Sustainable food systems | Desktop analysis | Review of special-issue literature in the journal Agriculture and Human Values | Sustainable development goals |
| The evolutionary emergence of quintuple helix coalitions: A case study of place-based sustainability transition [ ] | Local food production, short supply chains, production and sales of ancient grains (wheat varieties) | Case study | Value chain of ancient wheat varieties in Tuscany | Triple helix |
| High-tech urban agriculture in Amsterdam: An actor–network analysis [ ] | High-tech urban agriculture | Case study | Amsterdam: High-tech urban agriculture | Actor-network theory/multi-level perspective (MLP), technology-driven transition framework |
| Incumbents’ capabilities for sustainability-oriented innovation in the Norwegian food sector—An integrated framework [ ] | UN sustainable development goals | Multiple case study | Norwegian food sector | Theory of dynamic capabilities |
| China and changing food trends: A sustainability transition perspective [ ] | Reduced environmental footprint and better human diets. Planetary boundaries | Desktop analysis of literature and secondary data | Online sources related to major societal shifts in food consumption and production | Transition theories |
| Integrating sustainability transitions and food systems research to examine consultation failures in Canadian food policymaking [ ] | Food sovereignty | Case study | Canadian food policymaking | Multi-level perspective (MLP) |
| Digital fooding, cashless marketplaces and reconnection in intermediated third places: Conceptualizing metropolitan food provision in the age of prosumption [ ] | Local food (>250 km) | Case study | Ruche digital platform for food provisioning in France | Prosumption |
| Unlearning in sustainability transitions: Insight from two Dutch community-supported agriculture farms [ ] | Alternative food networks | Case study | Two Dutch community-supported agriculture groups | Organisational change theory, sustainability transitions perspective |
| A new green revolution or agribusiness as usual? Uncovering alignment issues and potential transition complications in agri-food system transitions [ ] | Identified issues: agrochemical use, Biodiversity, antibiotic uses | Vision documents for Dutch agricultural transition | Dutch agrifood system | Mission-orientated perspective, visioning |
| Sustainability buckets: A flexible heuristic for facilitating strategic investment on place-dependent sustainability narratives [ ] | Nourish the body, nourish the planet, socially just relationships, circular economy and economic viability | Two case studies | New Zealand egg sector and honey distributor | Multi-level perspective (MLP), sustainability cultures |
| Bui et al., 2019 [ ] | Innovation | |||
| Marletto and Sillig, 2019 [ ] | Innovation | |||
| Davidson et al., 2016 [ ] | Agriculture | |||
| Reconnecting farmers with Nature through agroecological transitions: Interacting niches and experimentation and the role of agricultural knowledge and innovation systems [ ] | Agroecology | Case study | Greenhouse sector, Almeria, Spain | Multi-level perspective (MLP), agroecological frameworks |
| Translating environmental potential to economic reality: Assessment of commercial aquaponics through sustainability transitions theory [ ] | Sustaining ecosystem services | Vision documents for Dutch agricultural transition | Dutch agrifood system | Mission-orientated perspective, visioning |
| The role of consumer-citizens and connectedness to nature in the sustainable transition to agroecological food systems: The mediation of innovative business models and a multi-level perspective (MLP) [ ] | Agroecology | Theoretical | Multi-level perspective (MLP) | |
| Sustainability transitions in the agrifood sector: How ecology affects transition dynamics [ ] | Biodiversity enhancement | Case study of four biodiversity initiatives | Dutch dairy sector | Multi-level perspective (MLP), innovation system perspective |
| Tackling material dependency in sustainability transition: Rationales and insights from the agriculture sector [ ] | Addressing the ecological crisis | Descriptive | Agriculture | Material dependency |
| Horn et al., 2023 [ ] | Governance/Agriculture | |||
- Ahmad, K. Global population will increase to nine billion by 2050. Lancet 2001 , 357 , 864. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Boliko, M.C. FAO and the situation of food security and nutrition in the world. J. Nutr. Sci. Vitaminol. 2019 , 65 , 4–8. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Ezeh, A.C.; Bongaarts, J.; Mberu, B. Global population trends and policy options. Lancet 2012 , 380 , 142–148. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- UNICEF. Brief to the State of Food Security and Nutrition in the World 2022 ; United Nations Children’s Fund: New York, NY, USA, 2022. [ Google Scholar ]
- United Nations. World Population Prospects 2022, Online Edition ; Department of Economic and Social Affairs, Population Division: New York, NY, USA, 2022; Available online: https://population.un.org/wpp/ (accessed on 30 August 2023).
- Godfray, H.C.J.; Crute, I.R.; Haddad, L.; Lawrence, D.; Muir, J.F.; Nisbett, N.; Pretty, J.; Robinson, S.; Toulmin, C.; Whiteley, R. The future of the global food system. Philos. Trans. R. Soc. Biol. Sci. 2010 , 365 , 2769–2777. [ Google Scholar ] [ CrossRef ]
- OECD. How We Feed the World Today ; OECD: Paris, France, 2022; Available online: https://www.oecd.org/agriculture/understanding-the-global-food-system/how-we-feed-the-world-today/ (accessed on 20 January 2024)).
- Gaitán-Cremaschi, D.; Klerkx, L.; Duncan, J.; Trienekens, J.H.; Huenchuleo, C.; Dogliotti, S.; Contesse, M.E.; Rossing, W.A.H. Characterizing diversity of food systems in view of sustainability transitions. A review. Agron. Sustain. Dev. 2019 , 39 , 1. [ Google Scholar ] [ CrossRef ]
- Hackfort, S. Unlocking sustainability? The power of corporate lock-ins and how they shape digital agriculture in Germany. J. Rural Stud. 2023 , 101 , 103065. [ Google Scholar ] [ CrossRef ]
- Mehrabi, S.; Perez-Mesa, J.C.; Giagnocavo, C. The Role of Consumer-Citizens and Connectedness to Nature in the Sustainable Transition to Agroecological Food Systems: The mediation of innovative business models and a multi-level perspective. Agriculture 2022 , 12 , 203. [ Google Scholar ] [ CrossRef ]
- Wojtynia, N.; van Dijk, J.; Derks, M.; Groot Koerkamp, P.; Hekkert, M. A new green revolution or agribusiness as usual? Uncovering alignment issues and potential transition complications in agri-food system transitions. Agron. Sustain. Dev. 2021 , 41 , 77. [ Google Scholar ] [ CrossRef ]
- Wojtynia, N.; van Dijk, J.; Derks, M.; Groot Koerkamp, P.; Hekkert, M. Spheres of transformation: Exploring personal, political and practical drivers of farmer agency and behaviour change in the Netherlands. Environ. Innov. Soc. Transit. 2023 , 49 , 100776. [ Google Scholar ] [ CrossRef ]
- Marinova, D.; Bogueva, D.; Wu, Y.; Guo, X. China and changing food trends: A sustainability transition perspective. Ukr. Food J. 2022 , 11 , 126–147. [ Google Scholar ] [ CrossRef ]
- Meynard, J.-M.; Jeuffroy, M.-H.; Le Bail, M.; Lefèvre, A.; Magrini, M.-B.; Michon, C. Designing coupled innovations for the sustainability transition of agrifood systems. Agric. Syst. 2017 , 157 , 330–339. [ Google Scholar ] [ CrossRef ]
- Vinnari, M.; Vinnari, E. A framework for sustainability transition: The case of plant-based diets. J. Agric. Environ. Ethics 2014 , 27 , 369–396. [ Google Scholar ] [ CrossRef ]
- Willett, W.; Rockström, J.; Loken, B.; Springmann, M.; Lang, T.; Vermeulen, S.; Garnett, T.; Tilman, D.; DeClerck, F.; Wood, A. Food in the Anthropocene: The EAT–Lancet Commission on healthy diets from sustainable food systems. Lancet 2019 , 393 , 447–492. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- El Bilali, H. Research on agro-food sustainability transitions: A systematic review of research themes and an analysis of research gaps. J. Clean. Prod. 2019 , 221 , 353–364. [ Google Scholar ] [ CrossRef ]
- FAO. Global Food Losses and Food Waste—Extent, Causes and Prevention ; Food and Agriculture Organization of the United Nations: Rome, Italy, 2011; Available online: https://www.fao.org/4/mb060e/mb060e.pdf (accessed on 10 November 2023).
- Kummu, M.; De Moel, H.; Porkka, M.; Siebert, S.; Varis, O.; Ward, P.J. Lost food, wasted resources: Global food supply chain losses and their impacts on freshwater, cropland, and fertiliser use. Sci. Total Environ. 2012 , 438 , 477–489. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Moult, J.; Allan, S.; Hewitt, C.; Berners-Lee, M. Greenhouse gas emissions of food waste disposal options for UK retailers. Food Policy 2018 , 77 , 50–58. [ Google Scholar ] [ CrossRef ]
- Guh, D.P.; Zhang, W.; Bansback, N.; Amarsi, Z.; Birmingham, C.L.; Anis, A.H. The incidence of co-morbidities related to obesity and overweight: A systematic review and meta-analysis. BMC Public Health 2009 , 9 , 88. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- WHO. Obesity and Overweight ; WHO: Geneva, Switzerland, 2021; Available online: https://www.who.int/news-room/fact-sheets/detail/obesity-and-overweight (accessed on 13 September 2023).
- Bui, S. Enacting transitions—The combined effect of multiple niches in whole system reconfiguration. Sustainability 2021 , 13 , 6135. [ Google Scholar ] [ CrossRef ]
- Aschemann-Witzel, J.; Mulders, M.D.G.H.; Mouritzen, S.L.T. Outside-in and bottom-up: Using sustainability transitions to understand the development phases of mainstreaming plant-based in the food sector in a meat and dairy focused economy. Technol. Forecast. Soc. Chang. 2023 , 197 , 122906. [ Google Scholar ] [ CrossRef ]
- Béné, C. Why the Great Food Transformation may not happen–A deep-dive into our food systems’ political economy, controversies and politics of evidence. World Dev. 2022 , 154 , 105881. [ Google Scholar ] [ CrossRef ]
- Kuokkanen, A.; Uusitalo, V.; Koistinen, K. A framework of disruptive sustainable innovation: An example of the Finnish food system. Technol. Anal. Strateg. Manag. 2019 , 31 , 749–764. [ Google Scholar ] [ CrossRef ]
- Sustainability Transition Research Network. (2021, 2022). Available online: https://transitionsnetwork.org/ (accessed on 28 October 2023).
- Gosnell, H.; Gill, N.; Voyer, M. Transformational adaptation on the farm: Processes of change and persistence in transitions to ‘climate-smart’regenerative agriculture. Glob. Environ. Chang. 2019 , 59 , 101965. [ Google Scholar ] [ CrossRef ]
- El Bilali, H.; Strassner, C.; Ben Hassen, T. Sustainable agri-food systems: Environment, economy, society, and policy. Sustainability 2021 , 13 , 6260. [ Google Scholar ] [ CrossRef ]
- Geels, F.W. Technological transitions as evolutionary reconfiguration processes: A multi-level perspective and a case-study. Res. Policy 2002 , 31 , 1257–1274. [ Google Scholar ] [ CrossRef ]
- Van den Ende, J.; Kemp, R. Technological transformations in history: How the computer regime grew out of existing computing regimes. Res. Policy 1999 , 28 , 833–851. [ Google Scholar ] [ CrossRef ]
- Verbong, G.; Geels, F. The ongoing energy transition: Lessons from a socio-technical, multi-level analysis of the Dutch electricity system (1960–2004). Energy Policy 2007 , 35 , 1025–1037. [ Google Scholar ] [ CrossRef ]
- Lachman, D.A. A survey and review of approaches to study transitions. Energy Policy 2013 , 58 , 269–276. [ Google Scholar ] [ CrossRef ]
- Loorbach, D.; Frantzeskaki, N.; Avelino, F. Sustainability transitions research: Transforming science and practice for societal change. Annu. Rev. Environ. Resour. 2017 , 42 , 599–626. [ Google Scholar ] [ CrossRef ]
- Köhler, J.; Geels, F.W.; Kern, F.; Markard, J.; Onsongo, E.; Wieczorek, A.; Alkemade, F.; Avelino, F.; Bergek, A.; Boons, F.; et al. An agenda for sustainability transitions research: State of the art and future directions. Environ. Innov. Soc. Transit. 2019 , 31 , 1–32. [ Google Scholar ] [ CrossRef ]
- Visser, M.; van Eck, N.J.; Waltman, L. Large-scale comparison of bibliographic data sources: Scopus, Web of Science, Dimensions, Crossref, and Microsoft Academic. Quant. Sci. Stud. 2021 , 2 , 20–41. [ Google Scholar ] [ CrossRef ]
- Van Eck, N.; Waltman, L. Software survey: VOSviewer, a computer program for bibliometric mapping. Scientometrics 2010 , 84 , 523–538. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Bui, S.; Costa, I.; De Schutter, O.; Dedeurwaerdere, T.; Hudon, M.; Feyereisen, M. Systemic ethics and inclusive governance: Two key prerequisites for sustainability transitions of agri-food systems. Agric. Hum. Values 2019 , 36 , 277–288. [ Google Scholar ] [ CrossRef ]
- Williams, T.G.; Bui, S.; Conti, C.; Debonne, N.; Levers, C.; Swart, R.; Verburg, P.H. Synthesising the diversity of European agri-food networks: A meta-study of actors and power-laden interactions. Glob. Environ. Chang. 2023 , 83 , 102746. [ Google Scholar ] [ CrossRef ]
- Bulah, B.M.; Tziva, M.; Bidmon, C.; Hekkert, M.P. Incumbent entry modes and entry timing in sustainable niches: The plant-based protein transition in the United States, Netherlands, and United Kingdom. Environ. Innov. Soc. Transit. 2023 , 48 , 100735. [ Google Scholar ] [ CrossRef ]
- Vermunt, D.A.; Negro, S.O.; van Laerhoven, F.; Verweij, P.A.; Hekkert, M.P. Sustainability transitions in the agri-food sector: How ecology affects transition dynamics. Environ. Innov. Soc. Transit. 2020 , 36 , 236–249. [ Google Scholar ] [ CrossRef ]
- Dedeurwaerdere, T.; De Schutter, O.; Hudon, M.; Mathijs, E.; Annaert, B.; Avermaete, T.; Bleeckx, T.; de Callataÿ, C.; De Snijder, P.; Fernández-Wulff, P.; et al. The governance features of social enterprise and social network activities of collective food buying groups. Ecol. Econ. 2017 , 140 , 123–135. [ Google Scholar ] [ CrossRef ]
- van Oers, L.; Feola, G.; Moors, E.; Runhaar, H. The politics of deliberate destabilisation for sustainability transitions. Environ. Innov. Soc. Transit. 2021 , 40 , 159–171. [ Google Scholar ] [ CrossRef ]
- van Oers, L.; Feola, G.; Runhaar, H.; Moors, E. Unlearning in sustainability transitions: Insight from two Dutch community-supported agriculture farms. Environ. Innov. Soc. Transit. 2023 , 46 , 100693. [ Google Scholar ] [ CrossRef ]
- Giagnocavo, C.; de Cara-García, M.; González, M.; Juan, M.; Marín-Guirao, J.I.; Mehrabi, S.; Rodríguez, E.; van der Blom, J.; Crisol-Martínez, E. Reconnecting farmers with Nature through agroecological transitions: Interacting niches and experimentation and the role of agricultural knowledge and innovation systems. Agriculture 2022 , 12 , 137. [ Google Scholar ] [ CrossRef ]
- Kuokkanen, A.; Nurmi, A.; Mikkilä, M.; Kuisma, M.; Kahiluoto, H.; Linnanen, L. Agency in regime destabilization through the selection environment: The Finnish food system’s sustainability transition. Res. Policy 2018 , 47 , 1513–1522. [ Google Scholar ] [ CrossRef ]
- Elsevier. Agricultural Systems—Aims & Scope ; Elsevier: Amsterdam, The Netherlands, 2024; Available online: https://www.sciencedirect.com/journal/agricultural-systems (accessed on 30 January 2024).
- El Bilali, H. Relation between innovation and sustainability in the agro-food system. Ital. J. Food Sci. 2018 , 30 , 200–225. [ Google Scholar ]
- Pant, L.P.; Bahadur, K.; Fraser, E.D.; Shrestha, P.K.; Lama, A.B.; Jirel, S.K.; Chaudhary, P. Adaptive transition management for transformations to agricultural sustainability in the Karnali mountains of Nepal. Agroecol. Sustain. Food Syst. 2014 , 38 , 1156–1183. [ Google Scholar ] [ CrossRef ]
- Smith, A.; Raven, R. What is protective space? Reconsidering niches in transitions to sustainability. Res. Policy 2012 , 41 , 1025–1036. [ Google Scholar ] [ CrossRef ]
- Geels, F.W.; Schot, J. Typology of socio-technical transition pathways. Res. Policy 2007 , 36 , 399–417. [ Google Scholar ] [ CrossRef ]
- Geels, F.W. Regime resistance against low-carbon transitions: Introducing politics and power into the multi-level perspective. Theory Cult. Soc. 2014 , 31 , 21–40. [ Google Scholar ] [ CrossRef ]
- Markard, J.; Raven, R.; Truffer, B. Sustainability transitions: An emerging field of research and its prospects. Res. Policy 2012 , 41 , 955–967. [ Google Scholar ] [ CrossRef ]
- Geels, F.W. Reconceptualising the co-evolution of firms-in-industries and their environments: Developing an inter-disciplinary Triple Embeddedness Framework. Res. Policy 2014 , 43 , 261–277. [ Google Scholar ] [ CrossRef ]
- Klerkx, L.; Aarts, N.; Leeuwis, C. Adaptive management in agricultural innovation systems: The interactions between innovation networks and their environment. Agric. Syst. 2010 , 103 , 390–400. [ Google Scholar ] [ CrossRef ]
- Geels, F.W. The multi-level perspective on sustainability transitions: Responses to seven criticisms. Environ. Innov. Soc. Transit. 2011 , 1 , 24–40. [ Google Scholar ] [ CrossRef ]
- Viola, E.; Mendes, V. Agriculture 4.0 and climate change in Brazil. Ambiente Soc. 2022 , 25 , e02462. [ Google Scholar ] [ CrossRef ]
- Farhangi, M.H.; Turvani, M.E.; van der Valk, A.; Carsjens, G.J. High-tech urban agriculture in Amsterdam: An actor network analysis. Sustainability 2020 , 12 , 3955. [ Google Scholar ] [ CrossRef ]
- Swaffield, S.R.; Corry, R.C.; Opdam, P.; McWilliam, W.; Primdahl, J. Connecting business with the agricultural landscape: Business strategies for sustainable rural development. Bus. Strategy Environ. 2019 , 28 , 1357–1369. [ Google Scholar ] [ CrossRef ]
- van der Gaast, K.; van Leeuwen, E.; Wertheim-Heck, S. Food systems in transition: Conceptualizing sustainable food entrepreneurship. Int. J. Agric. Sustain. 2022 , 20 , 705–721. [ Google Scholar ] [ CrossRef ]
- Horn, E.K.; Joyce, A.; Chowdhury, R.B.; Caputo, S.; Jacobs, B.; Winkler, M.; Proksch, G. Translating Environmental Potential to Economic Reality: Assessment of Commercial Aquaponics through Sustainability Transitions Theory. Circ. Econ. Sustain. 2023 , 4 , 523–554. [ Google Scholar ] [ CrossRef ]
- Davidson, D.J.; Jones, K.E.; Parkins, J.R. Food safety risks, disruptive events and alternative beef production: A case study of agricultural transition in Alberta. Agric. Hum. Values 2016 , 33 , 359–371. [ Google Scholar ] [ CrossRef ]
- Kinniburgh, F. The politics of expertise in assessing alternatives to glyphosate in France. Environ. Sci. Policy 2023 , 145 , 60–72. [ Google Scholar ] [ CrossRef ]
- Manuel-Navarrete, D.; Gallopín, G.C. Feeding the world sustainably: Knowledge governance and sustainable agriculture in the Argentine Pampas. Environ. Dev. Sustain. 2012 , 14 , 321–333. [ Google Scholar ] [ CrossRef ]
- McInnes, A. Integrating sustainability transitions and food systems research to examine consultation failures in Canadian food policymaking. J. Environ. Policy Plan. 2019 , 21 , 407–426. [ Google Scholar ] [ CrossRef ]
- Marletto, G.; Sillig, C. Lost in mainstreaming? Agrifood and urban mobility grassroots innovations with multiple pathways and outcomes. Ecol. Econ. 2019 , 158 , 88–100. [ Google Scholar ] [ CrossRef ]
- Block, C.; Rennings, M.; Bröring, S. Selecting technologies to engage in sustainability transitions—A multi-stakeholder perspective. Bus. Strategy Environ. 2023 , 32 , 3569–3595. [ Google Scholar ] [ CrossRef ]
- Long, T.B.; Blok, V.; Coninx, I. The diffusion of climate-smart agricultural innovations: Systems level factors that inhibit sustainable entrepreneurial action. J. Clean. Prod. 2019 , 232 , 993–1004. [ Google Scholar ] [ CrossRef ]
- Trotter, P.A.; Becker, T.; Renaldi, R.; Wang, X.; Khosla, R.; Walther, G. The role of supply chains for the sustainability transformation of global food systems: A large-scale, systematic review of food cold chains. J. Ind. Ecol. 2023 , 27 , 1429–1446. [ Google Scholar ] [ CrossRef ]
- Keating, C.B.; Katina, P.F.; Pyne, J.C.; Jaradat, R.M. Systemic intervention for complex system governance development. In Proceedings of the American Society for Engineering Management International Annual Conference, Huntsville, AL, USA, 18–21 October 2017. [ Google Scholar ]
- Halbe, J.; Pahl-Wostl, C. A methodological framework to initiate and design transition governance processes. Sustainability 2019 , 11 , 844. [ Google Scholar ] [ CrossRef ]
- Rossing, W.A.H.; Albicette, M.M.; Aguerre, V.; Leoni, C.; Ruggia, A.; Dogliotti, S. Crafting actionable knowledge on ecological intensification: Lessons from co-innovation approaches in Uruguay and Europe. Agric. Syst. 2021 , 190 , 103103. [ Google Scholar ] [ CrossRef ]
- Ribeiro, B.; Turner, J.A. Sustainability buckets: A flexible heuristic for facilitating strategic investment on place-dependent sustainability narratives. Sustainability 2021 , 13 , 9367. [ Google Scholar ] [ CrossRef ]
- Audet, R.; Lefèvre, S.; Brisebois, É.; El-Jed, M. Structuring tensions and key relations of Montreal seasonal food markets in the sustainability transition of the agri-food sector. Sustainability 2017 , 9 , 320. [ Google Scholar ] [ CrossRef ]
- Stephens, R.; Barbier, M. Digital fooding, cashless marketplaces and reconnection in intermediated third places: Conceptualizing metropolitan food provision in the age of prosumption. J. Rural Stud. 2021 , 82 , 366–379. [ Google Scholar ] [ CrossRef ]
- Desa, G.; Jia, X. Sustainability transitions in the context of pandemic: An introduction to the focused issue on social innovation and systemic impact. Agric. Hum. Values 2020 , 37 , 1207–1215. [ Google Scholar ] [ CrossRef ]
- Donati, L.; Stefani, G.; Bellandi, M. The evolutionary emergence of quintuple helix coalitions: A case study of place-based sustainability transition. Triple Helix 2023 , 10 , 125–155. [ Google Scholar ] [ CrossRef ]
- Ferguson, P. Productivity growth as a barrier to a sustainability transition. Environ. Innov. Soc. Transit. 2016 , 20 , 86–88. [ Google Scholar ] [ CrossRef ]
- Gonera, A.; Nykamp, H.A.; Carraresi, L. Incumbents’ capabilities for sustainability-oriented innovation in the Norwegian food sector—An integrated framework. Circ. Econ. Sustain. 2023 , 3 , 1299–1326. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Pellizzoni, L.; Centemeri, L. Tackling material dependency in sustainability transition: Rationales and insights from the agriculture sector. J. Environ. Policy Plan. 2022 , 24 , 355–366. [ Google Scholar ] [ CrossRef ]
- Hák, T.; Janoušková, S.; Moldan, B. Sustainable Development Goals: A need for relevant indicators. Ecol. Indic. 2016 , 60 , 565–573. [ Google Scholar ] [ CrossRef ]
Click here to enlarge figure
| Author | Publications |
|---|---|
| Bui, S. [ , , ] | 3 |
| Hekkert, M.P. [ , , ] | 3 |
| De Schutter, O. [ , ] | 2 |
| Dedeurwaerdere, T. [ , ] | 2 |
| Feola, G. [ , ] | 2 |
| Giagnocavo, C. [ , ] | 2 |
| Hudon, M. [ , ] | 2 |
| Kuokkanen, A. [ , ] | 2 |
| Mehrabi, S. [ , ] | 2 |
| Moors, E. [ , ] | 2 |
| Rank | Affiliation | No. of Papers | Country |
|---|---|---|---|
| 1 | Wageningen University and Research | 6 | The Netherlands |
| 2 | Universiteit Utrecht | 5 | The Netherlands |
| 3 | Copernicus Institute of Sustainable Development | 5 | Germany |
| 4 | CNRS Centre National de la Recherche Scientifique | 3 | France |
| 5 | University of Guelph | 3 | Canada |
| 6 | Université Libre de Bruxelles | 2 | Belgium |
| 7 | Université Catholique de Louvain | 2 | Belgium |
| 8 | Lincoln University | 2 | New Zealand |
| 9 | LUT University | 2 | Finland |
| 10 | Universidad de Almería | 2 | Spain |
| 11 | Deakin University | 2 | USA |
| 12 | Groupe de Recherche en Droit, Économie et Gestion | 2 | France |
| 13 | Institut National de la Recherche Agronomique | 2 | France |
| Country/Territory | No. Papers |
|---|---|
| The Netherlands | 11 |
| Germany | 7 |
| Canada | 6 |
| France | 5 |
| Australia | 3 |
| Belgium | 3 |
| Finland | 3 |
| Italy | 3 |
| New Zealand | 3 |
| Sweden | 3 |
| Source | Publications | Impact Factor | CiteScore | SJR | SNIP |
|---|---|---|---|---|---|
| Environmental Innovation and Societal Transitions | 6 | 9.4 | 13.1 | 2.4 | 1.9 |
| Sustainability | 5 | 3.9 | 5.8 | 0.7 | 1.2 |
| Agriculture and Human Values | 3 | 4.9 | 5.9 | 1.0 | 1.4 |
| Agricultural Systems | 2 | 6.8 | 11.9 | 1.6 | 2.0 |
| Agriculture | 2 | 3.4 | 3.6 | 0.6 | 1.2 |
| Business Strategy and the Environment | 2 | 10.8 | 17.8 | 2.9 | 2.8 |
| Circular Economy and Sustainability | 2 | N/A | N/A | N/A | N/A |
| Ecological Economics | 2 | 6.5 | 11.0 | 1.9 | 2.0 |
| Journal of Cleaner Production | 2 | 11.1 | 18.5 | 2.0 | 2.4 |
| Journal of Rural Studies | 2 | 5.1 | 8.1 | 1.3 | 1.9 |
| Rank | Authors | Title | Year | Source Title | Cited by |
|---|---|---|---|---|---|
| 1 | Meynard et al. [ ] | Designing coupled innovations for the sustainability transition of agrifood systems. | 2017 | Agricultural Systems | 171 |
| 2 | Vinnari and Vinnari [ ] | A framework for sustainability transition: The case of plant-based diets. | 2014 | Journal of Agricultural and Environmental Ethics | 65 |
| 3 | Kuokkanen et al. [ ] | Agency in regime destabilization through the selection environment: The Finnish food system’s sustainability transition. | 2018 | Research Policy | 43 |
| 4 | El Bilali [ ] | Relation between innovation and sustainability in the agro-food system. | 2018 | Italian Journal of Food Science | 41 |
| 5 | Bui [ ] | Systemic ethics and inclusive governance: Two key prerequisites for sustainability transitions of agrifood systems. | 2019 | Agriculture and Human Values | 40 |
| 6 | Vermunt et al. [ ] | Sustainability transitions in the agrifood sector: How ecology affects transition dynamics. | 2020 | Environmental Innovation and Societal Transitions | 38 |
| 7 | Dedeurwaerdere et al. [ ] | The governance features of social enterprise and social network activities of collective food buying groups. | 2017 | Ecological Economics | 35 |
| 8 | Pant et al. [ ] | Adaptive transition management for transformations to agricultural sustainability in the Karnali mountains of Nepal. | 2014 | Agroecology and Sustainable Food Systems | 29 |
| 9 | van Oers et al. [ ] | The politics of deliberate destabilisation for sustainability transitions. | 2021 | Environmental Innovation and Societal Transitions | 27 |
| 10 | Kuokkanen et al., 2019 [ ] | A framework of disruptive sustainable innovation: An example of the Finnish food system. | 2019 | Technology Analysis and Strategic Management | 26 |
| Author(s) | Cited Reference | Source | Citations | Total Link Strength |
|---|---|---|---|---|
| Geels [ ] | Technological transitions as evolutionary reconfiguration processes: a multi-level perspective and a case-study. | Research Policy | 8 | 18 |
| Smith and Raven [ ] | What is protective space? Reconsidering niches in transitions to sustainability. | Research Policy | 6 | 16 |
| Geels and Schot [ ] | Typology of socio-technical transition pathways. | Research Policy | 5 | 15 |
| Geels [ ] | Regime resistance against low-carbon transitions: introducing politics and power into the multi-level perspective. | Theory Cult. Soc | 3 | 11 |
| Markard et al. [ ] | Sustainability transitions: an emerging field of research and its prospects. | Research Policy | 7 | 9 |
| Geels [ ] | Reconceptualising the co-evolution of firms-in-industries and their environments: developing an inter-disciplinary triple embeddedness framework. | Research Policy | 3 | 8 |
| Klerkx et al. [ ] | Adaptive management in agricultural innovation systems: the interactions between innovation networks and their environment. | Agricultural Systems | 3 | 4 |
| Geels [ ] | The multi-level perspective on sustainability transitions: responses to seven criticisms. | Environmental Innovation and Societal Transitions | 4 | 1 |
| The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
Share and Cite
Lees, N.J.; Sivakumar, S.; Lucock, X. Agrifood Sustainability Transitions in Firms and Industry: A Bibliographic Analysis of Research Themes. Sustainability 2024 , 16 , 7079. https://doi.org/10.3390/su16167079
Lees NJ, Sivakumar S, Lucock X. Agrifood Sustainability Transitions in Firms and Industry: A Bibliographic Analysis of Research Themes. Sustainability . 2024; 16(16):7079. https://doi.org/10.3390/su16167079
Lees, Nic J., Sivashankar Sivakumar, and Xiaomeng Lucock. 2024. "Agrifood Sustainability Transitions in Firms and Industry: A Bibliographic Analysis of Research Themes" Sustainability 16, no. 16: 7079. https://doi.org/10.3390/su16167079
Article Metrics
Article access statistics, further information, mdpi initiatives, follow mdpi.

Subscribe to receive issue release notifications and newsletters from MDPI journals

Generating knowledge graphs through text mining of catalysis research related literature †

First published on 5th August 2024
Structured research data management in catalysis is crucial, especially for large amounts of data, and should be guided by FAIR principles for easy access and compatibility of data. Ontologies help to organize knowledge in a structured and FAIR way. The increasing numbers of scientific publications call for automated methods to preselect and access the desired knowledge while minimizing the effort to search for relevant publications. While ontology learning can be used to create structured knowledge graphs, named entity recognition allows detection and categorization of important information in text. This work combines ontology learning and named entity recognition for automated extraction of key data from publications and organization of the implicit knowledge in a machine- and user-readable knowledge graph and data. CatalysisIE is a pre-trained model for such information extraction for catalysis research. This model is used and extended in this work based on a new data set, increasing the precision and recall of the model with regard to the data set. Validation of the presented workflow is presented on two datasets regarding catalysis research. Preformulated SPARQL-queries are provided to show the usability and applicability of the resulting knowledge graph for researchers.
Introduction
Information structuring and systematization can be achieved through the utilization of ontologies. An ontology serves as a data model, depicting a collection of concepts and the relationships among these concepts within a specific domain. 7 In this framework, terms are organized hierarchically as classes and subclasses, with each class linked to other classes through properties. This allows for automated classification of research papers with regard to those classes, increasing FAIRness of the classified texts.
Ontology learning (OL) from text is the (semi-)automated process of ontology creation or reuse for enrichment or population purposes. In recent years, several OL approaches have been developed to automate the construction of ontologies. Heuristic and conceptual clustering is one of the statistical-based approaches used for grouping the concepts based on the semantic distance between them to build towards hierarchies. This method was employed in previous work 8 for knowledge extraction from catalysis-related texts for the automatic creation of a taxonomy with important terms extracted from texts. However, the resulting hierarchy is still missing specific interrelations between the terms, and concepts lack proper characterization through axioms. This proves that it is important to integrate the relation extraction in the process of OL. One other tool OntoCmaps 9 is an OL system, with which non-taxonomic relations can be recognized with dependency structure analysis and ontologies are constructed in the form of concept maps, which are not domain-specific and can contain not necessary information.
To extract valuable information from publications in the field of catalysis research, which can be considered as a named entity recognition (NER) task, a pre-trained model, CatalysisIE, 10 was used. This allows for identification of key information from a text based on pre-trained classes, such as names of people. The authors of this model constructed the first benchmark data set for knowledge extraction from the scientific literature in catalysis using active learning to generate a candidate sentence pool for annotation purposes. With this, extracted entities can be categorized into six categories: catalyst, reaction, reactant, product, characterization, and treatment. For the text span representation, pre-trained SciBERT 11 models were used. The parameters of SciBERT were optimized for catalysis-related information extraction (IE) by undergoing the domain adaptation using a corpus consisting of 10.4 million words. 10
The objective of this work is to facilitate the acquisition of information for catalysis research. This is obtained through the design of a tool for the automatic systematization of data extracted from scholarly publications into knowledge graphs. The construction of the knowledge graphs is based on an ontology, which allows for higher data FAIRness by structured relations and conceptual classification of knowledge. Additionally, the content of these publications is preserved in the form of terms deemed relevant to catalysis research. Utilizing CatalysisIE, the extracted entities can be categorized into the six concepts. After preprocessing, the abstracts from scholarly publications can be extracted with natural language processing (NLP) techniques. NLP techniques enable computers to interpret and generate human language, such as scientific texts. For IE, the pretrained model by the authors of CatalysisIE 10 can be used. Furthermore, the CatalysisIE model is trained on the complemented dataset presented in this work.
Ontologies can be queried using SPARQL 12 queries, e.g. formulated in Python functions. SPARQL is a structured query language used to retrieve data stored within databases, especially for triplet-based data, such as ontologies. Automatically generated knowledge graphs containing information for the retrieval of the publications can also later be queried for retrieval of publications.
| Overview of the proposed framework. | ||
Data retrieval
In this work, abstracts were processed assuming that this part of the initial text contains important information about the content of the article and the output is less affected by noisy and repetitive information. The noisy information could be, for example, the previous studies usually mentioned in the introduction section. Abstracts and publication titles were retrieved using text extraction techniques directly from PDF-files. Furthermore, the publicly available CrossRef REST API and publishers' API for metadata retrieval were used. 14 The CrossRef REST API was integrated with the habanero Python package, 15 which fulfils the role of a low-level client that provides functionalities for querying and response handling. The pybliometrics package, 16 an API-wrapper to access Scopus, is used for abstract scraping, which implies the process of automatically mining data or collecting information from the internet, along with pdfdataextractor 17 for abstract extraction directly from PDF files. When an abstract is not able to be retrieved from a PDF, the HTML-version of the publication and thus its abstract are retrieved based on the DOI.
After text preprocessing, entities relevant to the field of catalysis research are extracted from the text using a pre-trained model and labelled with one of the six categories: catalyst, reaction, reactant, product, characterization, and treatment.
Text mining and preprocessing
An example of compound identification with user feedback for “Rh 2 O 3 ” is shown in Fig. 2(a) . In cases where no compounds could be retrieved from PubChem and no synonyms were found in ChEBI, the user is prompted to confirm whether the compound exists. If the user answers “no”, the presumed chemical entity is skipped. By responding “yes”, the user confirms the existence of the entity and it will be identified with the name provided to the user (see Fig. 2(b) ).
| Example of a user-feedback process during preprocessing in Python. Identification of chemical compounds using queried compounds from PubChem (a) and confirmation of the compound existence (b). User input is marked in green. | ||
Ontology learning
As the main working ontology, the Allotrope Foundation Ontology (AFO) 22 is selected. In this ontology, object properties important for the ontology extension process are created manually ( Table 4 in the appendix), while classes and data properties are created automatically. After preprocessing of the extracted entities based on their labelled roles, the working ontology is extended with classes and instances using existing terms in other ontologies. Furthermore, rule-based approaches in combination with syntactic dependency parsing are pursued. The initial preprocessing of the extracted entities includes the POS taggers, lemmatization, tokenization, and the use of regular expressions. In specific steps, such as lemmatization, modules from spaCy 23 are used. Thus, the preprocessing allows prevention of the creation of synonyms as independent concepts in the ontology and creation of relations based on the context in the text. For example, given an entity “RhCo/SiO 2 ” labelled “catalyst”, “RhCo” is recognised as a catalyst, which is supported on “SiO 2 ”. This relation will be added later in the ontology to the corresponding classes of the compounds. To recreate the hierarchical structure within the ontology, syntactic dependency parsing was utilized as a preprocessing step to define the “head” – word of the entity and use it for search in ontologies. For example, the “head” of “packed bed methanation” is “methanation”.
In this work, some existing ontologies were used in the workflow for ontology extension. The primary selection criterion for these ontologies was their relevance to the domain of catalysis research. The selection of ontologies for this work, which are listed along with a short description in Table 1 , is based on the overview from the collection of ontologies for catalysis research presented in ref. 24 .
| Ontology acronym | Full name | Description |
|---|---|---|
| BFO | Basic Formal Ontology | A small ontology is designed for use in analysis and integration in scientific or other domains. It does not contain physical, chemical, or other domain-specific terms and is well used as a top-level ontology |
| AFO | Allotrope Foundation Ontology | AFO is a domain-specific ontology, offering a standardized vocabulary and semantic framework for the representation of laboratory analytical processes. AFO is aligned with BFO as a top-level ontology. Reasoning can be only provided by the HermiT reasoner |
| ChEBI | Chemical Entities of Biological Interest | ChEBI represents a vocabulary with a focus on small molecular entities and contains such information as InChiKeys, CAS numbers, and exact and related synonyms of chemical compounds |
| MOP | Molecular Process Ontology | Domain ontology contains good conceptual descriptions of molecular processes, such as crystallization and methylation |
| RXNO | Name Reaction Ontology | Domain ontology is strongly connected to MOP. It contains more than 500 classes representing organic reactions with good conceptual description |
Ontology extension is based on the reuse of the classes existing in other ontologies to properly characterize concepts and reuse existing frameworks and axioms. The reuse of ontology terms creates links between data, making the ontology more valuable. 28
For example, ChEBI contains more than 3 million axioms. Thus, only relevant subsets of ontologies were reused. After searching for classes within the working ontology using the owlready2 Python package, 29 missing chemical compounds and possible reaction types are searched for in other ontologies as listed in Table 1 . To accomplish this, a nested dictionary is used, containing all IRIs of terms alongside their corresponding labels, prefLabels, altLabels, and the names used within the ontology. Applying a class information extraction process that incorporates functions from the owlready2 package, the dictionary was generated for 22 ontologies relevant to the domain of catalysis research. 24 Once the dictionary is loaded into the Python environment, it is searched for classes missing in the working ontology. If one of the labels, prefLabels, altLabels, or names matches the searched entity, the corresponding IRI is added to the dictionary along with the matching value. The IRIs of found terms that are still missing in the working ontology are stored in automatically created text files. The names of the text files include the acronym of the source ontology for further reference.
Ontology extension
This ensures that the module retains all the same logical entailments in the full ontology, providing consistency in the ontology subset. The chosen SMLE approach is the BOTTOM module, which contains the terms in the seed, their corresponding superclasses, and the interrelations between them. As the name implies, the class hierarchy is built from the bottom up, gathering the superclasses of the selected class. Thus, for each ontology, a separate subset of relevant classes is created in rdf/xml (owl) format.
The second task for which ROBOT is used in this work is to merge the created subsets of classes and the main ontology into a single ontology with a single .owl-file. Thus, the merging process is used to update the working ontology within existing terms of other ontologies.
Because some of the merged ontologies are aligned with different top-level ontologies, terms that theoretically share the same definition are located at different positions within the class hierarchy. For example, the OBO is the top-level ontology of ChEBI, while the AFO is aligned with the BFO. Both ontologies have, for example, the term “atom”, but at different positions in the class hierarchy. Another factor why the same terms are represented differently is related to the granularity problem of ontologies. This issue arises because ontologies often adopt different levels of details when representing identical knowledge to support different applications. 30
Since all of the utilized ontologies are connected to the domain of catalysis 24 and chemistry, 31 terms with identical designation are assumed equivalent. The equivalence of classes indicates that respective classes share all their instances, and the descriptions of both classes are interlinked. However, the use of the equivalence relation does not imply class equality. Both relations are defined differently in OWL. Equality is denoted by “owl:sameAs”, while the equivalence is represented by owl:equivalentClass. Class equality can only be defined by the description language OWL-Full, and owlready2 supports only equivalence. 29
To identify terms with the same designation that originate from different ontologies and consequently have different IRIs, the mappings created in previous work 24 are used in the processing. These mappings represent all terms shared by two ontologies according to the same IRIs or the same set of labels, prefLabels, names, or altLabels. After merging ontologies to reuse existing terms, the process of creating new classes and subsequently populating the working ontology with new instances is initialized. First, a new instance of a publication is generated as an instance of the “publication” superclass. The DOI and title of the publication retrieved at the beginning of the process are added to the publication instance as datatypes using the “has doi” and “has title” datatype properties, respectively. Extracted chemical compounds that do not exist in the working ontology after merging are then created as new classes within the working ontology, utilizing the context information of the new compounds. Chemical compounds that can be further broken down into compounds and atoms, such as “Al 2 O 3 ” or “titanium dioxide”, or those that are recognized as compounds using pubchempy are created as subclasses of the “molecule” class.
Support material entities, which represent a combination of two or more carrier compounds, like “TiO 2 –SiO 2 ”, or materials such as “MCM-41” are created as subclasses of the “support material” class. Each newly created class and instance are automatically assigned a generated name linked to the number of the processed publications in the working ontology.
Entities from the “Reactant”, “Product” and “Catalyst” categories that represent specific types of chemical entities, such as “light olefin” and “vapour phase propene”, are created as instances of the corresponding chemical compound. Extracted and preprocessed catalyst entities are created as instances of the “chemical substance” class.
Chemical entities, which represent catalysts in the form of
“ <catalytic compound>/<support compound> ” or
“ <catalytic compound>@<support compound> ”
are labelled in the ontology
“ <catalytic compound> supported on <support compound> ”
and linked with their chemical compounds based on their roles in the entity using the “catalytic component of” and “support component of” object properties. A schematic example of interconnections within the ontology is shown in Fig. 3 .
| Examples of created entities and assigned relations. Entities within dashed boxes represent instances, while continuous bordered boxes represent classes. | ||
Table 4 in the appendix lists the object properties and their inverse properties that need to be defined within the working ontology in order to assign the relations between the newly created entities.
The creation of the classes corresponding to the catalyst types is based on the creation of subclasses of the term “catalyst role”, which already exists in the AFO ontology. Roles in ontologies are used to reduce the amount of object properties and thus to speed up reasoning. The corresponding roles of terms are provided as classes in the ontology and terms are linked to them via the “has role” object property. The hierarchical structuring of the catalyst roles is based on the content of the classes extracted from the entities. For instance, within the text corpus, the extracted class of an entity after preprocessing might be a “dispersed catalyst role”, while the catalyst type corresponding to another entity is an “atomically dispersed catalyst role”. Since the second class is identical to the first but with an additional word, it is considered a subclass of the first class. In case no entity from the text corpus has a “dispersed catalyst role” as an extracted class, then an “atomically dispersed catalyst role” is created as a subclass of the “catalyst role”.
Chemical reactions are created as subclasses of the previously extracted reaction “heads”. If there is no corresponding reaction found in other ontologies, a new class is created as a subclass of “chemical reaction (molecular)”, which is also a class within the AFO. For each class created after the merging that corresponds to an extracted entity or a chemical compound, an instance is created with an automatically generated name.
The label of the instance is the same as the label of its corresponding class. The same procedure is applied to the newly created classes. All classes and instances, once created, can be reused for ontology extension with the respective publications. The newly created classes of chemical compounds are linked to their corresponding components via “has part” relations at the instance level.
Created instances are linked to their roles according to the categories and the context using the “has role” relation. The used roles include the “support role”, “reactant role”, “product role”, and “catalyst role”, and all are created as subclasses of the “catalyst role”.
Finally, all created and used instances that are mentioned in the processed publication are linked to the instance of the publication through the “mentioned in” object property. Entities labelled “Characterization” and “Treatment” are added to annotations of publications as comment.
SPARQL queries
The following competency questions were implemented as SPARQL queries and can thus be easily retrieved from the knowledge graph resulting from the extension of the ontology. The corresponding SPARQL queries are numbered and exemplary input and output of the queries are listed in Table 5 in the appendix and an exemplary SPARQL query is listed in Table 6 in the appendix:
• Give me a list of reactions (1), reactants, support materials, catalysts, and products mentioned in one specific publication, which is a part of the knowledge graph, in one list (2) or separately,
• Retrieve the abstracts from publications in the ontology (3),
• Give me a list of DOIs of publications from the working ontology, which mention the same reactions (4) or the specific reaction (5) or catalyst (6),
• Give me a list of reactions, reactants, support materials, catalysts, and products mentioned in all publications of the knowledge graph (7),
• I need a list of all possible synonyms for the extracted reactants (8), support materials (9), catalysts (10), and products (11) in the form of chemical entities,
• I need possible catalysts where the support material from this paper can be used (12).
The retrieved entities can be used to query Scopus for new publications with similar context. Using the pybliometrics Python package, the search is performed, leading to a query, which has the same structure as a query that works in the Scopus advanced search. With the chosen query type ‘TITLE-ABS-KEY()’ (as depicted exemplarily in Fig. 4 ), the search is performed within the titles, abstracts, and keywords of the publication.
| Two types of query formulations for the advanced search for further publications in Scopus, executed by the Python API. | ||
Since there are multiple ways to name a specific chemical compound, to avoid a large number of possible queries and at the same time allow diversity in the naming of chemical compounds, a trade name or common name and a formula listed in the class annotations of a chemical compound are used for queries' formulation. Moreover to exclude mismatches, the publication will be skipped if during text mining no reaction was found in the text. After a query is executed, its results are downloaded and cached to speed up the subsequent analysis.
After the results are concatenated into one table, duplicates are removed from it. As Scopus contains records of articles published since 1970, an option to filter the results by publication date is integrated into the process, to allow for the inclusion of primarily newer publications into the knowledge graph. Utilization of the pandas Python module 33 allows the resulting DataFrames to be stored as sheets in an Excel file.
Results and discussion
Moreover, a set of 28 publications on methanation processes (dataset 2) was used to evaluate how well the created tool works on the different types of catalysed reactions. Hereby the focus was laid on the heterogeneously catalysed conversion of carbon monoxide and carbon dioxide to methane via hydrogenation, which is important for the production of synthetic natural gas. In particular, the valorization of CO 2 together with renewable hydrogen might be considered an integral sustainable path towards the production of renewable gaseous fuels. 36 For that, an extension of an alternate ontology setup similar to the first dataset was performed.
The dataset for training of the model was complemented with 151 sentences manually labelled in label-studio 37 from 18 abstracts of papers to the topic of hydroformylation in the liquid and gas phase. Checkpoints from the model trained by the authors of CatalysisIE and the model trained on the complemented dataset were compared with each other.
To evaluate the difference in the prediction of the checkpoints, ten manually labelled abstracts from papers to the same topic were compared to predictions of both models. Since it is important to gain as many correct distinct predictions from the text as possible to be able to describe the content of the publication using extracted entities, the recall R of the model was evaluated with the number of true positives TP and false negatives FN using eqn (1) . To obtain the true positives and false negatives, the amount of distinct entities was counted and compared with the number of distinct entities from the prediction after qualitative manual labelling of the texts. This comparison for each extracted abstract from dataset 1 is shown in Table 9 in the appendix.
Besides recall, the precision Pr was selected for evaluation of multi-label classification. Because class imbalances are present in the dataset, the precision was calculated using eqn (2) with the number of positives P i instead of true positives TP and the number of used labels N . Furthermore, the standard deviation σ of the precision was selected as a metric and calculated using eqn (3) . The sum of true positives corresponds to the number of correctly predicted instances. Precision and its standard deviation were calculated for the six categories for each of the abstracts.
| (1) |
| (2) |
| (3) |
Extraction of sequences was treated as a binary classification problem, where the sum of TP is correctly extracted from distinct entities and is independent from the assigned label. The sum of true positives and false negatives is the total number of distinct manually labelled entities in the text. The metrics were calculated for the ten manually labelled abstracts from papers to the same topic and compared to predictions of both checkpoints I and II. Here, checkpoint I addresses the complemented model, while checkpoint II addresses the pre-trained checkpoint, provided by the developers of CatalysisIE. The resulting metrics are listed in Table 2 . Deviations in the metrics of the fourth publication may be due to formatting errors in the retrieved abstract, causing extracted tokens to end with citation numbers ( e.g. , “catalysts9”), thus not being counted as found entities.
| TP + FN | CP | TP | R | Pr | σ | |
|---|---|---|---|---|---|---|
| 1 | 12 | I | 11 | 91.7 | 83.3 | 40.8 |
| II | 10 | 83.3 | 66.7 | 51.6 | ||
| 2 | 15 | I | 14 | 93.3 | 93.3 | 14.9 |
| II | 12 | 80.0 | 80.0 | 44.7 | ||
| 3 | 15 | I | 13 | 86.7 | 75.6 | 43.3 |
| II | 12 | 70.0 | 73.3 | 43.5 | ||
| 4 | 16 | I | 6 | 37.5 | 35.0 | 23.8 |
| II | 6 | 37.5 | 38.7 | 30.7 | ||
| 5 | 15 | I | 10 | 66.7 | 64.6 | 9.9 |
| II | 7 | 66.0 | 61.8 | 26.7 | ||
| 6 | 15 | I | 13 | 86.7 | 58.3 | 49.2 |
| II | 14 | 93.3 | 66.7 | 51.6 | ||
| 7 | 37 | I | 33 | 89.2 | 88.2 | 21.7 |
| II | 24 | 64.9 | 62.7 | 16.3 | ||
| 8 | 29 | I | 25 | 86.2 | 68.2 | 10.5 |
| II | 24 | 82.8 | 64.7 | 11.4 | ||
| 9 | 25 | I | 12 | 48.0 | 63.6 | 26.0 |
| II | 13 | 52.0 | 68.6 | 25.1 | ||
| 10 | 6 | I | 6 | 100.0 | 100.0 | 0.0 |
| II | 6 | 100.0 | 100.0 | 0.0 |
The entities labelled “Characterization” were predicted least accurately. Additionally, there were no “Treatment” labels in the evaluation dataset. Overall, the model trained on the expanded dataset (CP I) was better at predicting entities labelled “Catalyst”. The average recall of the newly trained model for ten abstracts is equal to 86.67% with a standard deviation of 20.85% and shows a high average precision of 71.90%. In comparison, the recall of the old model (CP II) is 80.00% with a standard deviation of 19.37% and an average precision of 66.67%. In both cases only in one text, precision and recall fall under 50%. Furthermore, for the ten publications shown in Table 2 , in the cases where CP I achieved higher precision, σ was lower. This indicates that the dispersion across the different classes in relation to Pr has decreased and therefore the model makes more stable predictions across the classes.
To investigate the performance of the extended model further, ten abstracts from dataset 2 are labelled manually and classified with CP I and CP II to evaluate the metrics as in Table 2 . The resulting metrics are presented in more detail in Table 7 . An average recall of 82.81% with a standard deviation of 22.39% and an average precision of 71.46% was achieved for CP I. Furthermore, an average recall of 79.47% with a standard deviation of 19.92% and an average precision of 73.20% was achieved for CP II. Thus, the extended model can also be applied on dataset 2.
Title recognition by 19 out of 23 processed PDFs from dataset 1 was successful and 26 from 28 publications from dataset 2 could be recognized correctly. Publications of “Royal Society of Chemistry” could not be correctly recognized because the layout of the publications is not integrated in the workflow of the used pdfdataextractor package.
The AFO was chosen as the initial ontology, because of its linkage to the chemical domain and well-defined structure in the class hierarchy. Table 8 lists the terms and textual definitions assigned as equivalent in ontologies for both datasets, which exist in the AFO and are merged into the working ontology from ChEBI.
Chemicals which could not be found in PubChem or in ChEBI are created as instances of the class “chemical substance”. For dataset 1, the ontology is extended with 53 instances of “chemical substance”. Dataset 2 results in 55 instances of “chemical substance” that were also created automatically. Each of the generated instances representing extracted entities and their chemical components is provided with a connection to the publication in which it is mentioned and linked to the corresponding roles as shown in an excerpt of the resulting ontology in Fig. 5 . The reactions that are mentioned within the publication are listed, including the respective participants of the reactions within the knowledge base (upper area of the figure). The individual “cobalt atom”, for example, is connected with the individual “Co-containing catalyst” via the object property “catalytic component of” (right area of the figure), thus indicating the suitable catalytic component of the concept extracted from text. Furthermore, the role of a “bimetallic catalyst role” is asserted to the three individuals on the bottom right of Fig. 5 . The class “bimetallic catalyst role” is created as a subclass of the “catalyst role”, which in turn also has an individual that is connected to other substances via the “has role” object property (bottom left of the figure).
| Excerpt from the created ontology for dataset 1 created with Protégé. Boxes marked with yellow circles represent classes and those with purple rhombi are instances. Arrows denote the relationships between them, color-coded as listed in the legend on the right. Small boxes with a plus (+) inside indicate that not all relations of the entity are shown in the figure. | ||
The knowledge graph with publications from dataset 1 was extended by 48 classes from the other ontologies, including their superclasses and interrelations. In total, 331 new classes, 9 new object properties, 2 new data properties, and 155 new individuals were added to the working ontology. From the new classes, 288 were merged from other ontologies, while none of the new individuals were merged from other ontologies, as expected. The new object and data properties were merged from other ontologies.
In the knowledge graph with dataset 2, 39 classes from other ontologies were imported from other ontologies together with their respective superclasses and interrelations. With this, 222 new classes, 4 new object properties, 2 new data properties, and 130 new individuals were added to the working ontology. Here, 198 from the 222 new classes were merged from other ontologies, while also none of the new individuals were merged from other ontologies. The new object and data properties were merged from the other ontologies listed in Table 1 and counted without the ones already presented in Table 4 in the appendix. The explained ontology metrics are listed in Table 3 .
| Metric | Initial ontology | Extended ontology dataset 1 | Extended ontology dataset 2 |
|---|---|---|---|
| Classes | 3116 | 3447 | 3338 |
| Instances | 47 | 203 | 178 |
| Logical axioms | 5755 | 6936 | 6596 |
| SubClassOf | 4823 | 5372 | 5174 |
| Equivalent classes | 178 | 188 | 185 |
Fig. 6 shows the individual “0.5% Co–0.5% Rh supported on Al 2 O 3 ” in an excerpt of Protégé after reasoning with HermiT. 38 The implicit knowledge is highlighted in yellow, showing an increased semantic expressiveness for the individual describing the catalyst complex. Thus, the individual now also can be found when searching the knowledge graph, e.g. , for catalysts that contain cobalt.
| Excerpt from Protégé with inferred relations after reasoning for the individual “0.5% Co–0.5% Rh supported on Al O ”. Knowledge inferred by the reasoner is highlighted in yellow, showing increased semantic expressiveness of the individual. | ||
Most of the terms in both knowledge graphs originate from the ChEBI ontology and identify chemical compounds and atoms. But also, the classes for such terms as “hydrogenation”, “hydroformylation”, and “acylation” are reused from the RXNO ontology. In the current process, entities representing some chemical groups, such as “phenolic substances” or “phenolic species”, can be recognized with the text mining module, but the extension of the ontology with them is not implemented. This includes, for example, entities such as “phenolic substances”, “phenolic species”, and “alkyl species” which are usually classified as products or reactants in the text. Such entities cannot be queried in PubChem, and in ChEBI, the presumed superclasses are placed in different positions. All queries are formulated within the functions in the module “queries” provided in the GitHub repository of this work 39 and can be executed by the Jupiter notebook “user_queries.ipynb” contained in the repository. It contains descriptions of code cells, which execute specific queries, which can answer competency questions formulated in Methods. In addition, some examples of executed functions are also provided in the notebook.
Listings of the resulting publications are found in the provided GitHub repository of this work 39 in the “output” directory.
To further rate the quality of the query result, a random sample in size of 50 publications from the resulting filtered list of 731 publications similar to those of dataset 1 was selected for the evaluation of the queried content. The list of chosen publications for evaluation is provided in the appendix in Table 9 .
These publications are rated as similar to the publications in the knowledge graph (of dataset 1) if the following requirements are fulfilled, based on the evaluation of their abstracts and titles:
• Heterogeneous or homogeneous catalysis or catalysts are mentioned.
• Hydroformylation or hydrogenation is mentioned.
• Rh-, Co-, and Ni-based catalysts with silica, zeolite or aluminium oxide as a support material are mentioned.
According to these restrictions, 34 out of 50 publications of the sample provided are rated as similar to the content of the publications within the knowledge graphs, which is equal to 68% accuracy.
Conclusions
The quality of matched articles to the content of the knowledge graphs was evaluated with dataset 1, dealing with hydroformylation reactions. A random sample of 50 publications was investigated in more detail and abstracts, keywords, and titles were screened for the mention of catalysts, hydrogenation or hydroformylation, and whether Rh-, Co-, and Ni-based catalysts with silica, zeolite or aluminium oxide as a support material were mentioned. With these quite strict criteria and with the lack of case sensitivity in the Scopus API, 68% accuracy was achieved for the random sample. This allows for more structured searches of relevant scientific literature in the domain of catalysis research, which is highly important, especially in this domain, as research is quite heterogeneous and the number of relevant publications in the field is quite high. However, a more thorough post-processing of these found publications needs to be conducted, e.g. by a post-processing that conducts a case-sensitive automated search of the respective keywords in the extracted abstracts from Scopus, to improve the accuracy of the output related publications.
| Object property | Reverse object property | ||
|---|---|---|---|
| Name | Rdfs:Label | Name | Rdfs:Label |
| supported_on | Supported on | support_material_of | Support material of |
| support_component_of | Support component of | has_support_component | Has support component |
| catalytic_component_of | Catalytic component of | has_catalytic_component | Has catalytic component |
| mentioned_in | Mentioned in | Mentions | Mentions |
| Query no. | Input parameters | Output |
|---|---|---|
| 1 | Doi = r‘10.1021/acsami.0c21749.s001’ | [‘hydroformylation’] |
| 2 | list_type = ‘all’ | [‘hydroformylation’, ‘olefin’, ‘rhodium atom’, ‘Rh-based atomically dispersed catalyst’, ‘Rh supported on ZnO modified with Pi’, ‘zinc oxide’, ‘phosphate ion’, ‘aldehyde’, ‘linear aldehyde’] |
| Doi = r‘10.1021/acsami.0c21749.s001’ | ||
| 3 | Doi = r‘10.1021/acsami.0c21749.s001’ | Abstract: in the study of heterogeneity of homogeneous processes, effective control of the microenvironment of active sites… |
| 4 | Doi = r‘10.1021/acsami.0c21749.s001’ | [[‘10.1021/acscatal.1c02014.s001’], [‘10.1021/acscatal.0c04684.s001’], [‘10.1021/acscatal.1c00705.s002’], [‘10.1021/acscatal.1c04359’],…] |
| 5 | Reac = “hydrogenation”, Doi = none | [[‘10.1021/acscatal.0c04684.s001’], [‘10.1021/acscatal.1c00705.s002’],…] |
| 6 | Cat = “RhCo”, Doi = none | [[‘10.1021/acscatal.0c04684.s001’], [‘10.1021/acscatal.1c00705.s002’], |
| 7 | list_type = ‘all’ | [‘styrene’, ‘cobalt atom’, ‘0.5% Co–0.5% Rh supported on Al O ’, ‘Co-containing catalyst’, ‘aluminum oxide’, ‘hydroformylation’,…] |
| Doi = none | ||
| 8 | list_type = ‘reactant’ | [‘olefin’] |
| Doi = r‘10.1021/acsami.0c21749.s001’ | ||
| 9 | list_type = ‘product’ | [‘Aldehyd’, ‘RCHO’, ‘aldehidos’, ‘aldehydes’, ‘Aldehyde’, ‘aldehydum’, ‘an aldehyde’, ‘RC( |
| Doi = r‘10.1021/acsami.0c21749.s001’ | ||
| 10 | Doi = r‘10.1021/acsami.0c21749.s001’ | [[‘45Rh’, ‘rhodium’, ‘Rh’, ‘rodio’, ‘Rh(111)’, ‘rhodium atom’]], [[‘Rh on ZnO modified with Pi’, ‘Rh supported on ZnO modified with Pi’]] |
| 11 | Doi = r‘10.1021/acsami.0c21749.s001’ | [[‘zinc oxide’, ‘oxyde de zinc’, ‘Zinkoxid’, ‘oxido de cinc’, ‘ZnO’], [‘phosphate ions’, ‘Pi’, ‘phosphate’, ‘phosphate ion’]] |
| 12 | Doi = r‘10.1021/acscatal.1c02014.s001’ | [[‘silicon dioxide’, ‘Rh supported on SiO catalyst’], [‘silicon dioxide’, ‘Rh P nanoparticle supported on SiO support material’], [‘silicon dioxide’, ‘Rh7Co1P4 supported on SiO ’],…] |
| only_doi = false |
| TP + FN | CP | TP | R | Pr | σ | |
|---|---|---|---|---|---|---|
| 1 | 15 | I | 13 | 86.7 | 68.3 | 32.5 |
| II | 12 | 80.0 | 61.7 | 36.1 | ||
| 2 | 19 | I | 15 | 78.9 | 82.7 | 28.9 |
| II | 15 | 78.9 | 82.7 | 28.9 | ||
| 3 | 9 | I | 9 | 100.0 | 93.3 | 14.9 |
| II | 9 | 100.0 | 93.3 | 14.9 | ||
| 4 | 11 | I | 6 | 54.5 | 66.7 | 28.9 |
| II | 7 | 63.6 | 70.8 | 26.0 | ||
| 5 | 6 | I | 6 | 100.0 | 100.0 | 0.0 |
| II | 6 | 100.0 | 100.0 | 0.0 | ||
| 6 | 14 | I | 8 | 57.1 | 43.8 | 51.5 |
| II | 8 | 57.1 | 43.8 | 51.5 | ||
| 7 | 12 | I | 12 | 100.0 | 73.6 | 29.7 |
| II | 11 | 91.7 | 72.6 | 42.6 | ||
| 8 | 12 | I | 7 | 58.3 | 69.3 | 41.3 |
| II | 6 | 50.0 | 49.3 | 46.6 | ||
| 9 | 9 | I | 4 | 44.4 | 69.0 | 27.0 |
| II | 5 | 55.6 | 73.8 | 25.1 | ||
| 10 | 15 | I | 15 | 100.0 | 92.6 | 13.5 |
| II | 15 | 100.0 | 92.6 | 13.5 |
| Term label | IRIs + Definitions | |
|---|---|---|
| In the AFO | In ChEBI | |
| ‘Chemical substance’ | http://purl.allotrope.org/ontologies/material#AFM_0001097 | http://purl.obolibrary.org/obo/CHEBI_33250 |
| A chemical substance is a portion of material that is matter of constant composition best characterized by the entities (molecules, formula units, atoms) it is composed of [IUPAC] | A chemical entity constituting the smallest component of an element having the chemical properties of the element | |
| ‘Anion’ | http://purl.allotrope.org/ontologies/material#AFM_0000161 | http://purl.obolibrary.org/obo/CHEBI_22563 |
| An anion (−) is an ion with more electrons than protons, giving it a net negative charge (since electrons are negatively charged and protons are positively charged) | A monoatomic or polyatomic species having one or more elementary charges of the electron | |
| ‘Ion’ | http://purl.allotrope.org/ontologies/material#AFM_0000077 | http://purl.obolibrary.org/obo/CHEBI_24870 |
| An ion is an atom or molecule in which the total number of electrons is not equal to the total number of protons, giving the atom or molecule a net positive or negative electrical charge | A molecular entity having a net electric charge | |
| ‘Role’ | http://purl.obolibrary.org/obo/BFO_0000023 | http://purl.obolibrary.org/obo/CHEBI_50906 |
| B is a role means: b is a realizable entity and b exists because there is some single bearer that is in some special physical, social, or institutional set of circumstances in which this bearer does not have to be and b is not such that, if it ceases to exist, then the physical make-up of the bearer is thereby changed [BFO] | A role is particular behavior which a material entity may exhibit | |
| ‘Cation’ | http://purl.allotrope.org/ontologies/material#AFM_0000189 | http://purl.obolibrary.org/obo/CHEBI_36916 |
| A cation (+) is an ion with fewer electrons than protons, giving it a positive charge | A monoatomic or polyatomic species having one or more elementary charges of the proton | |
| ‘Group’ | http://purl.obolibrary.org/obo/BFO_0000023 | http://purl.obolibrary.org/obo/CHEBI_24433 |
| A group is an aggregate of people | A defined linked collection of atoms or a single atom within a molecular entity | |
| ‘Atom’ | http://purl.allotrope.org/ontologies/material#AFM_0001028 | http://purl.obolibrary.org/obo/CHEBI_33250 |
| An atom is a smallest particle still characterizing a chemical element. It consists of a nucleus of a positive charge carrying almost all its mass (more than 99.9%) and Z electrons determining its size | A chemical entity constituting the smallest component of an element having the chemical properties of the element | |
| ‘Chemical substance’ | http://purl.allotrope.org/ontologies/material#AFM_0001097 | http://purl.obolibrary.org/obo/CHEBI_59999 |
| A chemical substance is a portion of material that is matter of constant composition best characterized by the entities (molecules, formula units, atoms) it is composed of | A chemical substance is a portion of matter of constant composition, composed of molecular entities of the same type or of different types | |
Data availability
Author contributions, conflicts of interest, acknowledgements.
- D. W. Hook, S. J. Porter and C. Herzog, Dimensions: Building Context for Search and Evaluation, Front. Res. Metr. Anal. , 2018, 3 DOI: 10.3389/frma.2018.00023 .
- M. D. Wilkinson, et al. , The FAIR Guiding Principles for scientific data management and stewardship, Sci. Data , 2016, 3 , 160018, DOI: 10.1038/sdata.2016.18 .
- A. Salazar, B. Wentzel, S. Schimmler, R. Gläser, S. Hanf and S. A. Schunk, How Research Data Management Plans Can Help in Harmonizing Open Science and Approaches in the Digital Economy, Chemistry , 2023, 29 (9), e202202720, DOI: 10.1002/chem.202202720 .
- B. V. Elsevier, Scopus, 2024. Accessed: February 2024. [Online]. Available: https://www.scopus.com/.
- C. P. Marshall, J. Schumann and A. Trunschke, Achieving Digital Catalysis: Strategies for Data Acquisition, Storage and Use, Angew. Chem., Int. Ed. , 2023, 62 (30), e202302971, DOI: 10.1002/anie.202302971 .
- M. Suvarna, A. C. Vaucher, S. Mitchell, T. Laino and J. Pérez-Ramírez, Language models and protocol standardization guidelines for accelerating synthesis planning in heterogeneous catalysis, Nat. Commun. , 2023, 14 (1), 7964, DOI: 10.1038/s41467-023-43836-5 .
- S. Mishra and S. Jain, A Study of Various Approaches and Tools on Ontology, in 2015 IEEE International Conference on Computational Intelligence & Communication Technology , Ghaziabad, India, 2015, pp. 57–61 Search PubMed .
- A. S. Behr, M. Völkenrath and N. Kockmann, Ontology extension with NLP-based concept extraction for domain experts in catalytic sciences, Knowl. Inf. Syst. , 2023, 65 (12), 5503–5522, DOI: 10.1007/s10115-023-01919-1 .
- A. Zouaq, D. Gasevic and M. Hatala, Towards open ontology learning and filtering, Information Systems , 2011, 36 (7), 1064–1081, DOI: 10.1016/j.is.2011.03.005 .
- Y. Zhang, C. Wang, M. Soukaseum, D. G. Vlachos and H. Fang, Unleashing the Power of Knowledge Extraction from Scientific Literature in Catalysis, J. Chem. Inf. Model. , 2022, 62 (14), 3316–3330, DOI: 10.1021/acs.jcim.2c00359 .
- I. Beltagy, K. Lo and A. Cohan, SciBERT: A Pretrained Language Model for Scientific Text, EMNLP. [Online]. Available: https://arxiv.org/pdf/1903.10676.pdf.
- W3C Sparql 1.1. [Online]. Available: https://www.w3.org/TR/sparql11-update/.
- J. Hastings, et al. , ChEBI in 2016: Improved services and an expanding collection of metabolites, Nucleic Acids Res. , 2016, 44 (D1), D1214–D1219, DOI: 10.1093/nar/gkv1031 .
- CrossRef, CrossRef API Documentation. Accessed: 2024.
- S. Chamberlain, J. Maupetit, S. Peak, C. Talbert, D. Himmelstein and K. Niemeyer, Habanero: Python client for the Crossref API, 2024, Accessed: 2024. [Online]. Available: https://github.com/sckott/habanero.
- M. E. Rose and J. R. Kitchin, pybliometrics: Scriptable bibliometrics using a Python interface to Scopus, SoftwareX , 2019, 10 , 100263, DOI: 10.1016/j.softx.2019.100263 .
- M. Zhu and J. M. Cole, PDFDataExtractor: A Tool for Reading Scientific Text and Interpreting Metadata from the Typeset Literature in the Portable Document Format, J. Chem. Inf. Model. , 2022, 62 (7), 1633–1643, DOI: 10.1021/acs.jcim.1c01198 .
- M. C. Swain and J. M. Cole, ChemDataExtractor: A Toolkit for Automated Extraction of Chemical Information from the Scientific Literature, J. Chem. Inf. Model. , 2016, 56 (10), 1894–1904, DOI: 10.1021/acs.jcim.6b00207 .
- Python Software Foundation, re - Regular expression operations, 2024.
- S. Kim, et al. , PubChem 2023 update, Nucleic Acids Res. , 2023, 51 (D1), D1373–D1380, DOI: 10.1093/nar/gkac956 .
- M. Swain, PubChemPy: A way to interact with PubChem in Python, 2014, [Online]. Available: https://github.com/mcs07/PubChemPy.
- Allotrope Foundation, Allotrope Foundation Ontologies. Accessed: 2022.
- I. Montani et al. , spaCy: Industrial-strength Natural Language Processing in Python, 2022.
- A. S. Behr, H. Borgelt and N. Kockmann, Ontologies4Cat: investigating the landscape of ontologies for catalysis research data management, J. Cheminf. , 2024, 16 (1), 16, DOI: 10.1186/s13321-024-00807-2 .
- R. Arp, B. Smith and A. D. Spear, Building ontologies with Basic Formal Ontology , Massachusetts Institute of Technology, Cambridge, Massachusetts, 2015 Search PubMed .
- C. Batchelor, Molecular Process Ontology (MOP). [Online]. Available: https://github.com/rsc-ontologies/rxno.
- C. Batchelor, Chemical Reactions Ontology (RXNO). [Online]. Available: https://github.com/rsc-ontologies/rxno.
- R. C. Jackson, J. P. Balhoff, E. Douglass, N. L. Harris, C. J. Mungall and J. A. Overton, ROBOT: A Tool for Automating Ontology Workflows, BMC Bioinf. , 2019, 20 (1), 407, DOI: 10.1186/s12859-019-3002-3 .
- J.-B. Lamy, Owlready: Ontology-oriented programming in Python with automatic classification and high level constructs for biomedical ontologies, Artif. Intell. Med. , 2017, 80 , 11–28, DOI: 10.1016/j.artmed.2017.07.002 .
- P. Sun and S. Zhang, Identifying Granularity Differences between Large Biomedical Ontologies through Rules, AMIA Annu. Symp. Proc. , 2010, 2010 , 927–931 Search PubMed .
- P. Strömert, J. Hunold, A. Castro, S. Neumann and O. Koepler, Ontologies4Chem: the landscape of ontologies in chemistry, Pure Appl. Chem. , 2022, 94 (6), 605–622, DOI: 10.1515/pac-2021-2007 .
- SPARQL 1.1 Query Language , ed. E. Prud'hommeaux, S. Harris and A. Seaborne, W3C, 2013, [Online] Available: https://www.w3.org/TR/sparql11-query Search PubMed .
- W. McKinney, Data Structures for Statistical Computing in Python, in Proceedings of the 9th Python in Science Conference , Austin, Texas, 2010, pp. 56–61 Search PubMed .
- Y. Liu, et al. , Rhodium nanoparticles supported on silanol-rich zeolites beyond the homogeneous Wilkinson's catalyst for hydroformylation of olefins, Nat. Commun. , 2023, 14 (1), 2531, DOI: 10.1038/s41467-023-38181-6 .
- S. Hanf, L. Alvarado Rupflin, R. Gläser and S. Schunk, Current State of the Art of the Solid Rh-Based Catalyzed Hydroformylation of Short-Chain Olefins, Catalysts , 2020, 10 (5), 510, DOI: 10.3390/catal10050510 .
- K. Ghaib, K. Nitz and F.-Z. Ben-Fares, Chemical Methanation of CO 2 : A Review, ChemBioEng Rev. , 2016, 3 (6), 266–275, DOI: 10.1002/cben.201600022 .
- M. Tkachenko, M. Malyuk, A. Holmanyuk and N. Liubimov, Label Studio: Data labeling software.
- B. Motik, R. Shearer, G. Stoils and I. Horrocks, HermiT OWL Reasoner: The New Kid on the OWL Block, University of Oxford, Accessed: May 14 2022. [Online]. Available: https://www.hermit-reasoner.com/.
- A. S. Behr and D. Chernenko, CatalysisIE Knowledge Graph Generator. [Online]. Available: https://github.com/AleSteB/CatalysisIE_Knowledge_Graph_Generator.
| † Electronic supplementary information (ESI) available: https://github.com/AleSteB/CatalysisIE_Knowledge_Graph_Generator. See DOI: |

IMAGES
COMMENTS
Learn what an appendix is and how to use it in your research paper. Find out what to include, how to format, and how to refer to appendices in your text.
Appendix format example. The appendix label appears at the top of the page, bold and centered. On the next line, include a descriptive title, also bold and centered. The text is presented in general APA format: left-aligned, double-spaced, and with page numbers in the top right corner. Start a new page for each new appendix.
Learn how to write an appendix for a research paper, including types, purpose, and examples. An appendix is a section that contains supplementary material that is not essential to the main body of the paper but is helpful to the reader.
Research Paper Appendix | Example & Templates. Published on 15 August 2022 by Kirsten Dingemanse and Tegan George. Revised on 25 October 2022. An appendix is a supplementary document that facilitates your reader's understanding of your research but is not essential to your core argument. Appendices are a useful tool for providing additional information or clarification in a research paper ...
Learn how to create an appendix for your research paper and what to include in it. Find out how to format, organize, and cite your appendix with examples and tips.
The order they are presented is dictated by the order they are mentioned in the text of your research paper. The heading should be "Appendix," followed by a letter or number [e.g., "Appendix A" or "Appendix 1"], centered and written in bold type. If there is a table of contents, the appendices must be listed.
Begin each appendix on a separate page. At the top of the page, center the word Appendix and the identifying capital letters (A, B, etc.) in the order in which they are mentioned in the text. Center the title of the appendix using uppercase and lowercase letter on the next line. Begin the text of the appendix flush left, followed by indented ...
Label the appendices: Label each appendix with a capital letter (e.g., "Appendix A," "Appendix B," etc.) and provide a brief descriptive title that summarizes the content. F ormat the appendices: Follow the same formatting style as the rest of your paper or report. Use the same font, margins, and spacing to maintain consistency.
Learn the functions, tips, format and examples of writing an appendix for a research paper. An appendix is a supplementary section that includes raw data, tables, figures, maps and other supporting materials.
Information in this section is as outlined in the APA Publication Manual (2020), sections 2.14, 2.17, 2.24, and 7.6. Appendices are used to include information that supplement the paper's content but are considered distracting or inappropriate for the overall topic. It is recommended to only include an appendix if it helps the reader ...
An appendix is a supplemental section of a research paper that provides additional information, data, or materials to support the main content. The appendix is usually placed at the end of the document and is numbered with letters or numbers, such as "Appendix A," "Appendix B," etc.
4. Add page numbers. You should make sure the appendix has page numbers at the bottom right corner or the center of the page. Use the same page number formatting for the appendix that you used for the rest of the paper. Continue the numbering from the text into the appendix so it feels like part of the whole.
A research paper appendix can include a wide range of supporting materials, such as: Raw data sets or statistical tables that are too extensive for the main text. Detailed descriptions of research methodologies, instruments, or protocols. Interview transcripts or survey questionnaires. Correspondence with research participants or collaborators.
An appendix is an academic work section that contains additional information (statistics, references, tables, figures, etc.) that cannot be included in the main text. This component is usually placed after the reference list at the end of a research paper or dissertation.
Each appendix begins on a new page. The order they are presented is dictated by the order they are mentioned in the text of your research paper. The heading should be "Appendix," followed by a letter or number [e.g., "Appendix A" or "Appendix 1"], centered and written in bold. Appendices must be listed in the table of contents [if used].
The appendix of a paper consists of supporting information for the research that is not necessary to include in the text. This section provides further insight into the topic of research but happens to be too complex or too broad to add to the body of the paper. A paper can have more than one appendix, as it is recommended to divide them ...
An appendix in writing is a supplementary section that is included at the end of a document, such as a research paper, report, or book. It contains additional information that is relevant to the main text but not essential for understanding the core content.
An appendix is a section added to the end of a research paper to give readers extra information. Appendices are labeled with numbers or letters and are often a good place to include data that might be distracting in the main text. The word appendix comes from the root word append, a verb meaning "to attach or add.".
An appendix or appendices should always be inserted after your Reference List; however, the appropriateness of appendix content really depends on the nature and scope of your research paper. For a more in-depth review of what supplemental materials might be included in a social science appendix, be sure to review Section 2.14 "Appendices ...
Title of the appendix can be in the same format as the title of the other sections of your research paper or presentation. You can write it in the same font style and size. It can also be written in all capital letters, i.e. APPENDIX or in title or sentence case, i.e. Appendix. Use Appendix A, Appendix B, Appendix C and so on to give them a ...
Each appendix begins on a new page. The order they are presented is dictated by the order they are mentioned in the text of your research paper. The heading should be "Appendix," followed by a letter or number [e.g., "Appendix A" or "Appendix 1"], centered and written in bold type. If there is a table of contents, the appendices must be listed.
Then, gather all the information you need to place in your appendix and assess their relevance to the research paper. Remember, your purpose should not be to attach each and every little detail you found in your research. Next, organize your appendix; the order it should appear should be properly prearranged. Use sections, headings, subheading ...
The appendix is a section at the end of the paper with additional information that doesn't belong in the main text. It shouldn't have any arguments, quotes, or conclusions. You must use it to support the claims you already made in your essay — appendices help illustrate everything better.
A research paper explores and evaluates previously and newly gathered information on a topic, then offers evidence for an argument. It follows academic writing standards, and virtually every college student will write at least one. Research papers are also integral to scientific fields, among others, as the most reliable way to share knowledge.
We have explored index additions and deletions and associated issues in an array of research papers over the years, and our findings have inspired a new approach to cap-weighted indexing. 1 In each paper, we keep a narrow focus on one aspect of this work: the reconstitution of a cap-weighted index. The reconstitution puzzle is a rich and nuanced topic, and our analysis has revealed several ...
Based on the work of thousands of volunteers, TRB delivers an extensive research program; convenes leaders, practitioners, and academics from around the world; and provides timely policy advice on issues facing the transportation community.
If you've used ChatGPT or other AI tools in your research, describe how you used the tool in your Method section or in a comparable section of your paper. ... You may also put the full text of long responses from ChatGPT in an appendix of your paper or in online supplemental materials, so readers have access to the exact text that was ...
The search for definitive biosignatures—unambiguous markers of past or present life—is a central goal of paleobiology and astrobiology. We used pyrolysis-gas chromatography coupled to mass spectrometry to analyze chemically disparate samples, including living cells, geologically processed fossil organic material, carbon-rich meteorites, and laboratory-synthesized organic compounds and ...
Feature papers represent the most advanced research with significant potential for high impact in the field. A Feature Paper should be a substantial original Article that involves several techniques or approaches, provides an outlook for future research directions and describes possible research applications.
This allows for automated classification of research papers with regard to those classes, increasing FAIRness of the classified texts. ... Table 4 in the appendix lists the object properties and their inverse properties that need to be defined within the working ontology in order to assign the relations between the newly created entities.